Solver for Ordinary Differential Equations (ODE)
lsode.RdSolves the initial value problem for stiff or nonstiff systems of ordinary differential equations (ODE) in the form: $$dy/dt = f(t,y)$$.
The R function lsode provides an interface to the FORTRAN ODE
solver of the same name, written by Alan C. Hindmarsh and Andrew
H. Sherman.
It combines parts of the code lsodar and can thus find the root
of at least one of a set of constraint functions g(i) of the independent
and dependent variables. This can be used to stop the simulation or to
trigger events, i.e. a sudden change in one of the state variables.
The system of ODE's is written as an R function or be defined in compiled code that has been dynamically loaded.
In contrast to lsoda, the user has to specify whether or
not the problem is stiff and choose the appropriate solution method.
lsode is very similar to vode, but uses a
fixed-step-interpolate method rather than the variable-coefficient
method in vode. In addition, in vode it is
possible to choose whether or not a copy of the Jacobian is saved for
reuse in the corrector iteration algorithm; In lsode, a copy is
not kept.
Usage
lsode(y, times, func, parms, rtol = 1e-6, atol = 1e-6,
jacfunc = NULL, jactype = "fullint", mf = NULL, rootfunc = NULL,
verbose = FALSE, nroot = 0, tcrit = NULL, hmin = 0, hmax = NULL,
hini = 0, ynames = TRUE, maxord = NULL, bandup = NULL, banddown = NULL,
maxsteps = 5000, dllname = NULL, initfunc = dllname,
initpar = parms, rpar = NULL, ipar = NULL, nout = 0,
outnames = NULL, forcings=NULL, initforc = NULL,
fcontrol=NULL, events=NULL, lags = NULL,...)Arguments
- y
the initial (state) values for the ODE system. If
yhas a name attribute, the names will be used to label the output matrix.- times
time sequence for which output is wanted; the first value of
timesmust be the initial time; if only one step is to be taken; settimes=NULL.- func
either an R-function that computes the values of the derivatives in the ODE system (the model definition) at time t, or a character string giving the name of a compiled function in a dynamically loaded shared library.
If
funcis an R-function, it must be defined as:func <- function(t, y, parms,...).tis the current time point in the integration,yis the current estimate of the variables in the ODE system. If the initial valuesyhas anamesattribute, the names will be available insidefunc.parmsis a vector or list of parameters; ... (optional) are any other arguments passed to the function.The return value of
funcshould be a list, whose first element is a vector containing the derivatives ofywith respect totime, and whose next elements are global values that are required at each point intimes. The derivatives must be specified in the same order as the state variablesy.If
funcis a string, thendllnamemust give the name of the shared library (without extension) which must be loaded beforelsode()is called. See package vignette"compiledCode"for more details.- parms
vector or list of parameters used in
funcorjacfunc.- rtol
relative error tolerance, either a scalar or an array as long as
y. See details.- atol
absolute error tolerance, either a scalar or an array as long as
y. See details.- jacfunc
if not
NULL, an R function that computes the Jacobian of the system of differential equations \(\partial\dot{y}_i/\partial y_j\), or a string giving the name of a function or subroutine indllnamethat computes the Jacobian (see vignette"compiledCode"for more about this option).In some circumstances, supplying
jacfunccan speed up the computations, if the system is stiff. The R calling sequence forjacfuncis identical to that offunc.If the Jacobian is a full matrix,
jacfuncshould return a matrix \(\partial\dot{y}/\partial y\), where the ith row contains the derivative of \(dy_i/dt\) with respect to \(y_j\), or a vector containing the matrix elements by columns (the way R and FORTRAN store matrices).
If the Jacobian is banded,jacfuncshould return a matrix containing only the nonzero bands of the Jacobian, rotated row-wise. See first example of lsode.- jactype
the structure of the Jacobian, one of
"fullint","fullusr","bandusr"or"bandint"- either full or banded and estimated internally or by user; overruled ifmfis notNULL.- mf
the "method flag" passed to function lsode - overrules
jactype- provides more options thanjactype- see details.- rootfunc
if not
NULL, an R function that computes the function whose root has to be estimated or a string giving the name of a function or subroutine indllnamethat computes the root function. The R calling sequence forrootfuncis identical to that offunc.rootfuncshould return a vector with the function values whose root is sought.- verbose
if TRUE: full output to the screen, e.g. will print the
diagnostiscsof the integration - see details.- nroot
only used if
dllnameis specified: the number of constraint functions whose roots are desired during the integration; ifrootfuncis an R-function, the solver estimates the number of roots.- tcrit
if not
NULL, thenlsodecannot integrate pasttcrit. The FORTRAN routinelsodeovershoots its targets (times points in the vectortimes), and interpolates values for the desired time points. If there is a time beyond which integration should not proceed (perhaps because of a singularity), that should be provided intcrit.- hmin
an optional minimum value of the integration stepsize. In special situations this parameter may speed up computations with the cost of precision. Don't use
hminif you don't know why!- hmax
an optional maximum value of the integration stepsize. If not specified,
hmaxis set to the largest difference intimes, to avoid that the simulation possibly ignores short-term events. If 0, no maximal size is specified.- hini
initial step size to be attempted; if 0, the initial step size is determined by the solver.
- ynames
logical, if
FALSEnames of state variables are not passed to functionfunc; this may speed up the simulation especially for multi-D models.- maxord
the maximum order to be allowed.
NULLuses the default, i.e. order 12 if implicit Adams method (meth = 1), order 5 if BDF method (meth = 2). Reduce maxord to save storage space.- bandup
number of non-zero bands above the diagonal, in case the Jacobian is banded.
- banddown
number of non-zero bands below the diagonal, in case the Jacobian is banded.
- maxsteps
maximal number of steps per output interval taken by the solver.
- dllname
a string giving the name of the shared library (without extension) that contains all the compiled function or subroutine definitions refered to in
funcandjacfunc. See package vignette"compiledCode".- initfunc
if not
NULL, the name of the initialisation function (which initialises values of parameters), as provided indllname. See package vignette"compiledCode".- initpar
only when
dllnameis specified and an initialisation functioninitfuncis in the dll: the parameters passed to the initialiser, to initialise the common blocks (FORTRAN) or global variables (C, C++).- rpar
only when
dllnameis specified: a vector with double precision values passed to the dll-functions whose names are specified byfuncandjacfunc.- ipar
only when
dllnameis specified: a vector with integer values passed to the dll-functions whose names are specified byfuncandjacfunc.- nout
only used if
dllnameis specified and the model is defined in compiled code: the number of output variables calculated in the compiled functionfunc, present in the shared library. Note: it is not automatically checked whether this is indeed the number of output variables calculated in the dll - you have to perform this check in the code - See package vignette"compiledCode".- outnames
only used if
dllnameis specified andnout> 0: the names of output variables calculated in the compiled functionfunc, present in the shared library. These names will be used to label the output matrix.- forcings
only used if
dllnameis specified: a list with the forcing function data sets, each present as a two-columned matrix, with (time,value); interpolation outside the interval [min(times), max(times)] is done by taking the value at the closest data extreme.See forcings or package vignette
"compiledCode".- initforc
if not
NULL, the name of the forcing function initialisation function, as provided indllname. It MUST be present ifforcingshas been given a value. See forcings or package vignette"compiledCode".- fcontrol
A list of control parameters for the forcing functions. See forcings or vignette
compiledCode.- events
A matrix or data frame that specifies events, i.e. when the value of a state variable is suddenly changed. See events for more information.
- lags
A list that specifies timelags, i.e. the number of steps that has to be kept. To be used for delay differential equations. See timelags, dede for more information.
- ...
additional arguments passed to
funcandjacfuncallowing this to be a generic function.
Value
A matrix of class deSolve with up to as many rows as elements
in times and as many columns as elements in y plus the number of "global"
values returned in the next elements of the return from func,
plus and additional column for the time value. There will be a row
for each element in times unless the FORTRAN routine `lsode'
returns with an unrecoverable error. If y has a names
attribute, it will be used to label the columns of the output value.
References
Alan C. Hindmarsh, "ODEPACK, A Systematized Collection of ODE Solvers," in Scientific Computing, R. S. Stepleman, et al., Eds. (North-Holland, Amsterdam, 1983), pp. 55-64.
Details
The work is done by the FORTRAN subroutine lsode, whose
documentation should be consulted for details (it is included as
comments in the source file src/opkdmain.f). The implementation
is based on the November, 2003 version of lsode, from Netlib.
Before using the integrator lsode, the user has to decide
whether or not the problem is stiff.
If the problem is nonstiff, use method flag mf = 10, which
selects a nonstiff (Adams) method, no Jacobian used.
If the problem
is stiff, there are four standard choices which can be specified with
jactype or mf.
The options for jactype are
- jactype = "fullint"
a full Jacobian, calculated internally by lsode, corresponds to
mf= 22,- jactype = "fullusr"
a full Jacobian, specified by user function
jacfunc, corresponds tomf= 21,- jactype = "bandusr"
a banded Jacobian, specified by user function
jacfunc; the size of the bands specified bybandupandbanddown, corresponds tomf= 24,- jactype = "bandint"
a banded Jacobian, calculated by lsode; the size of the bands specified by
bandupandbanddown, corresponds tomf= 25.
More options are available when specifying mf directly.
The
legal values of mf are 10, 11, 12, 13, 14, 15, 20, 21, 22, 23,
24, 25.mf is a positive two-digit integer, mf =
(10*METH + MITER), where
- METH
indicates the basic linear multistep method: METH = 1 means the implicit Adams method. METH = 2 means the method based on backward differentiation formulas (BDF-s).
- MITER
indicates the corrector iteration method: MITER = 0 means functional iteration (no Jacobian matrix is involved). MITER = 1 means chord iteration with a user-supplied full (NEQ by NEQ) Jacobian. MITER = 2 means chord iteration with an internally generated (difference quotient) full Jacobian (using NEQ extra calls to
funcper df/dy value). MITER = 3 means chord iteration with an internally generated diagonal Jacobian approximation (using 1 extra call tofuncper df/dy evaluation). MITER = 4 means chord iteration with a user-supplied banded Jacobian. MITER = 5 means chord iteration with an internally generated banded Jacobian (using ML+MU+1 extra calls tofuncper df/dy evaluation).
If MITER = 1 or 4, the user must supply a subroutine jacfunc.
Inspection of the example below shows how to specify both a banded and full Jacobian.
The input parameters rtol, and atol determine the
error control performed by the solver. See lsoda
for details.
The diagnostics of the integration can be printed to screen
by calling diagnostics. If verbose = TRUE,
the diagnostics will written to the screen at the end of the integration.
See vignette("deSolve") for an explanation of each element in the vectors containing the diagnostic properties and how to directly access them.
Models may be defined in compiled C or FORTRAN code, as well as
in an R-function. See package vignette "compiledCode" for details.
More information about models defined in compiled code is in the package vignette ("compiledCode"); information about linking forcing functions to compiled code is in forcings.
Examples in both C and FORTRAN are in the dynload subdirectory
of the deSolve package directory.
lsode can find the root of at least one of a set of constraint functions
rootfunc of the independent and dependent variables. It then returns the
solution at the root if that occurs sooner than the specified stop
condition, and otherwise returns the solution according the specified
stop condition.
Caution: Because of numerical errors in the function
rootfun due to roundoff and integration error, lsode may
return false roots, or return the same root at two or more
nearly equal values of time.
See also
rk,lsoda,lsodes,lsodar,vode,daspkfor other solvers of the Livermore family,odefor a general interface to most of the ODE solvers,ode.bandfor solving models with a banded Jacobian,ode.1Dfor integrating 1-D models,ode.2Dfor integrating 2-D models,ode.3Dfor integrating 3-D models,
diagnostics to print diagnostic messages.
Examples
## =======================================================================
## Example 1:
## Various ways to solve the same model.
## =======================================================================
## the model, 5 state variables
f1 <- function (t, y, parms) {
ydot <- vector(len = 5)
ydot[1] <- 0.1*y[1] -0.2*y[2]
ydot[2] <- -0.3*y[1] +0.1*y[2] -0.2*y[3]
ydot[3] <- -0.3*y[2] +0.1*y[3] -0.2*y[4]
ydot[4] <- -0.3*y[3] +0.1*y[4] -0.2*y[5]
ydot[5] <- -0.3*y[4] +0.1*y[5]
return(list(ydot))
}
## the Jacobian, written as a full matrix
fulljac <- function (t, y, parms) {
jac <- matrix(nrow = 5, ncol = 5, byrow = TRUE,
data = c(0.1, -0.2, 0 , 0 , 0 ,
-0.3, 0.1, -0.2, 0 , 0 ,
0 , -0.3, 0.1, -0.2, 0 ,
0 , 0 , -0.3, 0.1, -0.2,
0 , 0 , 0 , -0.3, 0.1))
return(jac)
}
## the Jacobian, written in banded form
bandjac <- function (t, y, parms) {
jac <- matrix(nrow = 3, ncol = 5, byrow = TRUE,
data = c( 0 , -0.2, -0.2, -0.2, -0.2,
0.1, 0.1, 0.1, 0.1, 0.1,
-0.3, -0.3, -0.3, -0.3, 0))
return(jac)
}
## initial conditions and output times
yini <- 1:5
times <- 1:20
## default: stiff method, internally generated, full Jacobian
out <- lsode(yini, times, f1, parms = 0, jactype = "fullint")
## stiff method, user-generated full Jacobian
out2 <- lsode(yini, times, f1, parms = 0, jactype = "fullusr",
jacfunc = fulljac)
## stiff method, internally-generated banded Jacobian
## one nonzero band above (up) and below(down) the diagonal
out3 <- lsode(yini, times, f1, parms = 0, jactype = "bandint",
bandup = 1, banddown = 1)
## stiff method, user-generated banded Jacobian
out4 <- lsode(yini, times, f1, parms = 0, jactype = "bandusr",
jacfunc = bandjac, bandup = 1, banddown = 1)
## non-stiff method
out5 <- lsode(yini, times, f1, parms = 0, mf = 10)
## =======================================================================
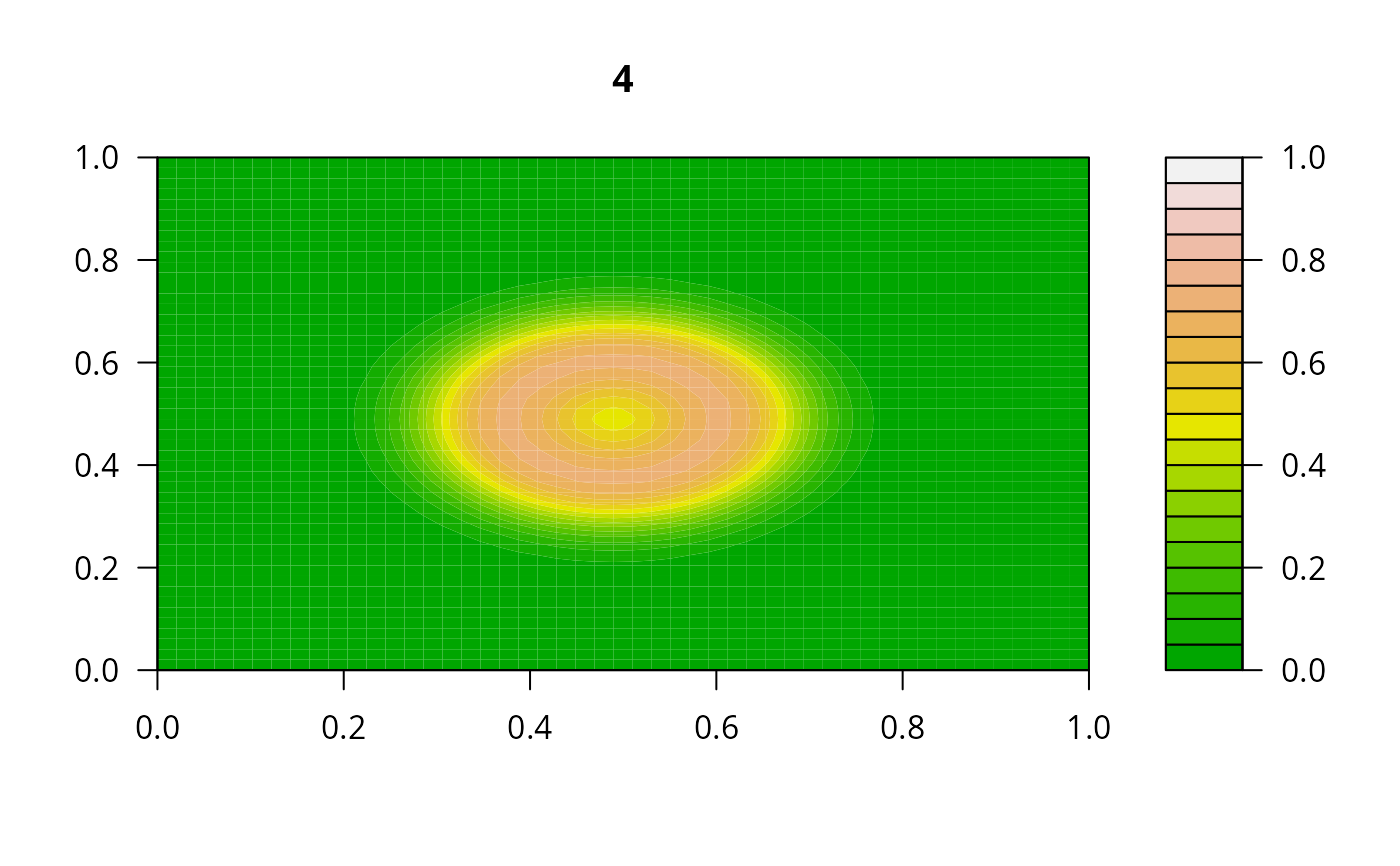
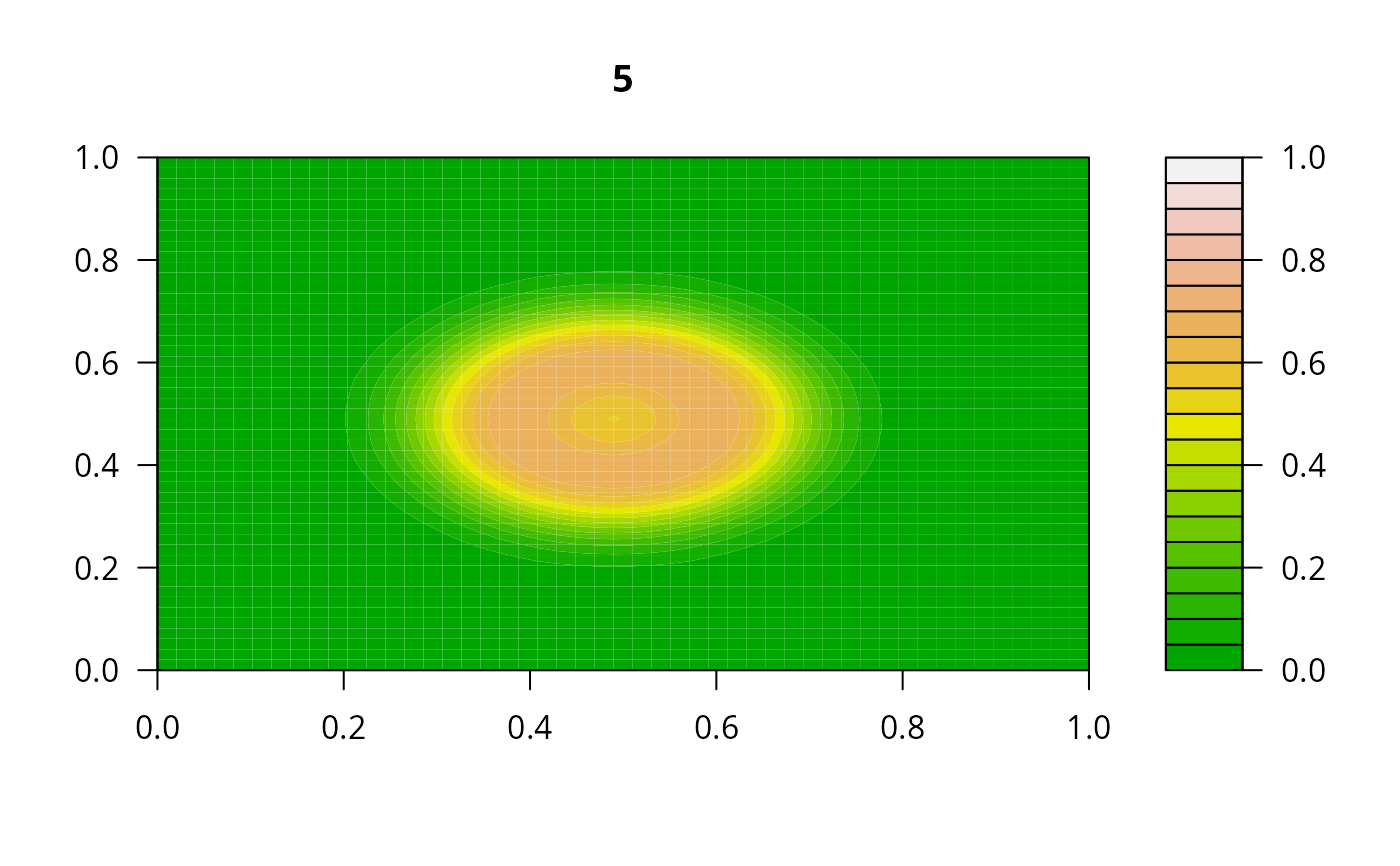
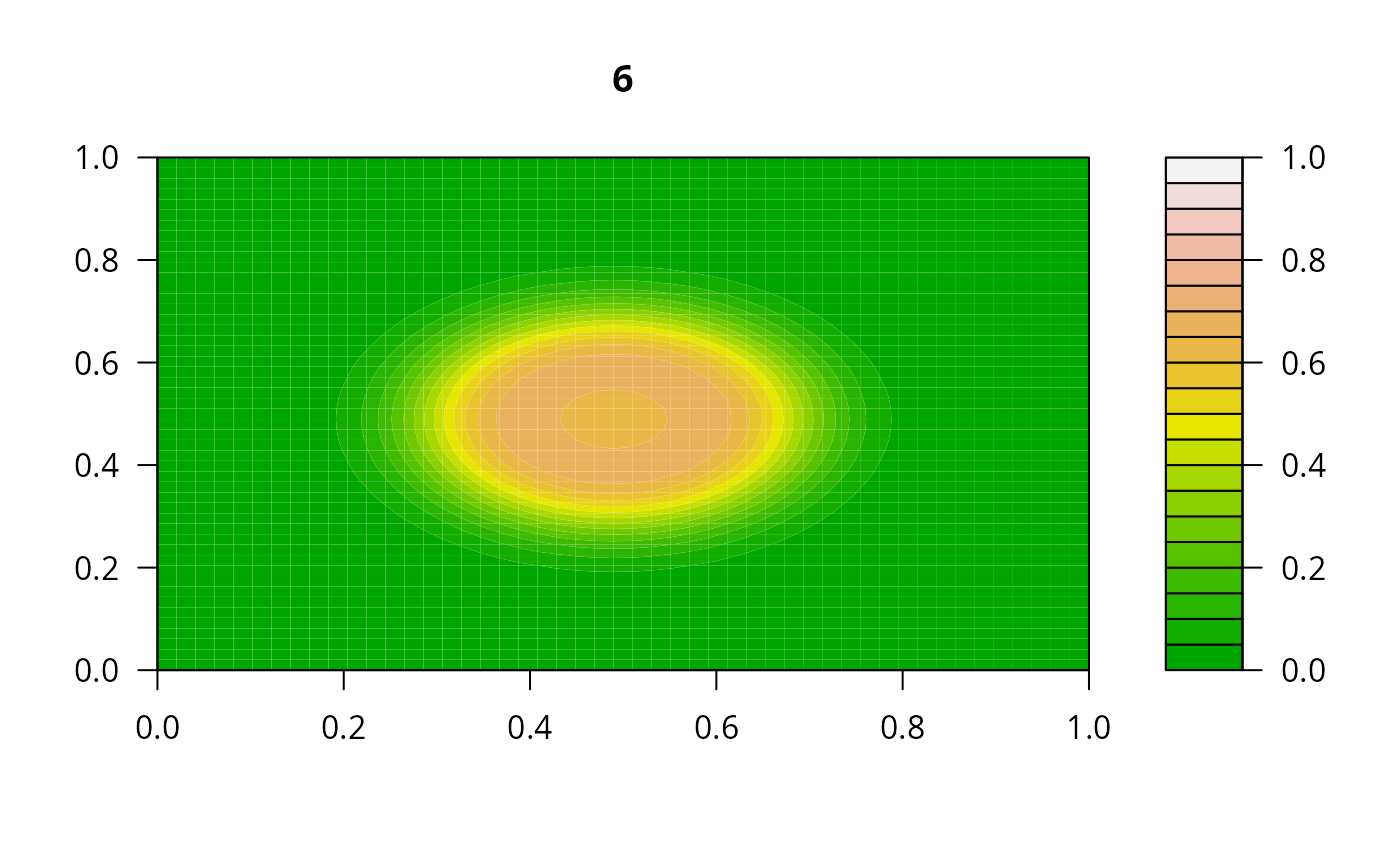
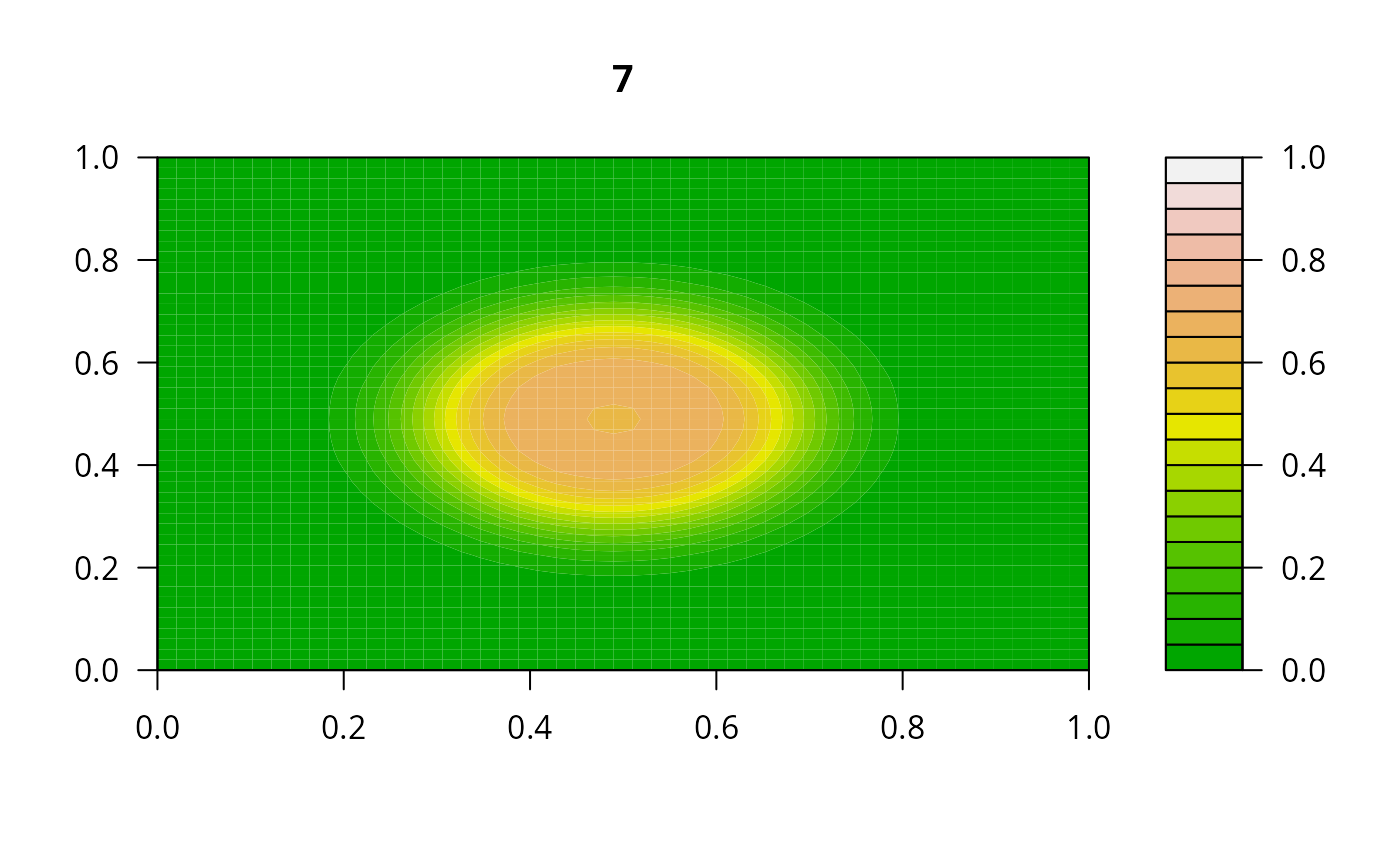
## Example 2:
## diffusion on a 2-D grid
## partially specified Jacobian
## =======================================================================
diffusion2D <- function(t, Y, par) {
y <- matrix(nrow = n, ncol = n, data = Y)
dY <- r*y # production
## diffusion in X-direction; boundaries = 0-concentration
Flux <- -Dx * rbind(y[1,],(y[2:n,]-y[1:(n-1),]),-y[n,])/dx
dY <- dY - (Flux[2:(n+1),]-Flux[1:n,])/dx
## diffusion in Y-direction
Flux <- -Dy * cbind(y[,1],(y[,2:n]-y[,1:(n-1)]),-y[,n])/dy
dY <- dY - (Flux[,2:(n+1)]-Flux[,1:n])/dy
return(list(as.vector(dY)))
}
## parameters
dy <- dx <- 1 # grid size
Dy <- Dx <- 1 # diffusion coeff, X- and Y-direction
r <- 0.025 # production rate
times <- c(0, 1)
n <- 50
y <- matrix(nrow = n, ncol = n, 0)
pa <- par(ask = FALSE)
## initial condition
for (i in 1:n) {
for (j in 1:n) {
dst <- (i - n/2)^2 + (j - n/2)^2
y[i, j] <- max(0, 1 - 1/(n*n) * (dst - n)^2)
}
}
filled.contour(y, color.palette = terrain.colors)
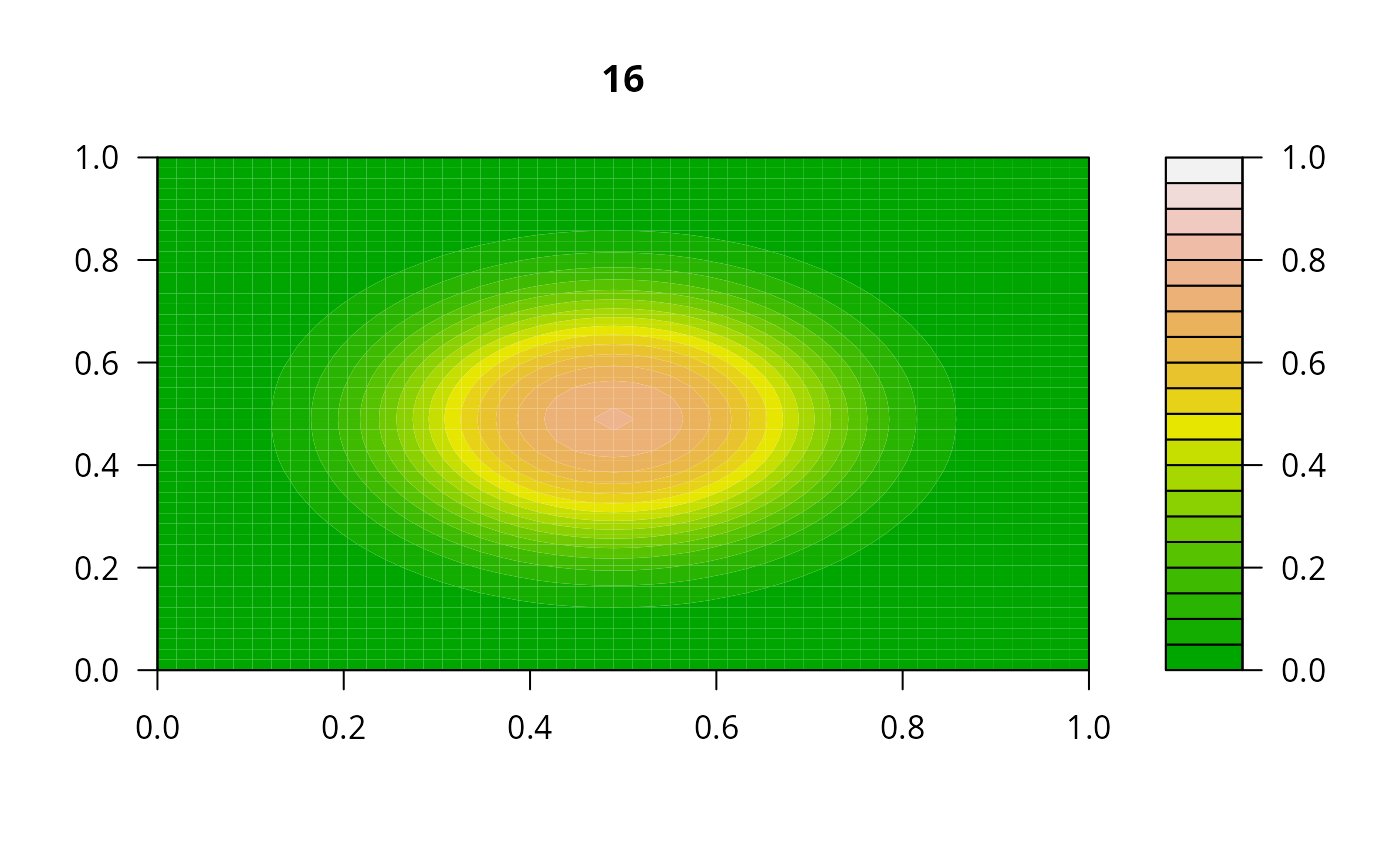
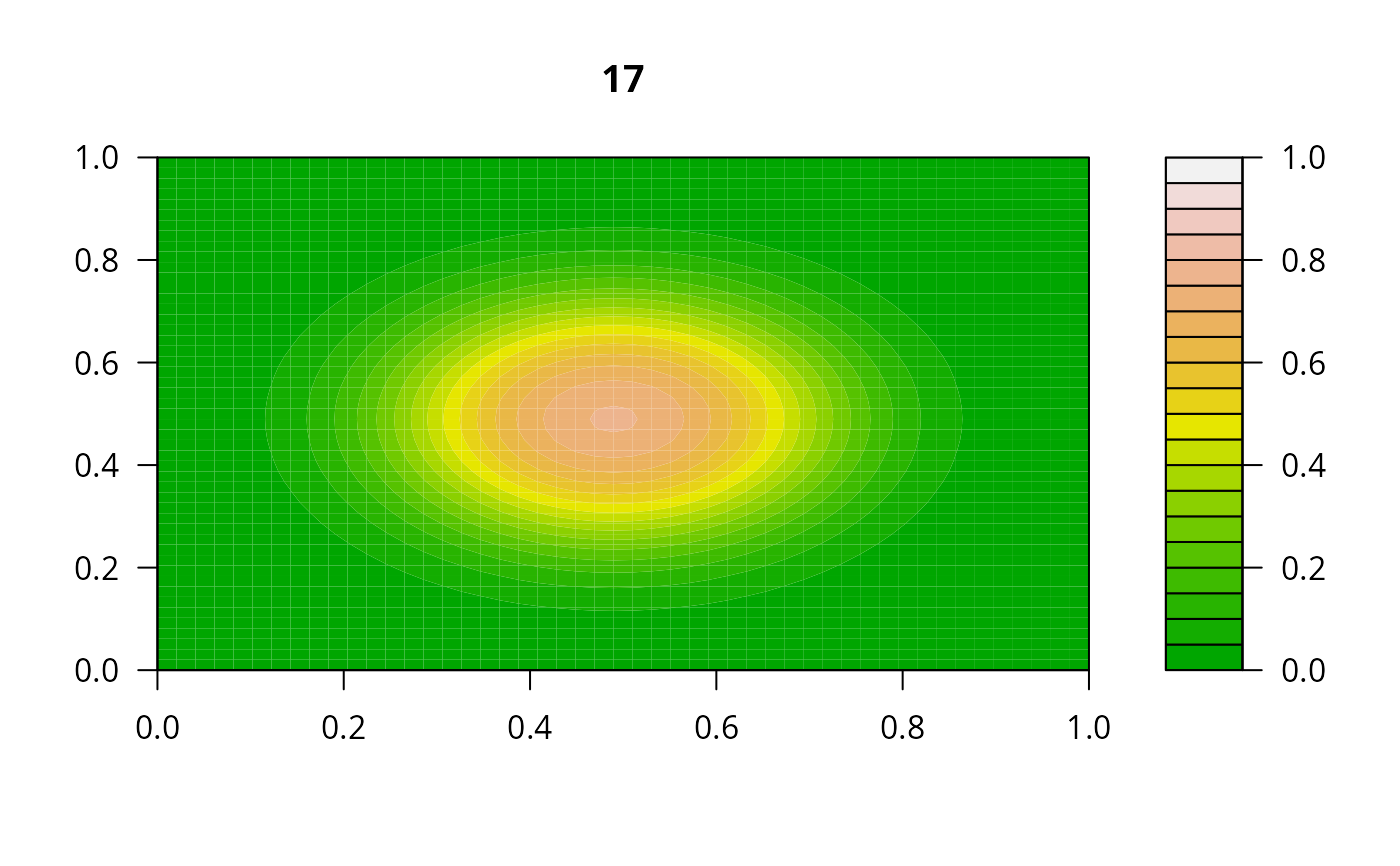
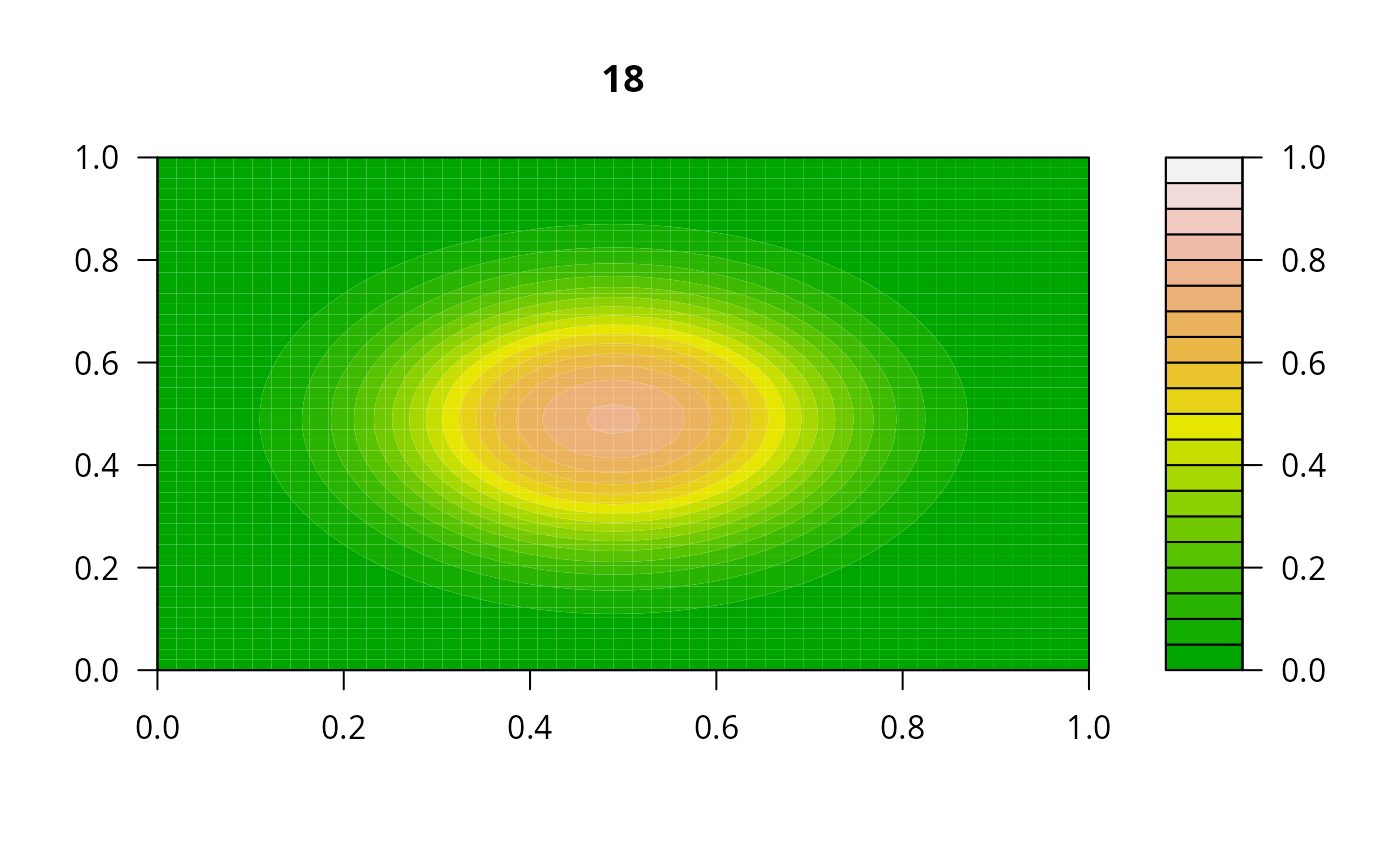
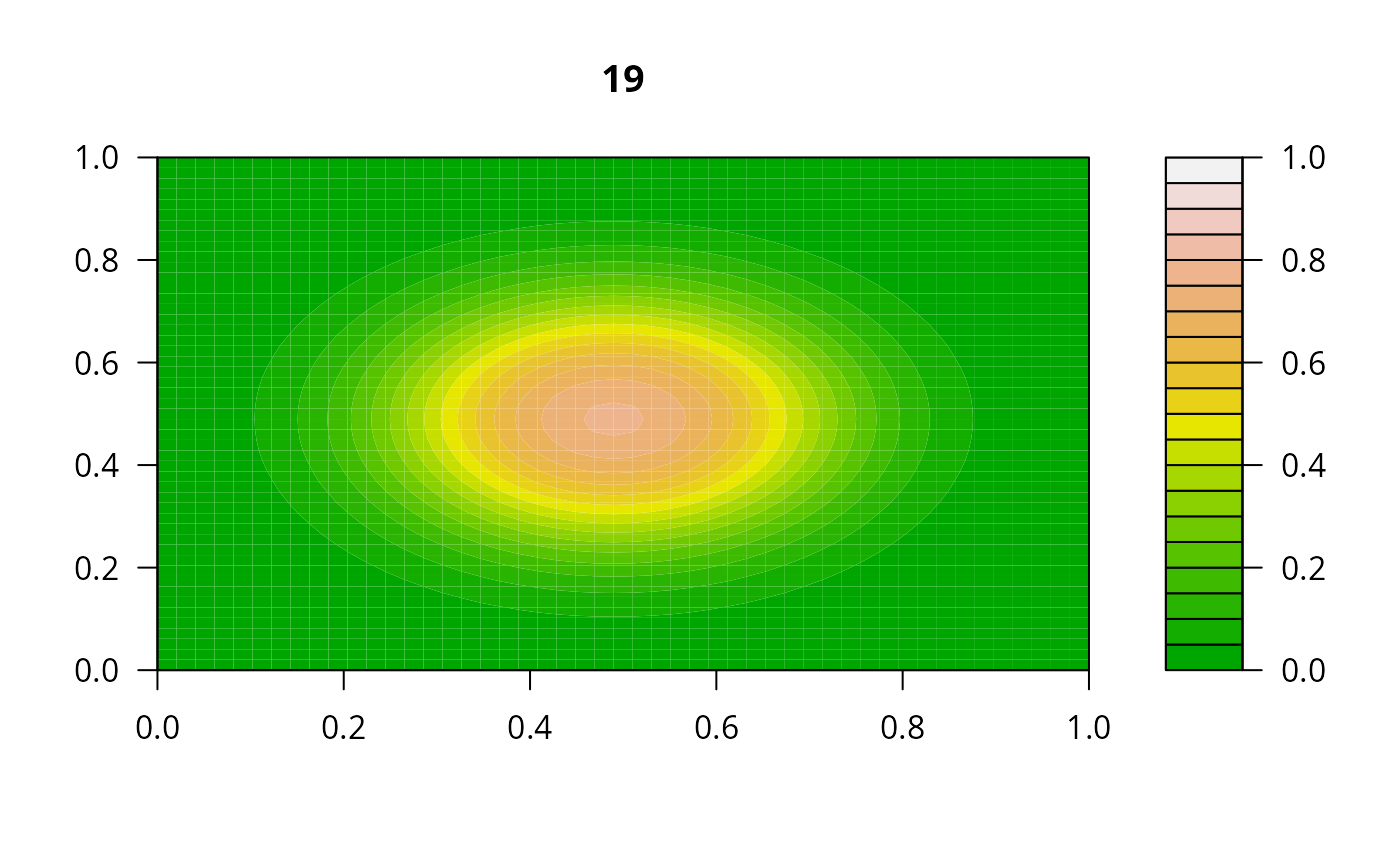
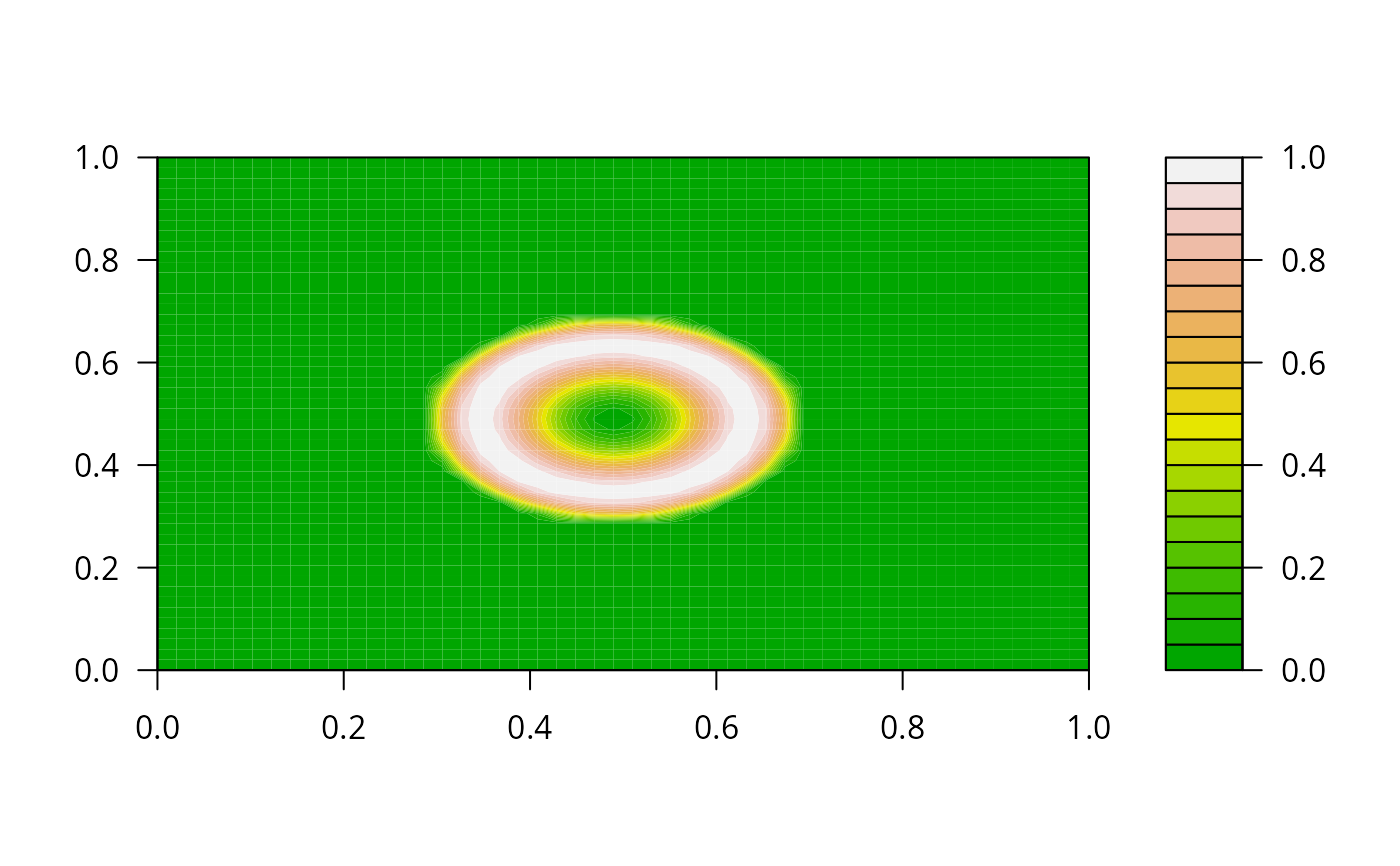 ## =======================================================================
## jacfunc need not be estimated exactly
## a crude approximation, with a smaller bandwidth will do.
## Here the half-bandwidth 1 is used, whereas the true
## half-bandwidths are equal to n.
## This corresponds to ignoring the y-direction coupling in the ODEs.
## =======================================================================
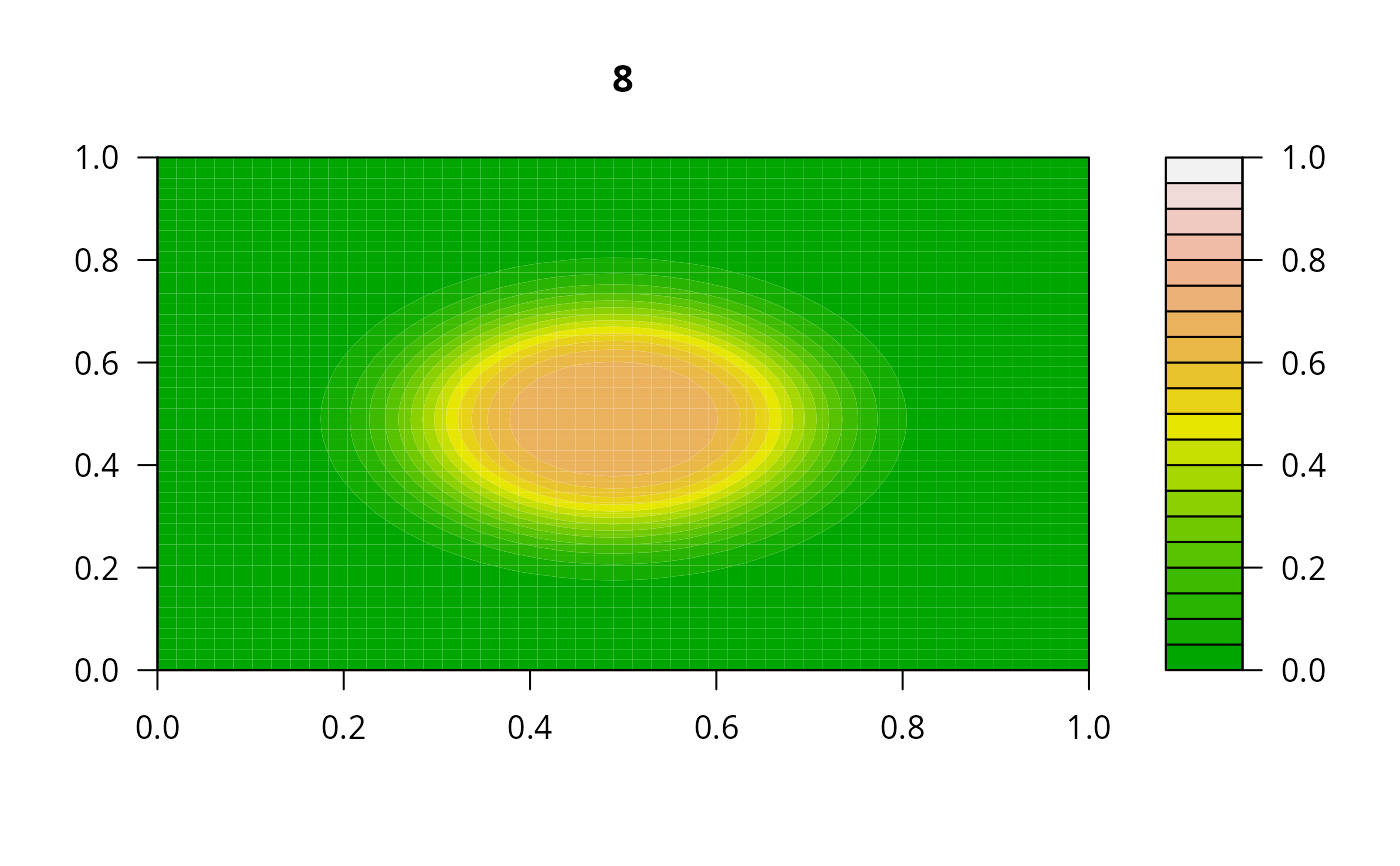
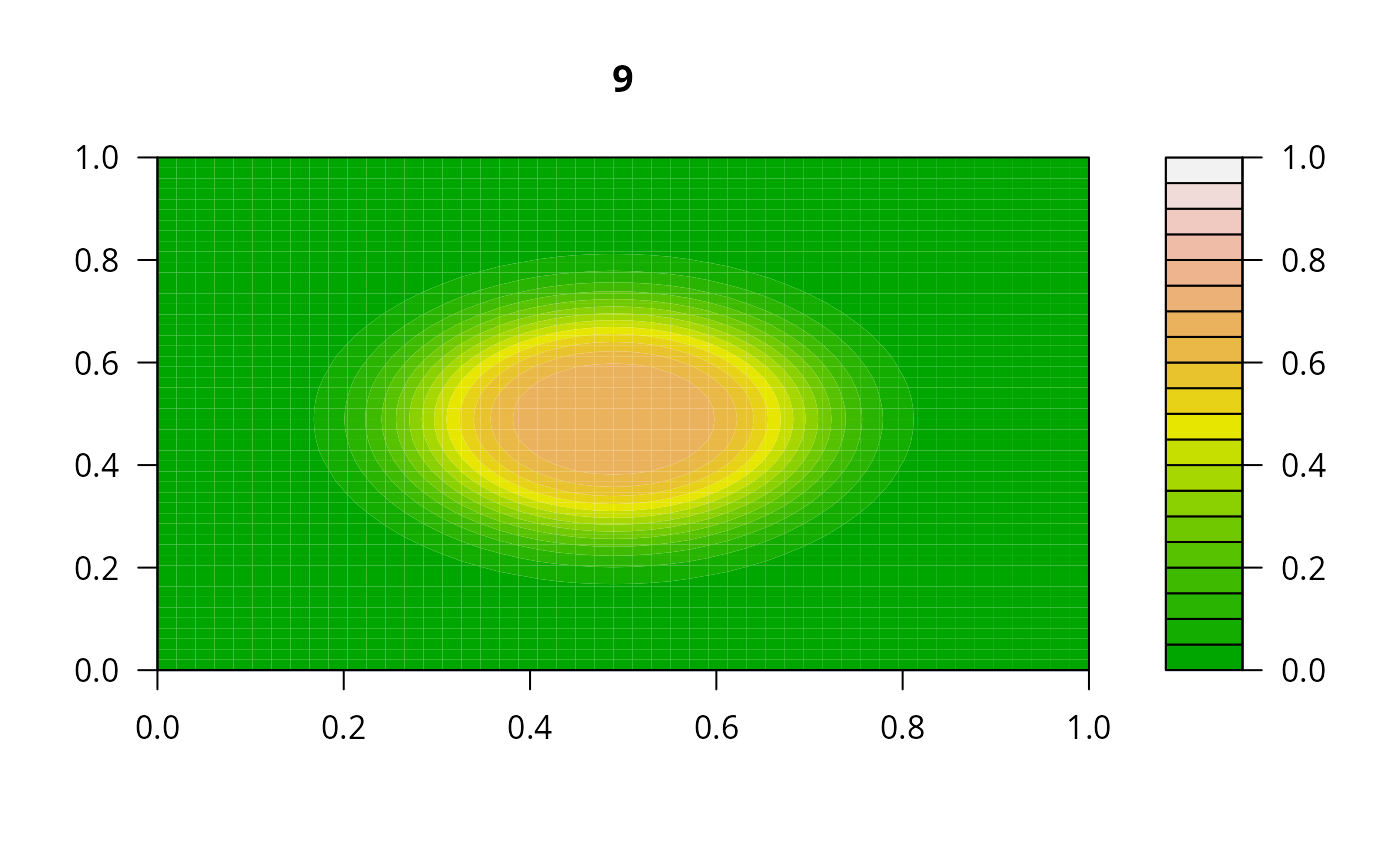
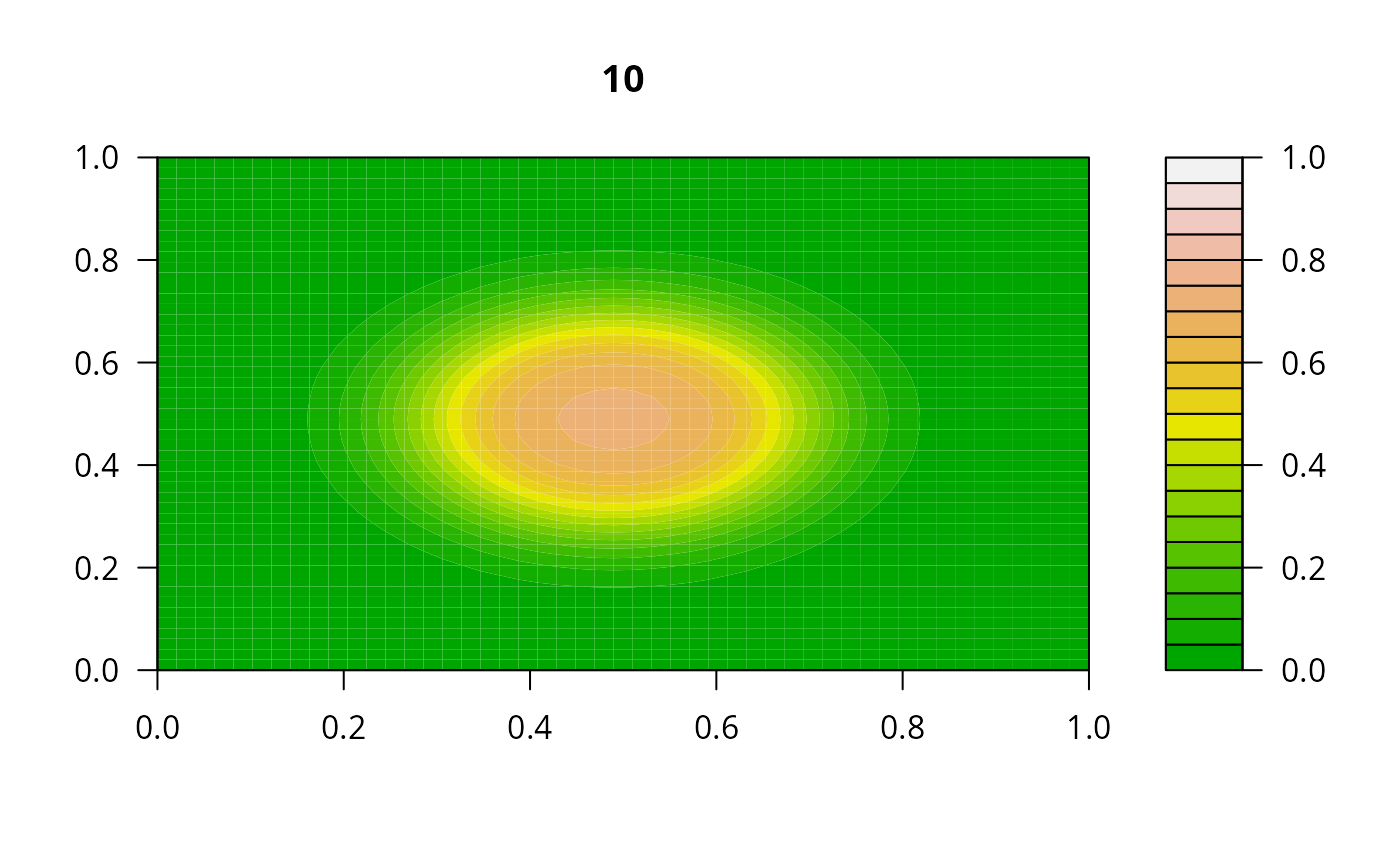
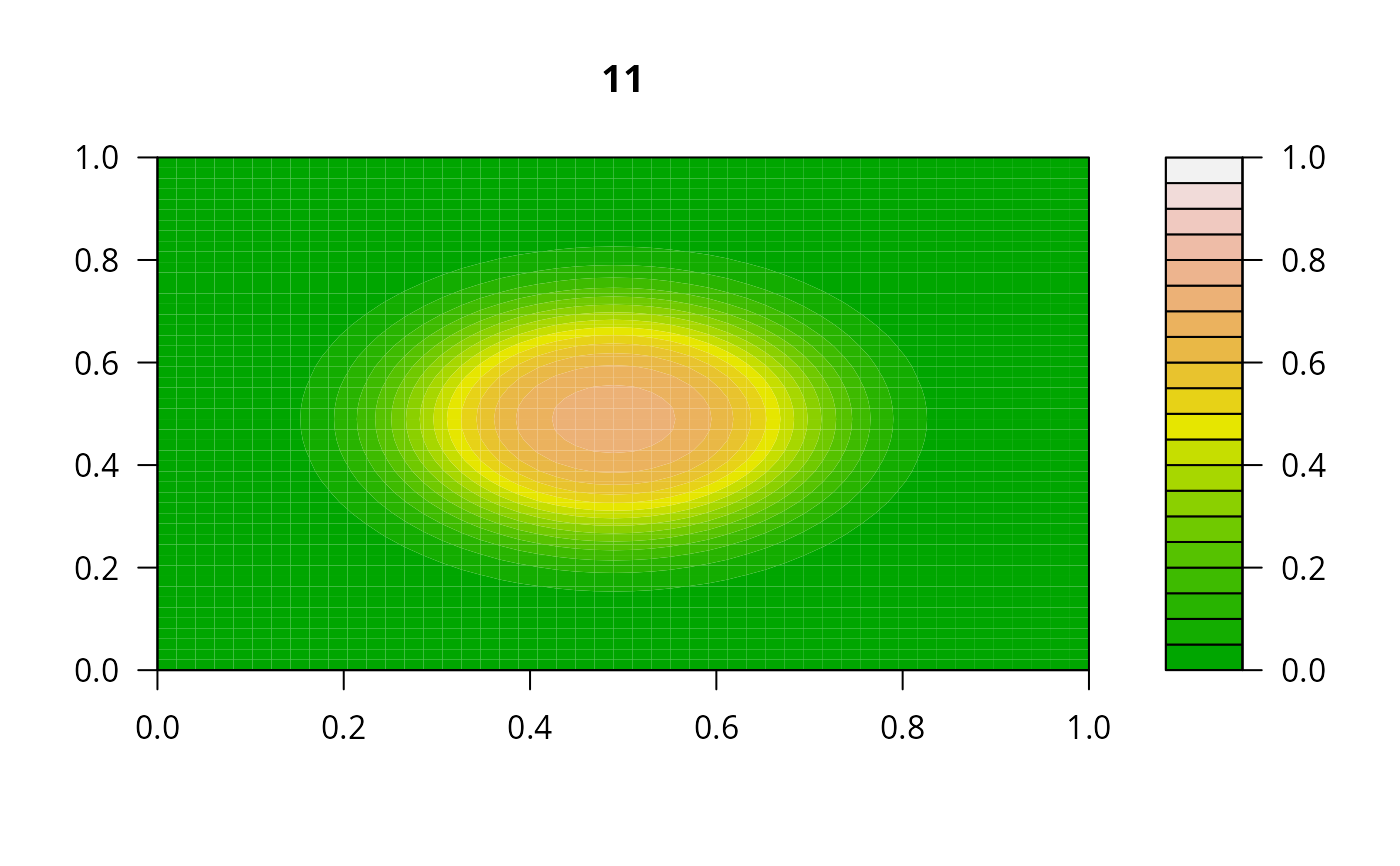
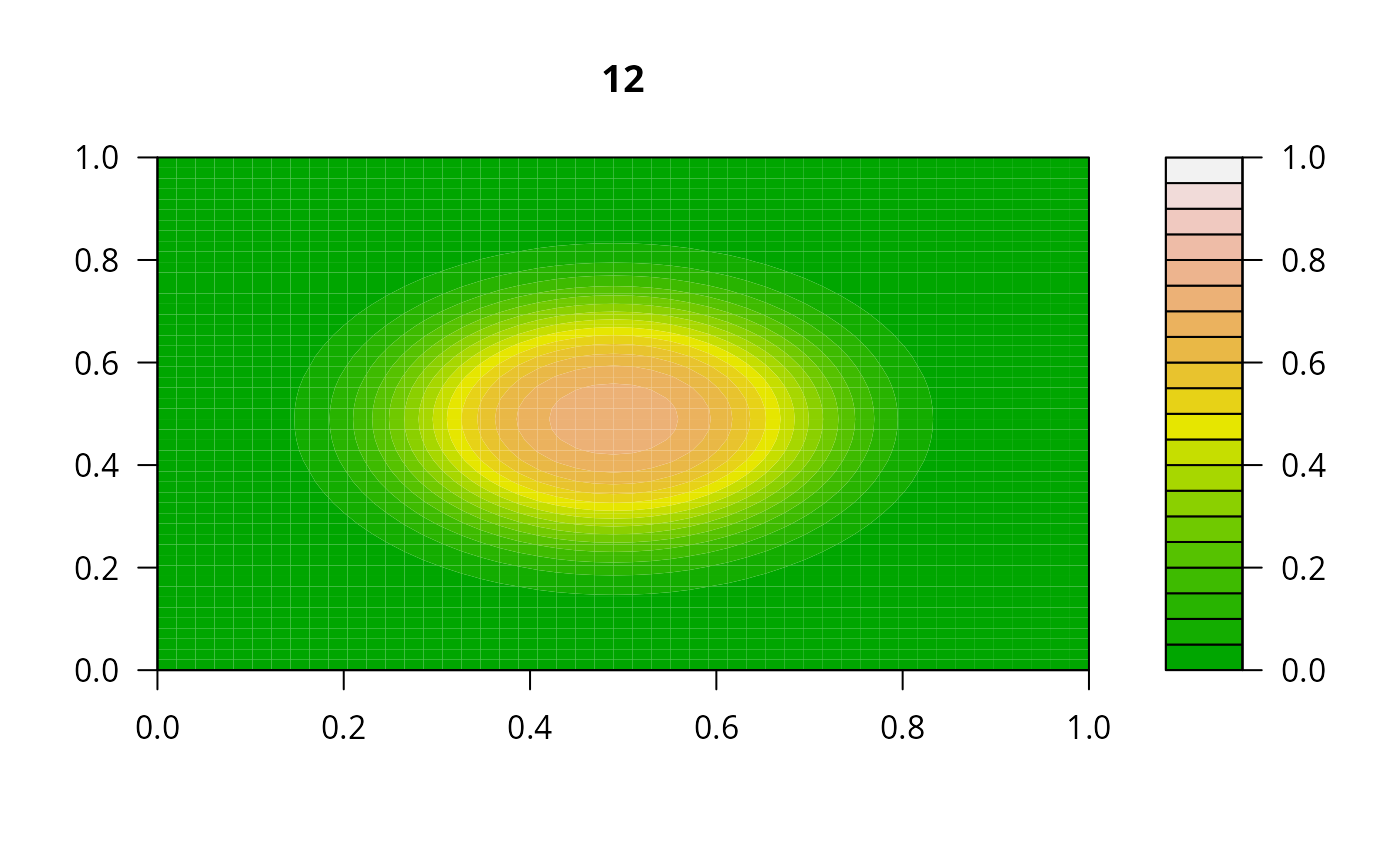
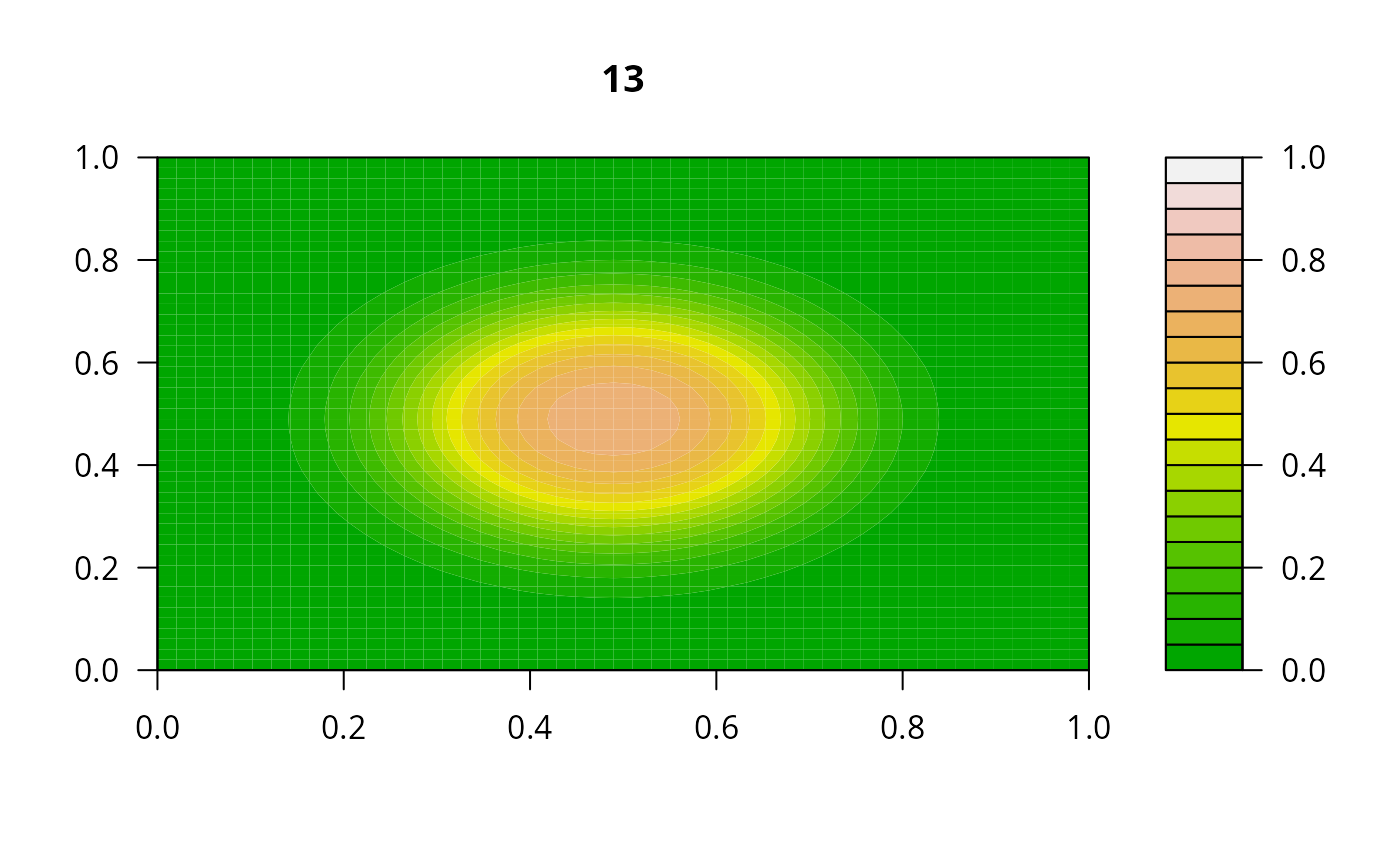
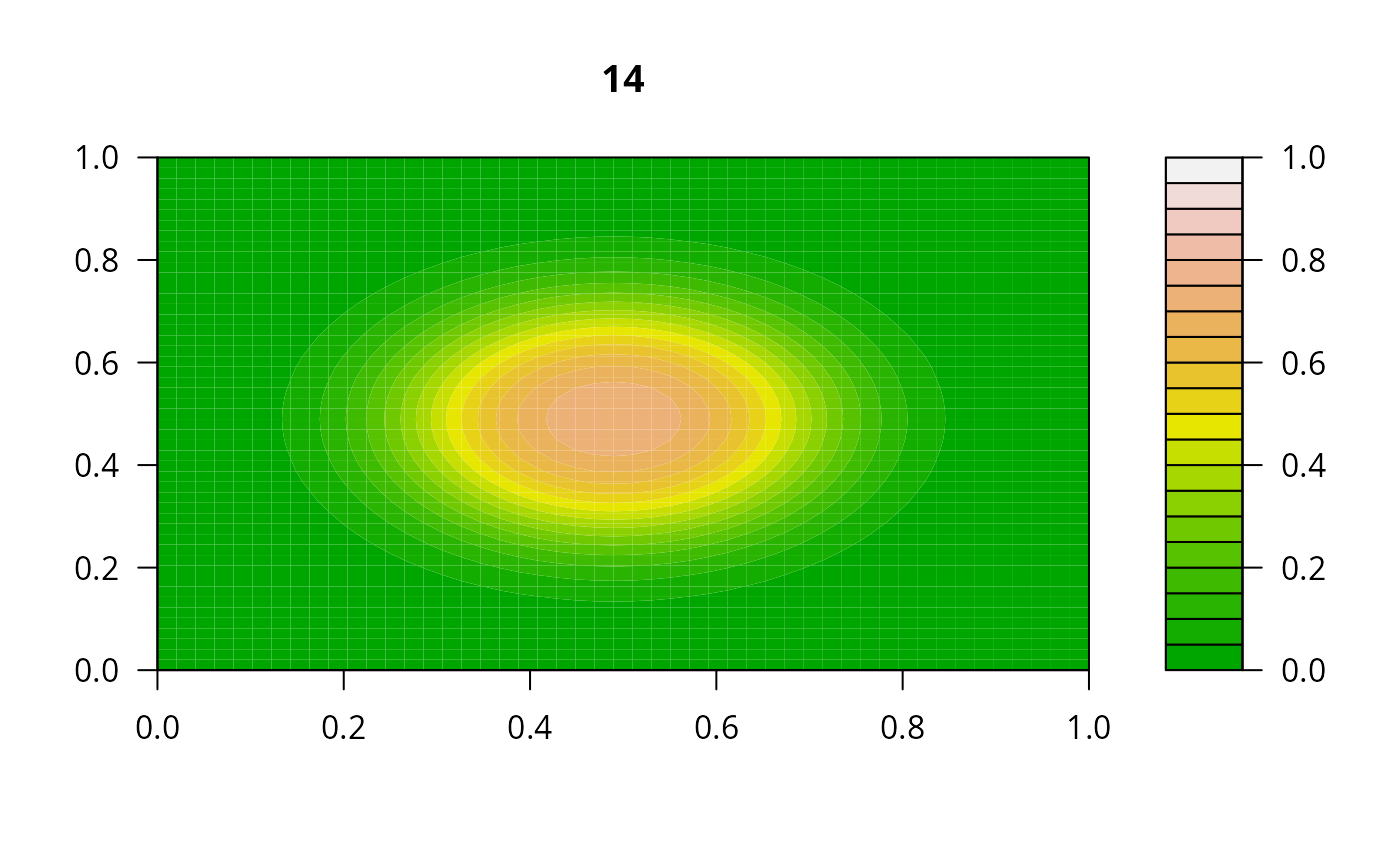
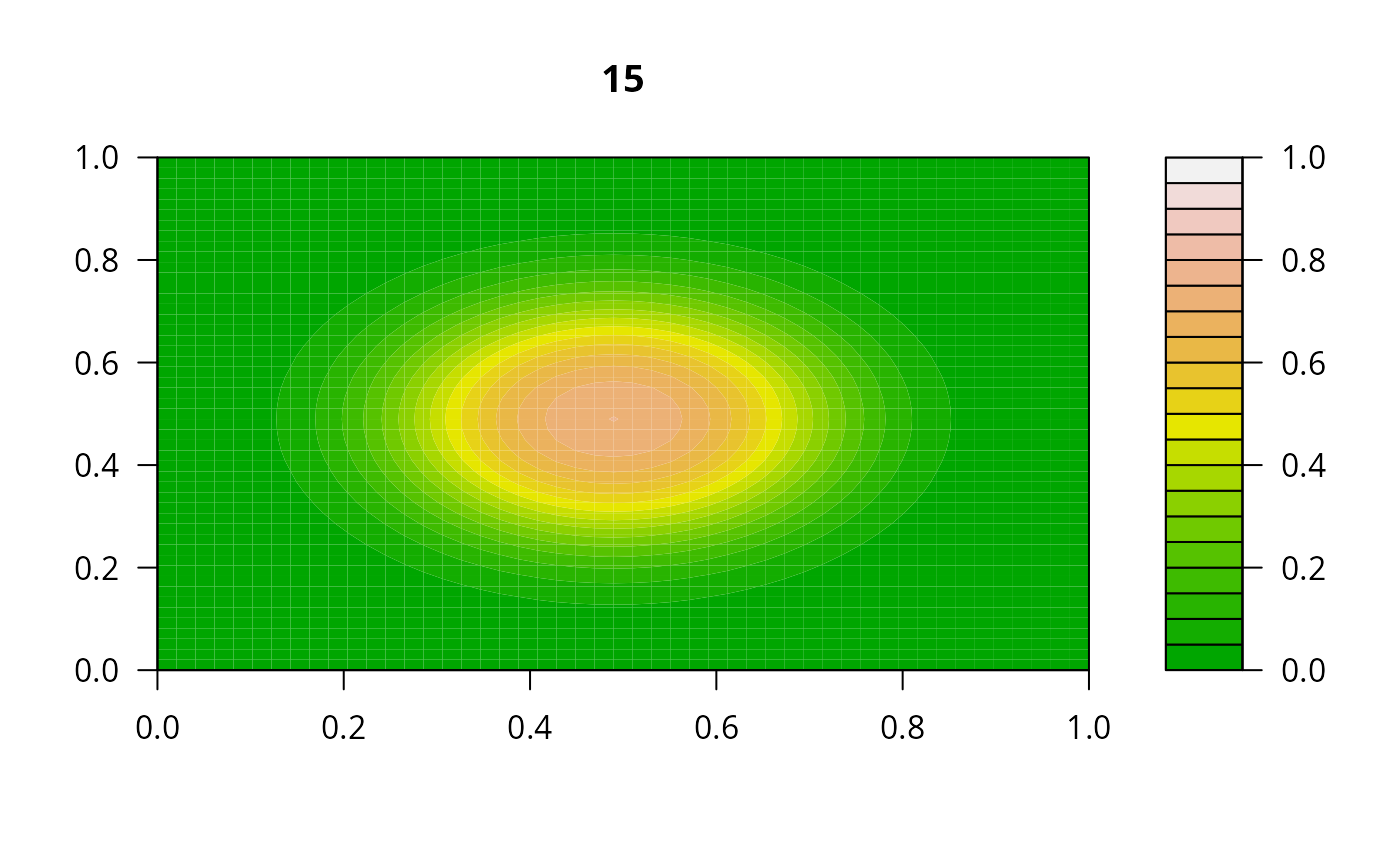
print(system.time(
for (i in 1:20) {
out <- lsode(func = diffusion2D, y = as.vector(y), times = times,
parms = NULL, jactype = "bandint", bandup = 1, banddown = 1)
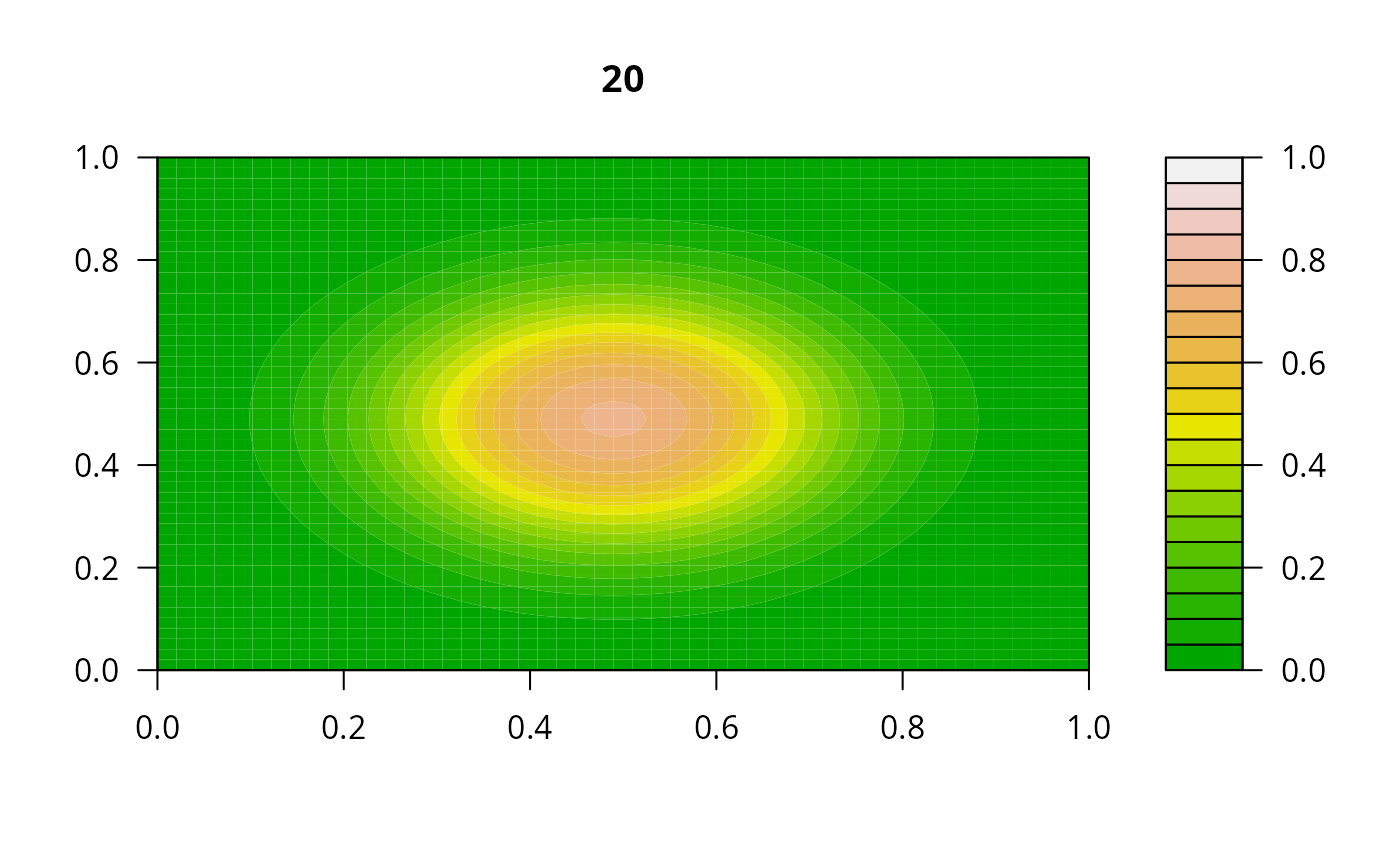
filled.contour(matrix(nrow = n, ncol = n, out[2,-1]), zlim = c(0,1),
color.palette = terrain.colors, main = i)
y <- out[2, -1]
}
))
## =======================================================================
## jacfunc need not be estimated exactly
## a crude approximation, with a smaller bandwidth will do.
## Here the half-bandwidth 1 is used, whereas the true
## half-bandwidths are equal to n.
## This corresponds to ignoring the y-direction coupling in the ODEs.
## =======================================================================
print(system.time(
for (i in 1:20) {
out <- lsode(func = diffusion2D, y = as.vector(y), times = times,
parms = NULL, jactype = "bandint", bandup = 1, banddown = 1)
filled.contour(matrix(nrow = n, ncol = n, out[2,-1]), zlim = c(0,1),
color.palette = terrain.colors, main = i)
y <- out[2, -1]
}
))
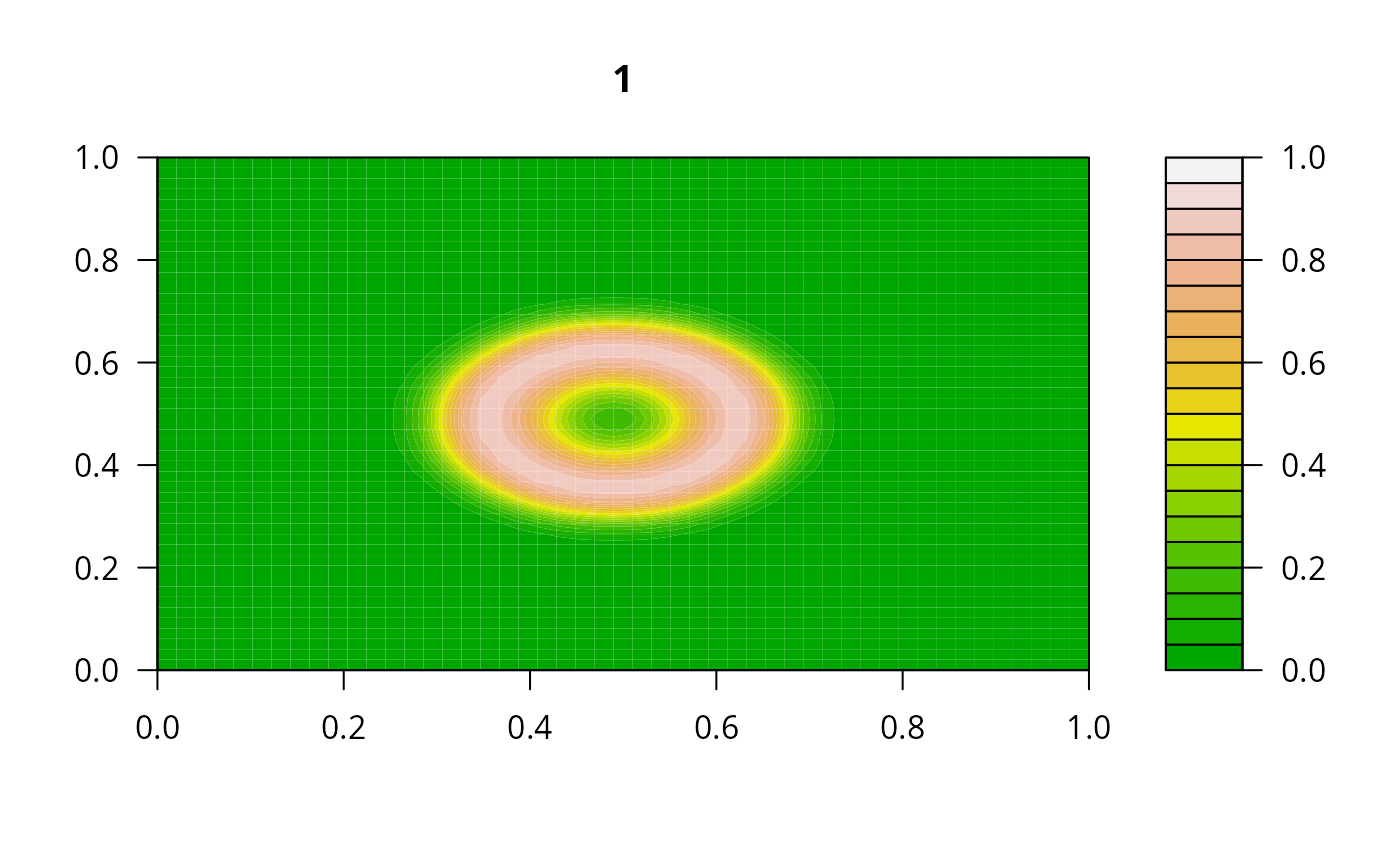
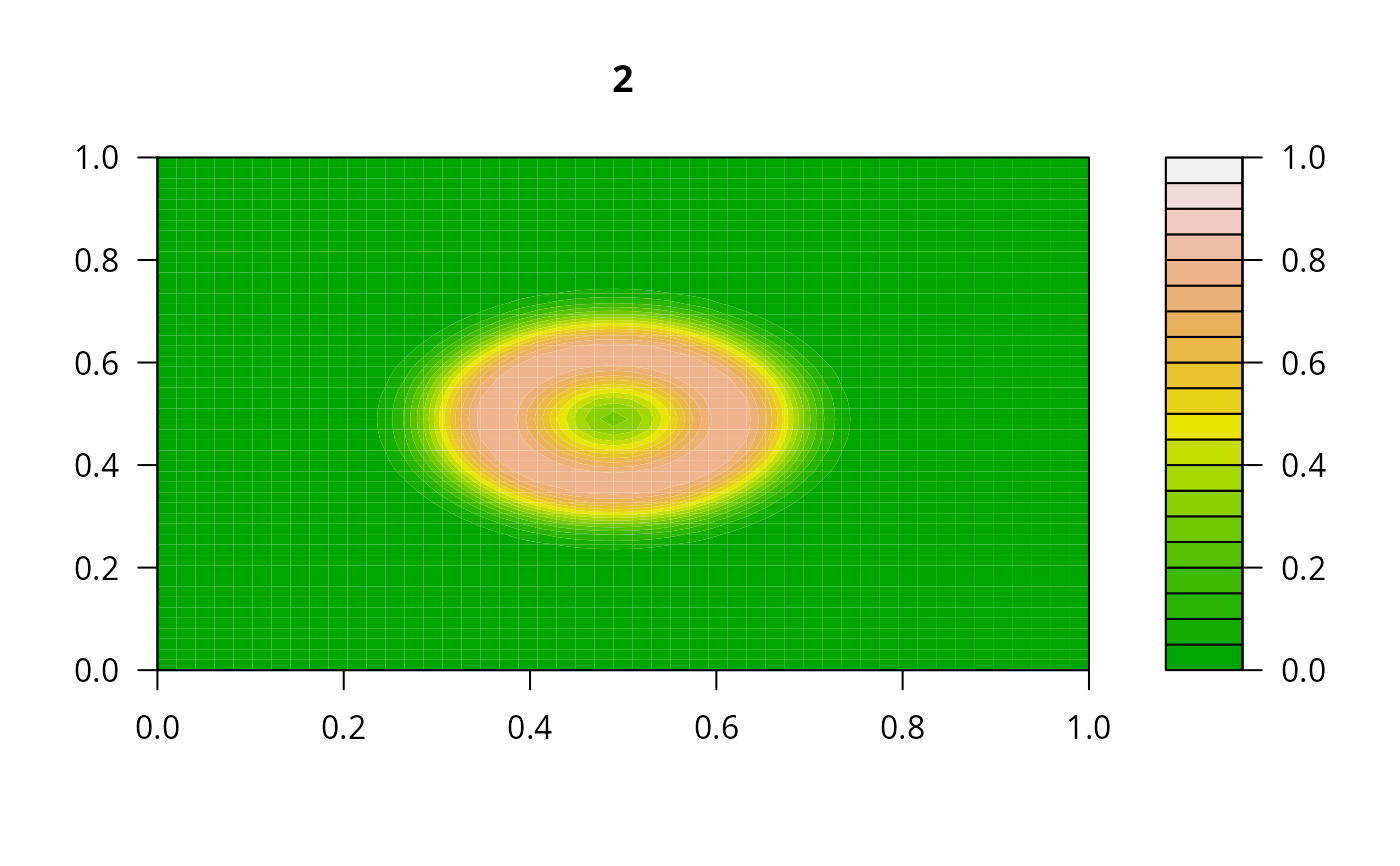
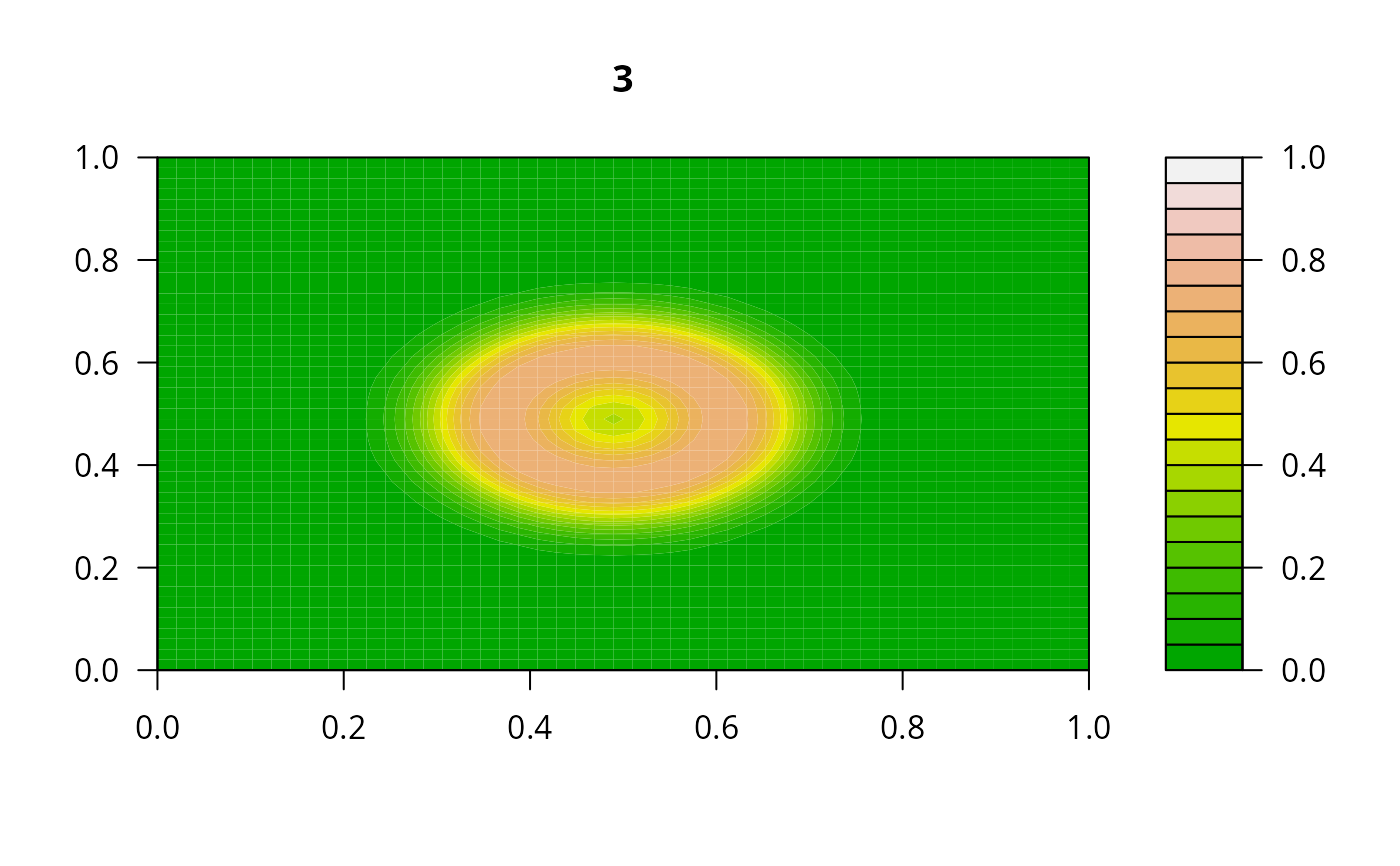
 #> user system elapsed
#> 0.252 0.009 0.261
par(ask = pa)
#> user system elapsed
#> 0.252 0.009 0.261
par(ask = pa)