General Solver for Delay Differential Equations.
dede.RdFunction dede is a general solver for delay differential equations, i.e.
equations where the derivative depends on past values of the state variables
or their derivatives.
Usage
dede(y, times, func=NULL, parms,
method = c( "lsoda", "lsode", "lsodes", "lsodar", "vode",
"daspk", "bdf", "adams", "impAdams", "radau"), control = NULL, ...)Arguments
- y
the initial (state) values for the DE system, a vector. If
yhas a name attribute, the names will be used to label the output matrix.- times
time sequence for which output is wanted; the first value of
timesmust be the initial time.- func
an R-function that computes the values of the derivatives in the ODE system (the model definition) at time \(t\).
funcmust be defined as:func <- function(t, y, parms, ...).tis the current time point in the integration,yis the current estimate of the variables in the DE system. If the initial valuesyhas anamesattribute, the names will be available insidefunc.parmsis a vector or list of parameters;...(optional) are any other arguments passed to the function.The return value of
funcshould be a list, whose first element is a vector containing the derivatives ofywith respect totime, and whose next elements are global values that are required at each point intimes.The derivatives must be specified in the same order as the state variablesy.If method "daspk" is used, then
funccan beNULL, in which caseresshould be used.- parms
parameters passed to
func.- method
the integrator to use, either a string (
"lsoda","lsode","lsodes","lsodar","vode","daspk","bdf","adams","impAdams","radau") or a function that performs the integration. The default integrator used is lsoda.- control
a list that can supply (1) the size of the history array, as
control$mxhist; the default is 1e4 and (2) how to interpolate, ascontrol$interpol, where1is hermitian interpolation,2is variable order interpolation, using the Nordsieck history array. Only for the two Adams methods is the second option recommended.- ...
additional arguments passed to the integrator.
Value
A matrix of class deSolve with up to as many rows as elements in
times and as many
columns as elements in y plus the number of "global" values
returned in the second element of the return from func, plus an
additional column (the first) for the time value. There will be one
row for each element in times unless the integrator returns
with an unrecoverable error. If y has a names attribute, it
will be used to label the columns of the output value.
Details
Functions lagvalue and lagderiv are to be used with dede
as they provide access to past (lagged)
values of state variables and derivatives. The number of past values that
are to be stored in a history matrix, can be specified in control$mxhist.
The default value (if unspecified) is 1e4.
Cubic Hermite interpolation is used by default to obtain an accurate
interpolant at the requested lagged time. For methods adams, impAdams,
a more accurate interpolation method can be triggered by setting
control$interpol = 2.
dede does not deal explicitly with propagated derivative discontinuities,
but relies on the integrator to control the stepsize in the region of a
discontinuity.
dede does not include methods to deal with delays that are smaller than the
stepsize, although in some cases it may be possible to solve such models.
For these reasons, it can only solve rather simple delay differential equations.
When used together with integrator lsodar, or lsode, dde
can simultaneously locate a root, and trigger an event. See last example.
Examples
## =============================================================================
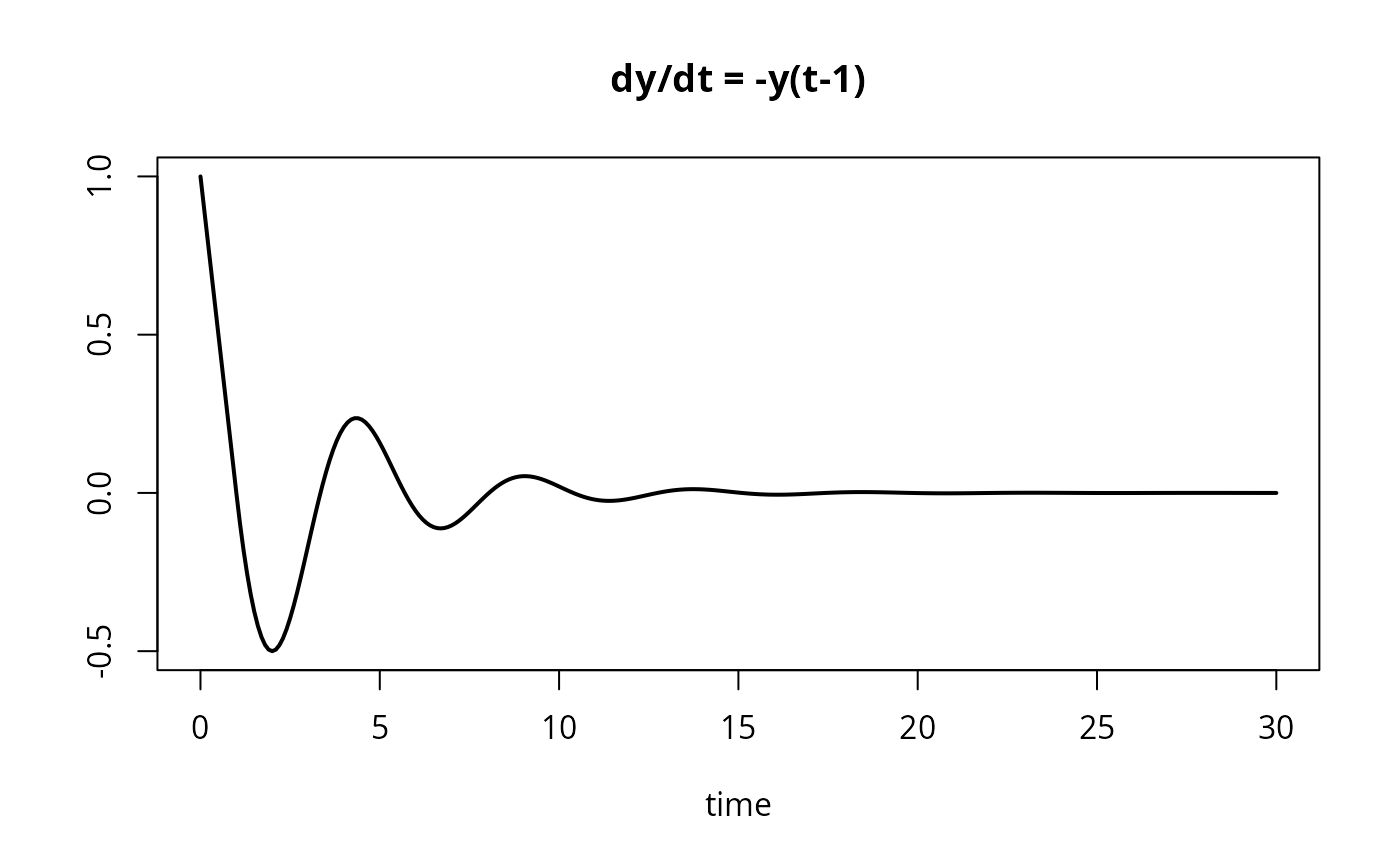
## A simple delay differential equation
## dy(t) = -y(t-1) ; y(t<0)=1
## =============================================================================
##-----------------------------
## the derivative function
##-----------------------------
derivs <- function(t, y, parms) {
if (t < 1)
dy <- -1
else
dy <- - lagvalue(t - 1)
list(c(dy))
}
##-----------------------------
## initial values and times
##-----------------------------
yinit <- 1
times <- seq(0, 30, 0.1)
##-----------------------------
## solve the model
##-----------------------------
yout <- dede(y = yinit, times = times, func = derivs, parms = NULL)
##-----------------------------
## display, plot results
##-----------------------------
plot(yout, type = "l", lwd = 2, main = "dy/dt = -y(t-1)")
 ## =============================================================================
## The infectuous disease model of Hairer; two lags.
## example 4 from Shampine and Thompson, 2000
## solving delay differential equations with dde23
## =============================================================================
##-----------------------------
## the derivative function
##-----------------------------
derivs <- function(t,y,parms) {
if (t < 1)
lag1 <- 0.1
else
lag1 <- lagvalue(t - 1,2)
if (t < 10)
lag10 <- 0.1
else
lag10 <- lagvalue(t - 10,2)
dy1 <- -y[1] * lag1 + lag10
dy2 <- y[1] * lag1 - y[2]
dy3 <- y[2] - lag10
list(c(dy1, dy2, dy3))
}
##-----------------------------
## initial values and times
##-----------------------------
yinit <- c(5, 0.1, 1)
times <- seq(0, 40, by = 0.1)
##-----------------------------
## solve the model
##-----------------------------
system.time(
yout <- dede(y = yinit, times = times, func = derivs, parms = NULL)
)
#> user system elapsed
#> 0.037 0.000 0.036
##-----------------------------
## display, plot results
##-----------------------------
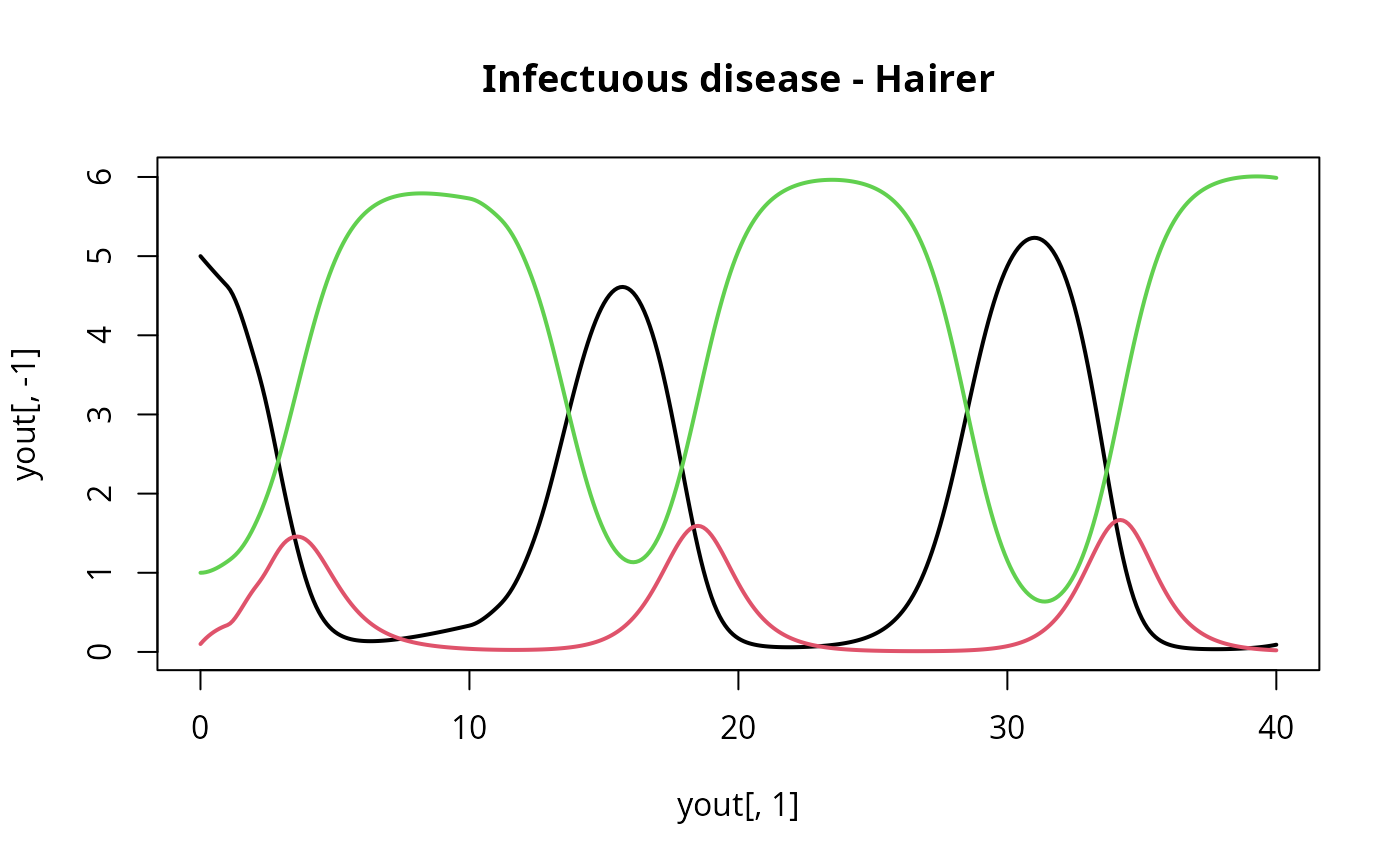
matplot(yout[,1], yout[,-1], type = "l", lwd = 2, lty = 1,
main = "Infectuous disease - Hairer")
## =============================================================================
## The infectuous disease model of Hairer; two lags.
## example 4 from Shampine and Thompson, 2000
## solving delay differential equations with dde23
## =============================================================================
##-----------------------------
## the derivative function
##-----------------------------
derivs <- function(t,y,parms) {
if (t < 1)
lag1 <- 0.1
else
lag1 <- lagvalue(t - 1,2)
if (t < 10)
lag10 <- 0.1
else
lag10 <- lagvalue(t - 10,2)
dy1 <- -y[1] * lag1 + lag10
dy2 <- y[1] * lag1 - y[2]
dy3 <- y[2] - lag10
list(c(dy1, dy2, dy3))
}
##-----------------------------
## initial values and times
##-----------------------------
yinit <- c(5, 0.1, 1)
times <- seq(0, 40, by = 0.1)
##-----------------------------
## solve the model
##-----------------------------
system.time(
yout <- dede(y = yinit, times = times, func = derivs, parms = NULL)
)
#> user system elapsed
#> 0.037 0.000 0.036
##-----------------------------
## display, plot results
##-----------------------------
matplot(yout[,1], yout[,-1], type = "l", lwd = 2, lty = 1,
main = "Infectuous disease - Hairer")
 ## =============================================================================
## time lags + EVENTS triggered by a root function
## The two-wheeled suitcase model
## example 8 from Shampine and Thompson, 2000
## solving delay differential equations with dde23
## =============================================================================
##-----------------------------
## the derivative function
##-----------------------------
derivs <- function(t, y, parms) {
if (t < tau)
lag <- 0
else
lag <- lagvalue(t - tau)
dy1 <- y[2]
dy2 <- -sign(y[1]) * gam * cos(y[1]) +
sin(y[1]) - bet * lag[1] + A * sin(omega * t + mu)
list(c(dy1, dy2))
}
## root and event function
root <- function(t,y,parms) ifelse(t>0, return(y), return(1))
event <- function(t,y,parms) return(c(y[1], y[2]*0.931))
gam = 0.248; bet = 1; tau = 0.1; A = 0.75
omega = 1.37; mu = asin(gam/A)
##-----------------------------
## initial values and times
##-----------------------------
yinit <- c(y = 0, dy = 0)
times <- seq(0, 12, len = 1000)
##-----------------------------
## solve the model
##-----------------------------
## Note: use a solver that supports both root finding and events,
## e.g. lsodar, lsode, lsoda, adams, bdf
yout <- dede(y = yinit, times = times, func = derivs, parms = NULL,
method = "lsodar", rootfun = root, events = list(func = event, root = TRUE))
##-----------------------------
## display, plot results
##-----------------------------
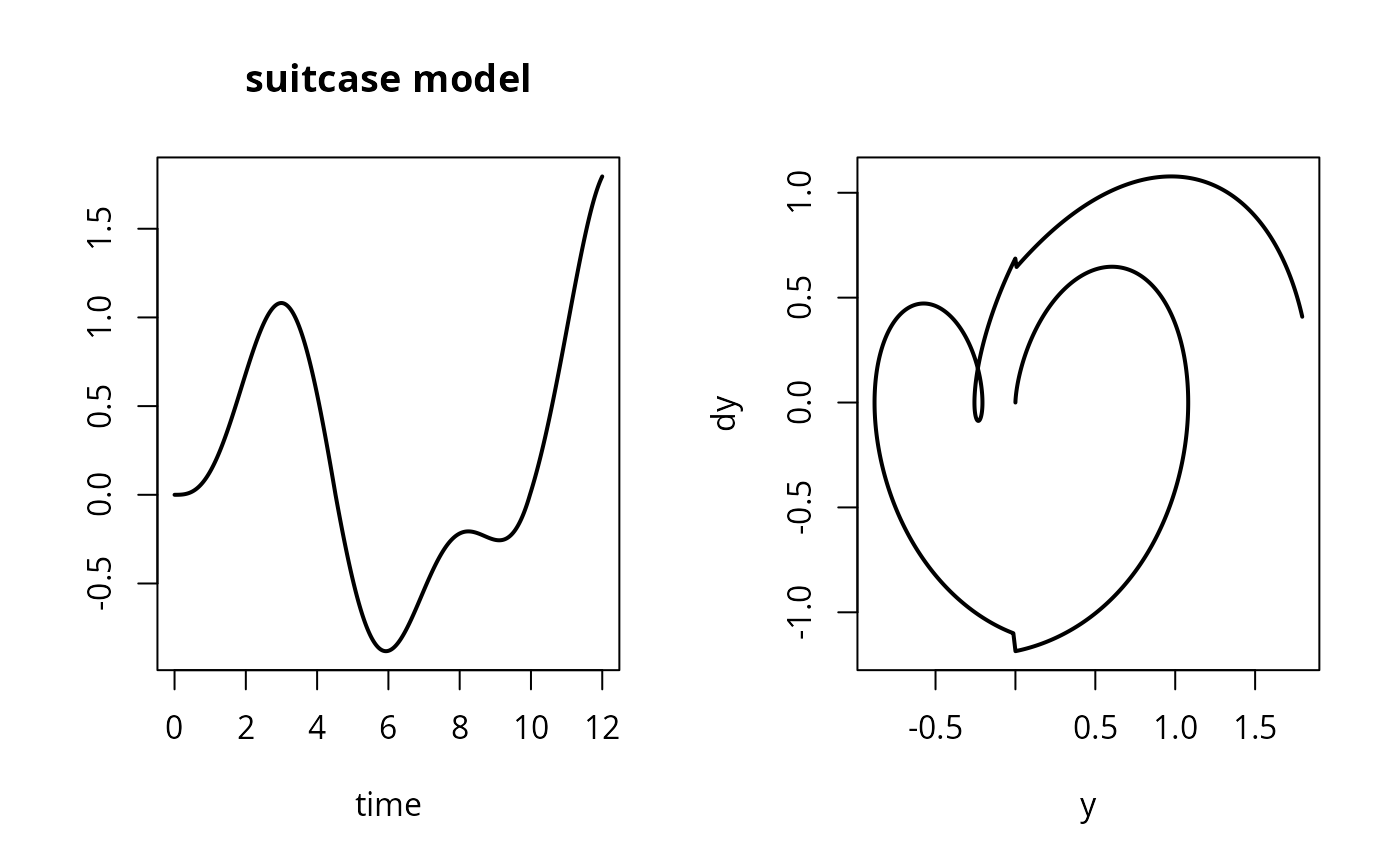
plot(yout, which = 1, type = "l", lwd = 2, main = "suitcase model", mfrow = c(1,2))
plot(yout[,2], yout[,3], xlab = "y", ylab = "dy", type = "l", lwd = 2)
## =============================================================================
## time lags + EVENTS triggered by a root function
## The two-wheeled suitcase model
## example 8 from Shampine and Thompson, 2000
## solving delay differential equations with dde23
## =============================================================================
##-----------------------------
## the derivative function
##-----------------------------
derivs <- function(t, y, parms) {
if (t < tau)
lag <- 0
else
lag <- lagvalue(t - tau)
dy1 <- y[2]
dy2 <- -sign(y[1]) * gam * cos(y[1]) +
sin(y[1]) - bet * lag[1] + A * sin(omega * t + mu)
list(c(dy1, dy2))
}
## root and event function
root <- function(t,y,parms) ifelse(t>0, return(y), return(1))
event <- function(t,y,parms) return(c(y[1], y[2]*0.931))
gam = 0.248; bet = 1; tau = 0.1; A = 0.75
omega = 1.37; mu = asin(gam/A)
##-----------------------------
## initial values and times
##-----------------------------
yinit <- c(y = 0, dy = 0)
times <- seq(0, 12, len = 1000)
##-----------------------------
## solve the model
##-----------------------------
## Note: use a solver that supports both root finding and events,
## e.g. lsodar, lsode, lsoda, adams, bdf
yout <- dede(y = yinit, times = times, func = derivs, parms = NULL,
method = "lsodar", rootfun = root, events = list(func = event, root = TRUE))
##-----------------------------
## display, plot results
##-----------------------------
plot(yout, which = 1, type = "l", lwd = 2, main = "suitcase model", mfrow = c(1,2))
plot(yout[,2], yout[,3], xlab = "y", ylab = "dy", type = "l", lwd = 2)