Plot Estimated Systematical Bias with Confidence Bounds
MCResult.plotBias.RdThis function plots the estimated systematical bias \(( Intercept + Slope * Refrencemethod ) - Referencemethod\) with confidence bounds, covering the whole range of reference method X or only part of it.
MCResult.plotBias(
x,
xn = 100,
alpha = 0.05,
add = FALSE,
prop = FALSE,
xlim = NULL,
ylim = NULL,
bias = TRUE,
bias.lty = 1,
bias.lwd = 2,
bias.col = NULL,
ci.area = TRUE,
ci.area.col = NULL,
ci.border = FALSE,
ci.border.col = NULL,
ci.border.lwd = 1,
ci.border.lty = 2,
zeroline = TRUE,
zeroline.col = NULL,
zeroline.lty = 2,
zeroline.lwd = 1,
main = NULL,
sub = NULL,
add.grid = TRUE,
xlab = NULL,
ylab = NULL,
cut.point = NULL,
cut.point.col = "red",
cut.point.lwd = 2,
cut.point.lty = 1,
...
)Arguments
- x
object of class "MCResult".
- xn
# number of poits for drawing of confidence bounds/area.
- alpha
numeric value specifying the 100(1-
alpha)% confidence level of confidence intervals (Default is 0.05).- add
logical value. If
add=TRUE, the grafic will be drawn in current grafical window.- prop
a logical value. If
prop=TRUEthe proportional bias \( \%bias(Xc) = [ Intercept + (Slope-1) * Xc ] / Xc\) will be drawn.- xlim
limits of the x-axis. If
xlim=NULLthe x-limits will be calculated automatically.- ylim
limits of the y-axis. If
ylim=NULLthe y-limits will be calculated automatically.- bias
logical value. If
identity=TRUEthe bias line will be drawn. If ci.bounds=FALSE and ci.area=FALSE the bias line will be drawn always.- bias.lty
type of the bias line.
- bias.lwd
width of the bias line.
- bias.col
color of the bias line.
- ci.area
logical value. If
ci.area=TRUE(default) the confidence area will be drawn.- ci.area.col
color of the confidence area.
- ci.border
logical value. If ci.border=TRUE the confidence limits will be drawn.
- ci.border.col
color of the confidence limits.
- ci.border.lwd
line width of confidence limits.
- ci.border.lty
line type of confidence limits.
- zeroline
logical value. If
zeroline=TRUEthe zero-line will be drawn.- zeroline.col
color of the zero-line.
- zeroline.lty
type of the zero-line.
- zeroline.lwd
width of the zero-line.
- main
character string. The main title of plot. If
main = NULLit will include regression name.- sub
character string. The subtitle of plot. If
sub=NULLandci.border=TRUEorci.area=TRUEit will include the art of confidence bounds calculation.- add.grid
logical value. If
grid=TRUE(default) the gridlines will be drawn.- xlab
label for the x-axis
- ylab
label for the y-axis
- cut.point
numeric value. Decision level of interest.
- cut.point.col
color of the confidence bounds at the required decision level.
- cut.point.lwd
line width of the confidence bounds at the required decision level.
- cut.point.lty
line type of the confidence bounds at the required decision level.
- ...
further graphical parameters
Value
No return value, instead a plot is generated
See also
Examples
#library("mcr")
data(creatinine,package="mcr")
creatinine <- creatinine[complete.cases(creatinine),]
x <- creatinine$serum.crea
y <- creatinine$plasma.crea
# Calculation of models
m1 <- mcreg(x,y,method.reg="WDeming", method.ci="jackknife",
mref.name="serum.crea",mtest.name="plasma.crea", na.rm=TRUE)
#> Jackknife based calculation of standard error and confidence intervals according to Linnet's method.
m2 <- mcreg(x,y,method.reg="WDeming", method.ci="bootstrap",
method.bootstrap.ci="BCa",mref.name="serum.crea",
mtest.name="plasma.crea", na.rm=TRUE)
#> The global.sigma is calculated with Linnet's method
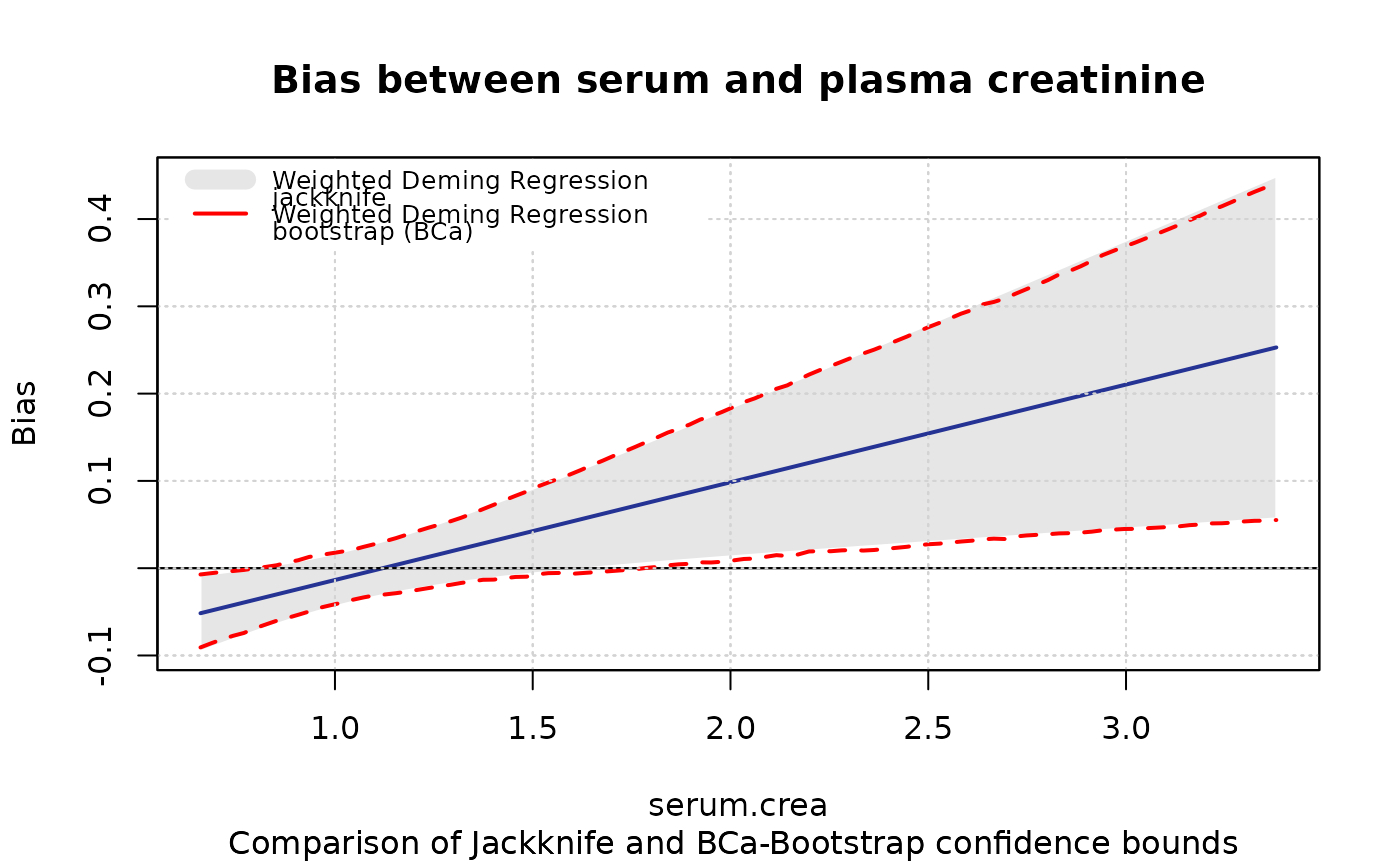
# Grafical comparison of systematical Bias of two models
plotBias(m1, zeroline=TRUE,zeroline.col="black",zeroline.lty=1,
ci.area=TRUE,ci.border=FALSE, ci.area.col=grey(0.9),
main = "Bias between serum and plasma creatinine",
sub="Comparison of Jackknife and BCa-Bootstrap confidence bounds ")
plotBias(m2, ci.area=FALSE, ci.border=TRUE, ci.border.lwd=2,
ci.border.col="red",bias=FALSE ,add=TRUE)
includeLegend(place="topleft",models=list(m1,m2), lwd=c(10,2),
lty=c(2,1),colors=c(grey(0.9),"red"), bias=TRUE,
design="1", digits=4)
 # Drawing of proportional bias
plotBias(m1, ci.area=FALSE, ci.border=TRUE)
# Drawing of proportional bias
plotBias(m1, ci.area=FALSE, ci.border=TRUE)
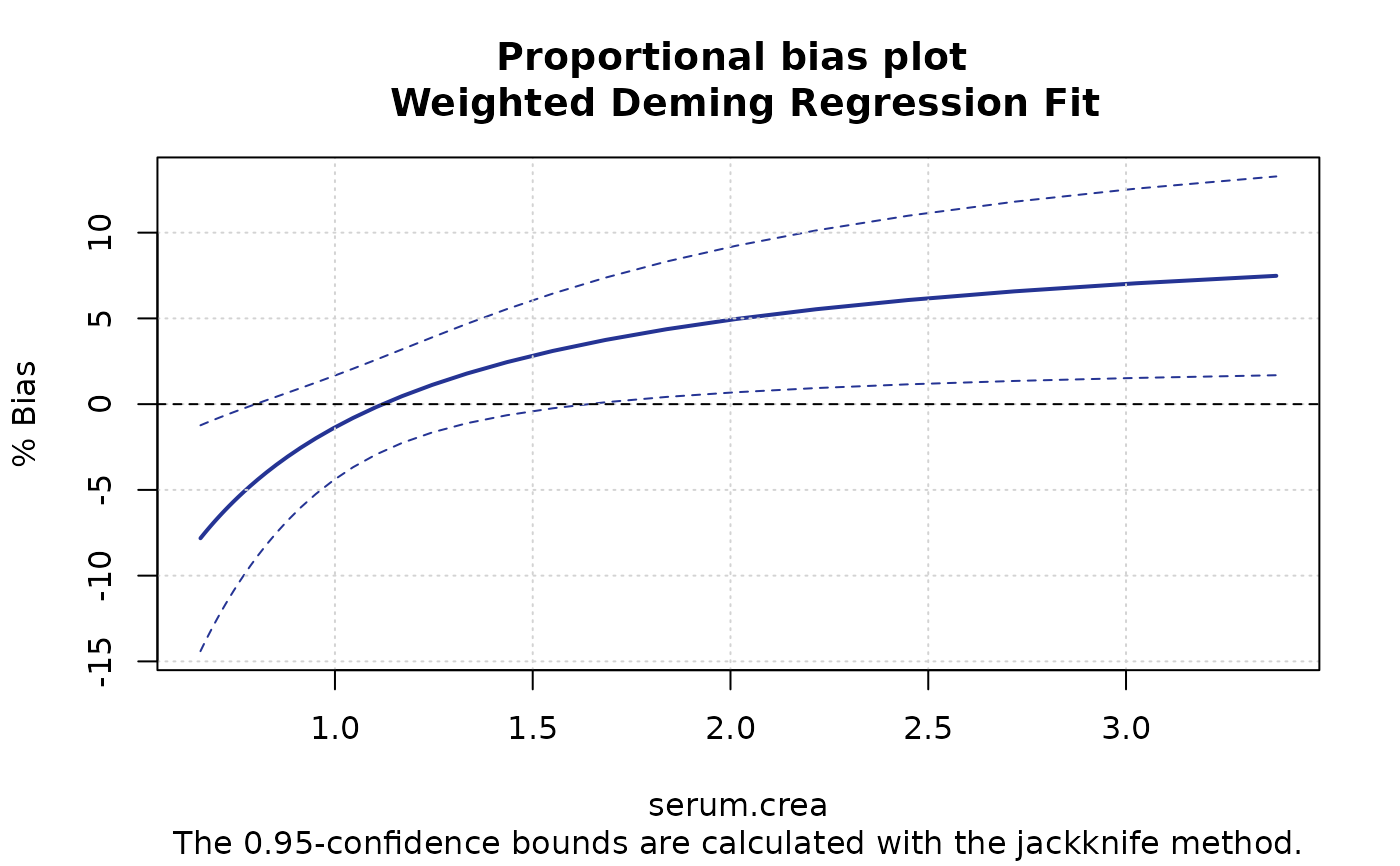
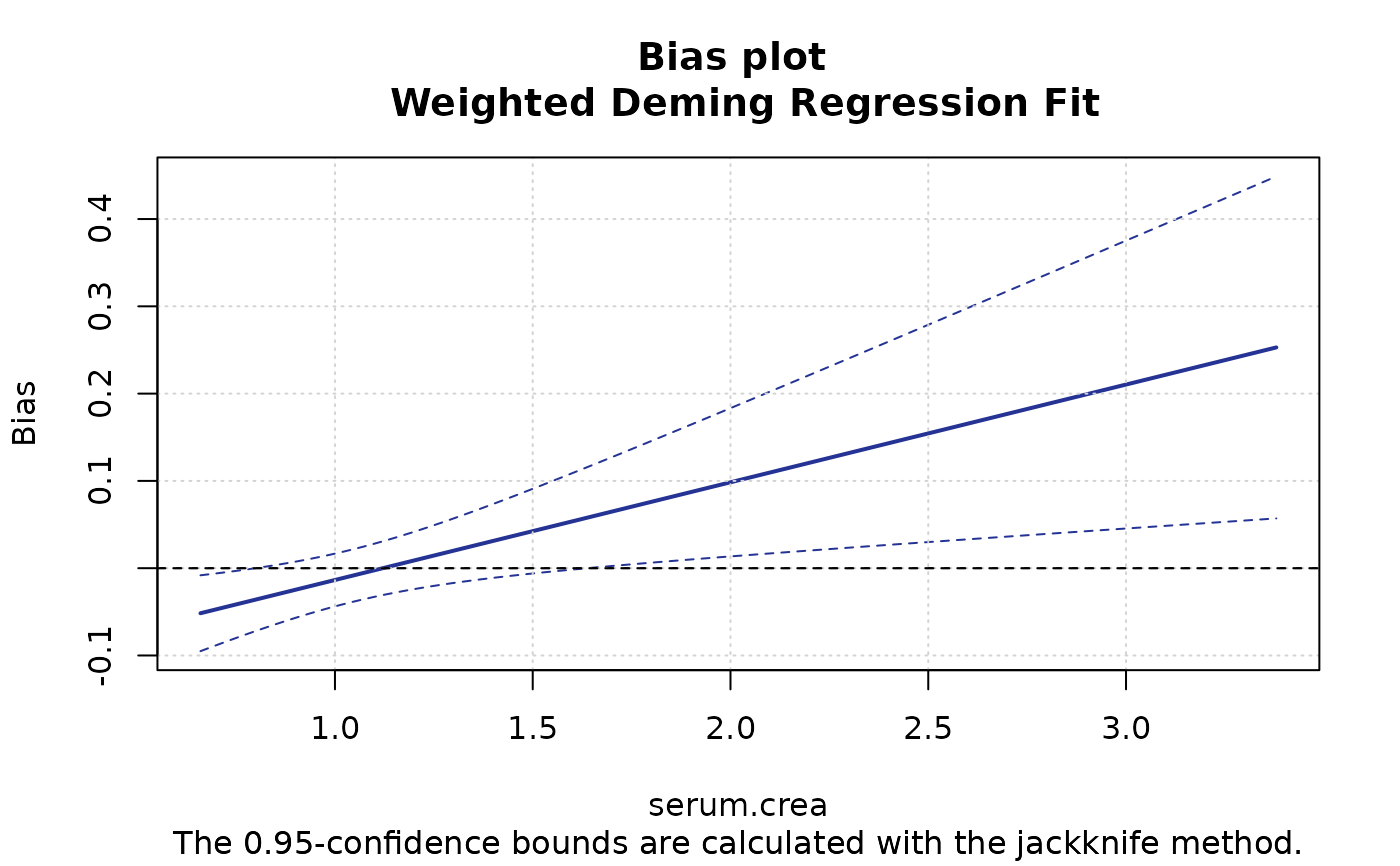 plotBias(m1, ci.area=FALSE, ci.border=TRUE, prop=TRUE)
plotBias(m1, ci.area=FALSE, ci.border=TRUE, prop=TRUE)
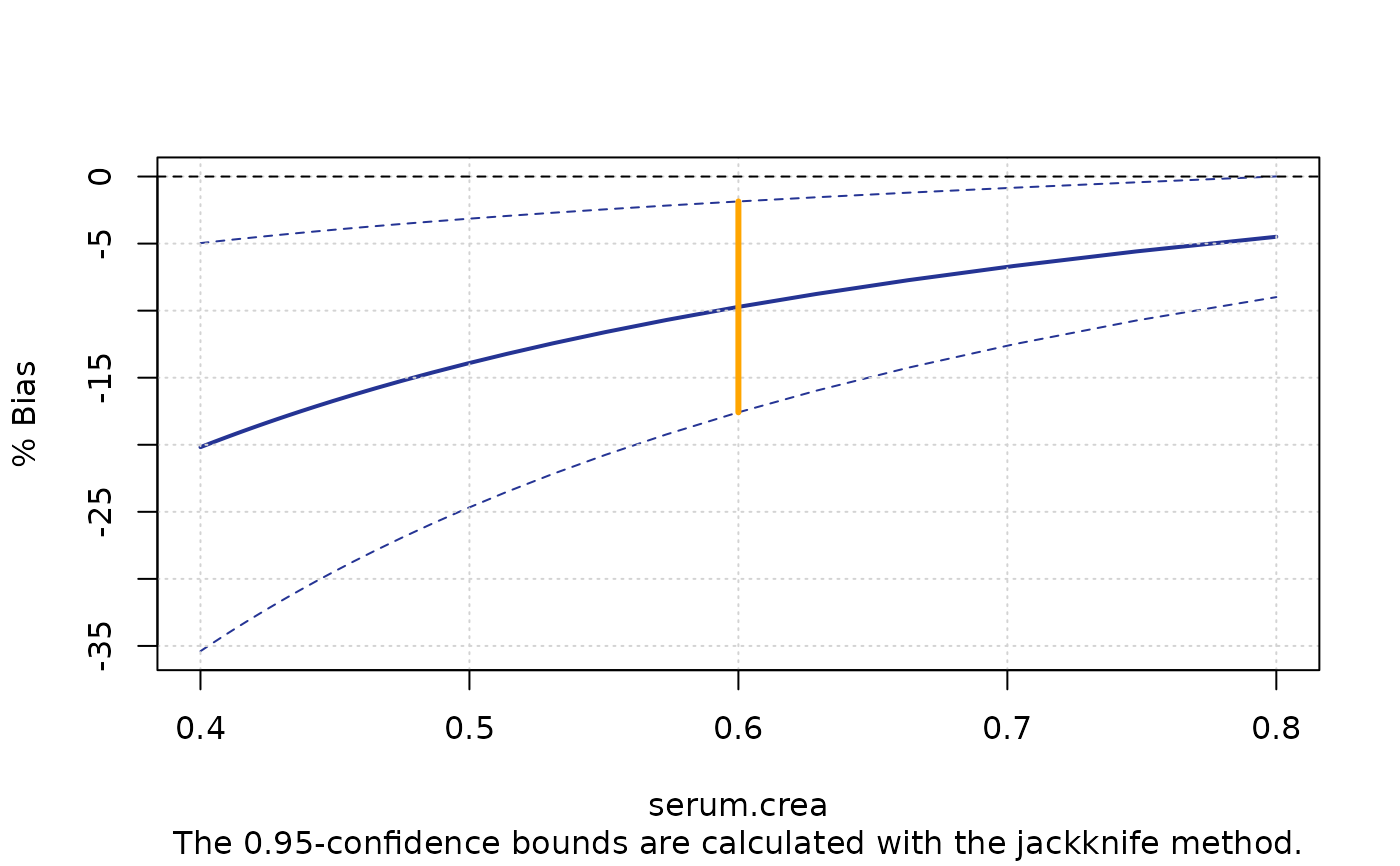
 plotBias(m1, ci.area=FALSE, ci.border=TRUE, prop=TRUE, cut.point=0.6)
plotBias(m1, ci.area=FALSE, ci.border=TRUE, prop=TRUE, cut.point=0.6)
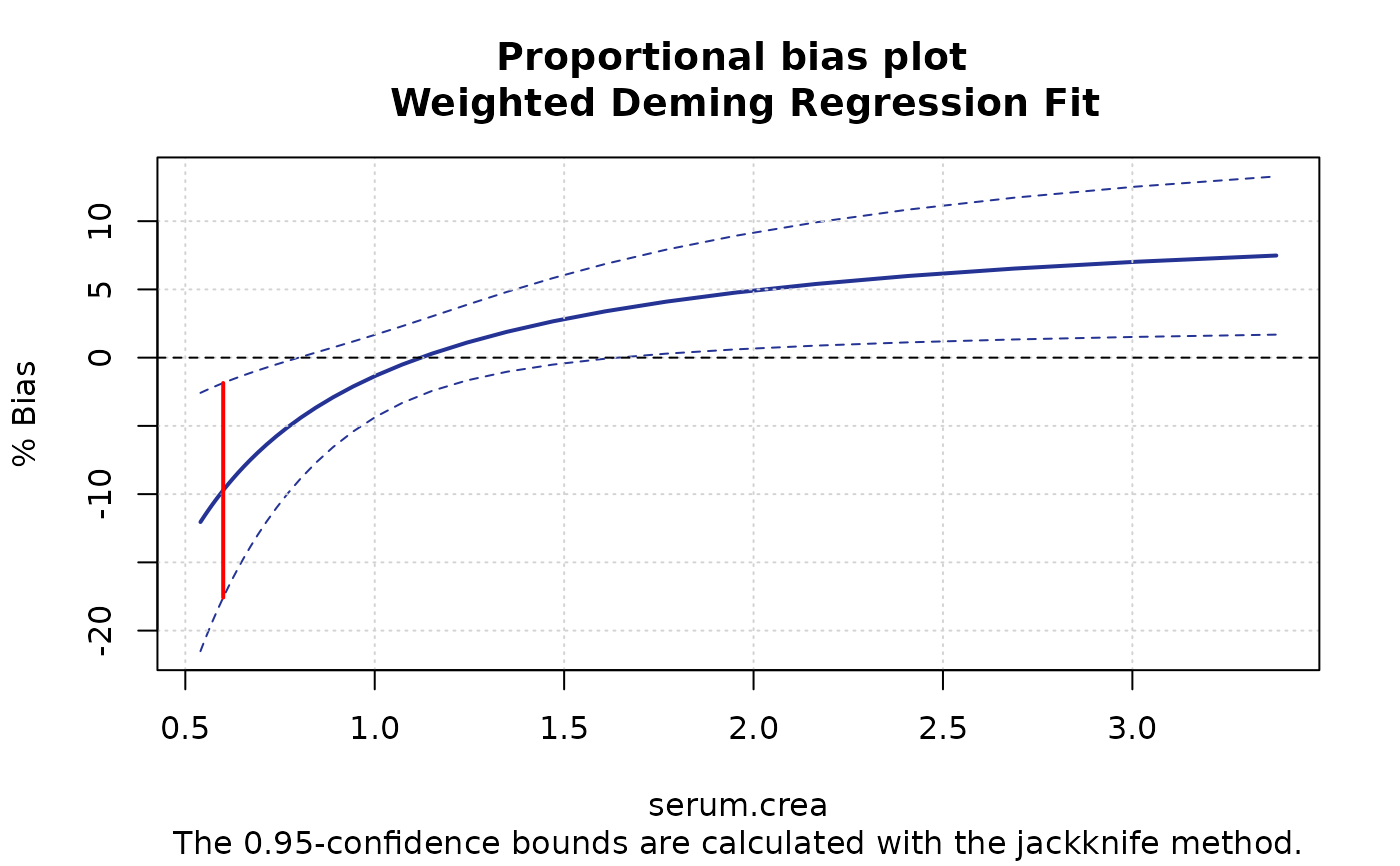 plotBias(m1, ci.area=FALSE, ci.border=TRUE, prop=TRUE, cut.point=0.6,
xlim=c(0.4,0.8),cut.point.col="orange", cut.point.lwd=3, main ="")
plotBias(m1, ci.area=FALSE, ci.border=TRUE, prop=TRUE, cut.point=0.6,
xlim=c(0.4,0.8),cut.point.col="orange", cut.point.lwd=3, main ="")