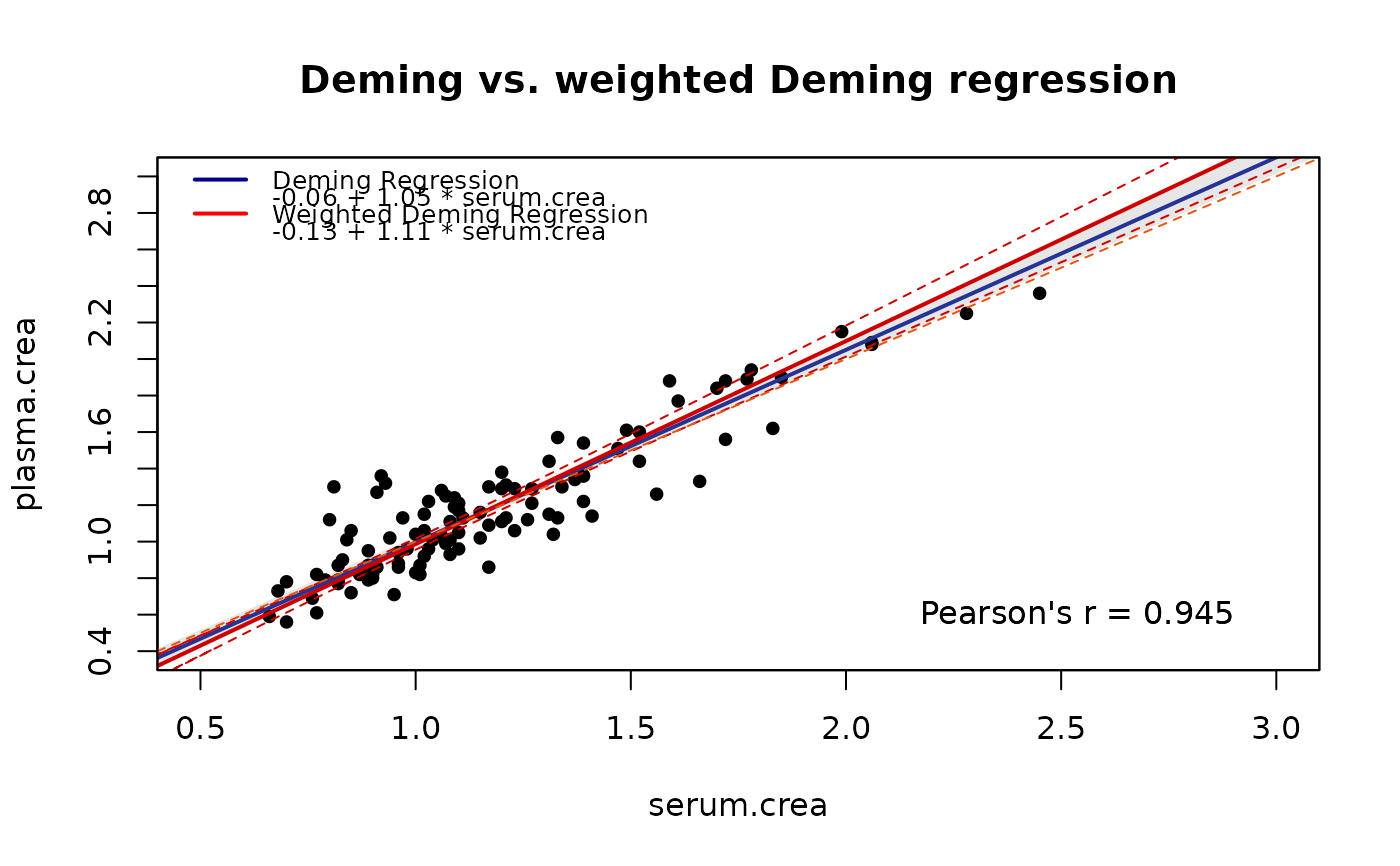
Scatter Plot Method X vs. Method Y
MCResult.plot.RdPlot method X (reference) vs. method Y (test) with (optional) line of identity, regression line and confidence bounds for response.
MCResult.plot(
x,
alpha = 0.05,
xn = 20,
equal.axis = FALSE,
xlim = NULL,
ylim = NULL,
xaxp = NULL,
yaxp = NULL,
x.lab = x@mnames[1],
y.lab = x@mnames[2],
add = FALSE,
draw.points = TRUE,
points.col = "black",
points.pch = 1,
points.cex = 0.8,
reg = TRUE,
reg.col = NULL,
reg.lty = 1,
reg.lwd = 2,
identity = TRUE,
identity.col = NULL,
identity.lty = 2,
identity.lwd = 1,
ci.area = TRUE,
ci.area.col = NULL,
ci.border = FALSE,
ci.border.col = NULL,
ci.border.lty = 2,
ci.border.lwd = 1,
add.legend = TRUE,
legend.place = c("topleft", "topright", "bottomleft", "bottomright"),
main = NULL,
sub = NULL,
add.cor = TRUE,
cor.method = c("pearson", "kendall", "spearman"),
add.grid = TRUE,
digits = list(coef = 2, cor = 3),
...
)Arguments
- x
object of class "MCResult".
- alpha
numeric value specifying the 100(1-
alpha)% confidence bounds.- xn
number of points (default 20) for calculation of confidence bounds.
- equal.axis
logical value. If
equal.axis=TRUEx-axis will be equal to y-axis.- xlim
limits of the x-axis. If
xlim=NULLthe x-limits will be calculated automatically.- ylim
limits of the y-axis. If
ylim=NULLthe y-limits will be calculated automatically.- xaxp
ticks of the x-axis. If
xaxp=NULLthe x-ticks will be calculated automatically.- yaxp
ticks of the y-axis. If
yaxp=NULLthe y-ticks will be calculated automatically.- x.lab
label of x-axis. Default is the name of reference method.
- y.lab
label of y-axis. Default is the name of test method.
- add
logical value. If
add=TRUE, the plot will be drawn in current graphical window.- draw.points
logical value. If
draw.points=TRUE, the data points will be drawn.- points.col
Color of data points.
- points.pch
Type of data points (see
par()).- points.cex
Size of data points (see
par()).- reg
Logical value. If
reg=TRUE, the regression line will be drawn.- reg.col
Color of regression line.
- reg.lty
Type of regression line.
- reg.lwd
The width of regression line.
- identity
logical value. If
identity=TRUEthe identity line will be drawn.- identity.col
The color of identity line.
- identity.lty
The type of identity line.
- identity.lwd
the width of identity line.
- ci.area
logical value. If
ci.area=TRUE(default) the confidence area will be drawn.- ci.area.col
the color of confidence area.
- ci.border
logical value. If
ci.border=TRUEthe confidence limits will be drawn.- ci.border.col
The color of confidence limits.
- ci.border.lty
The line type of confidence limits.
- ci.border.lwd
The line width of confidence limits.
- add.legend
logical value. If
add.legend=FALSEthe plot will not have any legend.- legend.place
The position of legend: "topleft","topright","bottomleft","bottomright".
- main
String value. The main title of plot. If
main=NULLit will include regression name.- sub
String value. The subtitle of plot. If
sub=NULLandci.border=TRUEorci.area=TRUEit will include the art of confidence bounds calculation.- add.cor
Logical value. If
add.cor=TRUEthe correlation coefficient will be shown.- cor.method
a character string indicating which correlation coefficient is to be computed. One of "pearson" (default), "kendall", or "spearman", can be abbreviated.
- add.grid
Logical value. If
add.grid=TRUE(default) the gridlines will be drawn.- digits
list with the number of digits for the regression equation and the correlation coefficient.
- ...
further graphical parameters
Value
No return value, instead a plot is generated
See also
Examples
library(mcr)
data(creatinine,package="mcr")
creatinine <- creatinine[complete.cases(creatinine),]
x <- creatinine$serum.crea
y <- creatinine$plasma.crea
m1 <- mcreg(x,y,method.reg="Deming", mref.name="serum.crea",
mtest.name="plasma.crea", na.rm=TRUE)
m2 <- mcreg(x,y,method.reg="WDeming", method.ci="jackknife",
mref.name="serum.crea",
mtest.name="plasma.crea", na.rm=TRUE)
#> Jackknife based calculation of standard error and confidence intervals according to Linnet's method.
plot(m1, xlim=c(0.5,3),ylim=c(0.5,3), add.legend=FALSE,
main="Deming vs. weighted Deming regression",
points.pch=19,ci.area=TRUE, ci.area.col=grey(0.9),
identity=FALSE, add.grid=FALSE, sub="")
plot(m2, ci.area=FALSE, ci.border=TRUE, ci.border.col="red3",
reg.col="red3", add.legend=FALSE,
draw.points=FALSE,add=TRUE)
includeLegend(place="topleft",models=list(m1,m2),
colors=c("darkblue","red"), design="1", digits=2)