Resampling Model Calibration
calibrate.RdUses bootstrapping or cross-validation to get bias-corrected (overfitting-
corrected) estimates of predicted vs. observed values based on
subsetting predictions into intervals or better on
nonparametric or adaptive parametric smoothers. There are calibration
functions for Cox (cph), parametric survival models (psm),
binary and ordinal logistic models (lrm, orm) and ordinary least
squares (ols). For survival models and orm,
"predicted" means predicted survival probability at a single
time point, and "observed" refers to the corresponding Kaplan-Meier
survival estimate, stratifying on intervals of predicted survival, or,
the predicted survival
probability as a function of transformed predicted survival probability
using the flexible hazard regression approach or for orm and probably
better, smoothed overlapping moving Kaplan-Meier estimates
(see the val.surv.args argument and val.surv
function for details). Nonparametric
calibration curves are estimated over a regular sequence of predicted values. The
fit must have specified x=TRUE, y=TRUE.
See predab.resample for information about confidence limits. Confidence limits for bootstrap overfitting-corrected calibration curves are not computed for psm fits. This is because of calibrate.psm averages over multiple bootstrap loops. This can probably be changed.
The print and
plot methods print the mean absolute error in predictions,
the mean squared error, and the 0.9 quantile of the absolute error.
Here, error refers to the difference between the predicted values and
the corresponding bias-corrected calibrated values.
Below, calibrate.default is for the ols and lrm.
Usage
calibrate(fit, ...)
# Default S3 method
calibrate(fit, predy,
method=c("boot","crossvalidation",".632","randomization"),
B=40, bw=FALSE, rule=c("aic","p"),
type=c("residual","individual"),
sls=.05, aics=0, force=NULL, estimates=TRUE, pr=FALSE, kint,
smoother="lowess", digits=NULL, ...)
# S3 method for class 'cph'
calibrate(fit, cmethod=c('hare', 'KM'),
method="boot", u, m=150, pred, cuts, B=40,
bw=FALSE, rule="aic", type="residual", sls=0.05, aics=0, force=NULL,
estimates=TRUE,
pr=FALSE, what="observed-predicted", tol=1e-12, maxdim=5, ...)
# S3 method for class 'orm'
calibrate(fit,
method="boot", u, m=150, pred, B=40,
bw=FALSE, rule="aic",
type="residual", sls=.05, aics=0, force=NULL,
estimates=TRUE, pr=FALSE, what="observed-predicted",
val.surv.args=list(method='smoothkm', eps=30),
...)
# S3 method for class 'psm'
calibrate(fit, cmethod=c('hare', 'KM'),
method="boot", u, m=150, pred, cuts, B=40,
bw=FALSE,rule="aic",
type="residual", sls=.05, aics=0, force=NULL, estimates=TRUE,
pr=FALSE, what="observed-predicted", tol=1e-12, maxiter=15,
rel.tolerance=1e-5, maxdim=5, ...)
# S3 method for class 'calibrate'
print(x, B=Inf, ...)
# S3 method for class 'calibrate.default'
print(x, B=Inf, ...)
# S3 method for class 'calibrate'
plot(x, xlab, ylab, subtitles=TRUE, conf.int=TRUE,
cex.subtitles=.75, riskdist=TRUE, add=FALSE,
scat1d.opts=list(nhistSpike=200), par.corrected=NULL, ...)
# S3 method for class 'calibrate.default'
plot(x, xlab, ylab, xlim, ylim,
legend=TRUE, subtitles=TRUE, cex.subtitles=.75, riskdist=TRUE,
scat1d.opts=list(nhistSpike=200), ...)Arguments
- fit
a fit from
ols,lrm,cphorpsm- x
an object created by
calibrate- method, B, bw, rule, type, sls, aics, force, estimates
see
validate. Forprint.calibrate,Bis an upper limit on the number of resamples for which information is printed about which variables were selected in each model re-fit. Specify zero to suppress printing. Default is to print all re-samples.- cmethod
method for validating survival predictions using right-censored data. The default is
cmethod='hare'to use theharefunction in thepolsplinepackage. Specifycmethod='KM'to use less precision stratified Kaplan-Meier estimates. If thepolsplinepackage is not available, the procedure reverts tocmethod='KM'.- u
the time point for which to validate predictions for survival models. For
cphfits, you must have specifiedsurv=TRUE, time.inc=u, whereuis the constant specifying the time to predict.- m
group predicted
u-time units survival into intervals containingmsubjects on the average (for survival models only)- pred
vector of predicted survival probabilities at which to evaluate the calibration curve. By default, the low and high prediction values from
datadistare used, which for large sample size is the 10th smallest to the 10th largest predicted probability.- cuts
actual cut points for predicted survival probabilities. You may specify only one of
mandcuts(for survival models only)- pr
set to
TRUEto print intermediate results for each re-sample- what
The default is
"observed-predicted", meaning to estimate optimism in this difference. This is preferred as it accounts for skewed distributions of predicted probabilities in outer intervals. You can also specify"observed". This argument applies to survival models only.- tol
criterion for matrix singularity (default is
1e-12)- maxdim
see
hare- maxiter
for
psm, this is passed tosurvreg.control(default is 15 iterations)- rel.tolerance
parameter passed to
survreg.controlforpsm(default is 1e-5).- predy
a scalar or vector of predicted values to calibrate (for
lrm,ols). Default is 50 equally spaced points between the 5th smallest and the 5th largest predicted values. Forlrmthe predicted values are probabilities (seekint).- kint
For an ordinal logistic model the default predicted probability that \(Y\geq\) the middle level. Specify
kintto specify the intercept to use, e.g.,kint=2means to calibrate \(Prob(Y\geq b)\), where \(b\) is the second level of \(Y\).- val.surv.args
a list containing arguments to send to
val.survwhen runningcalibrate.orm. By default smoothed overlapping windows of Kaplan-Meier estimates are used fororm. Theval.surv.argsargument is especially useful for specifying bandwidths and themovStatsepsargument.- smoother
a function in two variables which produces \(x\)- and \(y\)-coordinates by smoothing the input
y. The default is to uselowess(x, y, iter=0).- digits
If specified, predicted values are rounded to
digitsdigits before passing to the smoother. Occasionally, large predicted values on the logit scale will lead to predicted probabilities very near 1 that should be treated as 1, and theroundfunction will fix that. Applies tocalibrate.default.- ...
other arguments to pass to
predab.resample, such asconf.int,group,cluster, andsubset. Also, other arguments forplot.- xlab
defaults to "Predicted x-units Survival" or to a suitable label for other models
- ylab
defaults to "Fraction Surviving x-units" or to a suitable label for other models
- xlim,ylim
2-vectors specifying x- and y-axis limits, if not using defaults
- subtitles
set to
FALSEto suppress subtitles in plot describing method and forlrmandolsthe mean absolute error and original sample size- conf.int
set to
FALSEto suppress plotting 0.95 confidence intervals for Kaplan-Meier estimates- cex.subtitles
character size for plotting subtitles
- riskdist
set to
FALSEto suppress the distribution of predicted risks (survival probabilities) from being plotted- add
set to
TRUEto add the calibration plot to an existing plot- scat1d.opts
a list specifying options to send to
scat1difriskdist=TRUE. Seescat1d.- par.corrected
a list specifying graphics parameters
col,lty,lwd,pchto be used in drawing overfitting-corrected estimates. Default iscol="blue",lty=1,lwd=1,pch=4.- legend
set to
FALSEto suppress legends (forlrm,olsonly) on the calibration plot, or specify a list with elementsxandycontaining the coordinates of the upper left corner of the legend. By default, a legend will be drawn in the lower right 1/16th of the plot.
Value
matrix specifying mean predicted survival in each interval, the
corresponding estimated bias-corrected Kaplan-Meier estimates,
number of subjects, and other statistics. For linear and logistic models,
the matrix instead has rows corresponding to the prediction points, and
the vector of predicted values being validated is returned as an attribute.
The returned object has class "calibrate" or
"calibrate.default".
plot.calibrate.default invisibly returns the vector of estimated
prediction errors corresponding to the dataset used to fit the model.
Details
If the fit was created using penalized maximum likelihood estimation,
the same penalty and penalty.scale parameters are used during
validation.
See https://www.fharrell.com/post/bootcal/ for simulations of the accuracy of various smoothers for binary logistic model calibration, as well as simulations of confidence interval coverage.
Examples
require(survival)
set.seed(1)
n <- 200
d.time <- rexp(n)
x1 <- runif(n)
x2 <- factor(sample(c('a', 'b', 'c'), n, TRUE))
f <- cph(Surv(d.time) ~ pol(x1,2) * x2, x=TRUE, y=TRUE, surv=TRUE, time.inc=1.5)
#or f <- psm(S ~ \dots)
pa <- requireNamespace('polspline')
if(pa) {
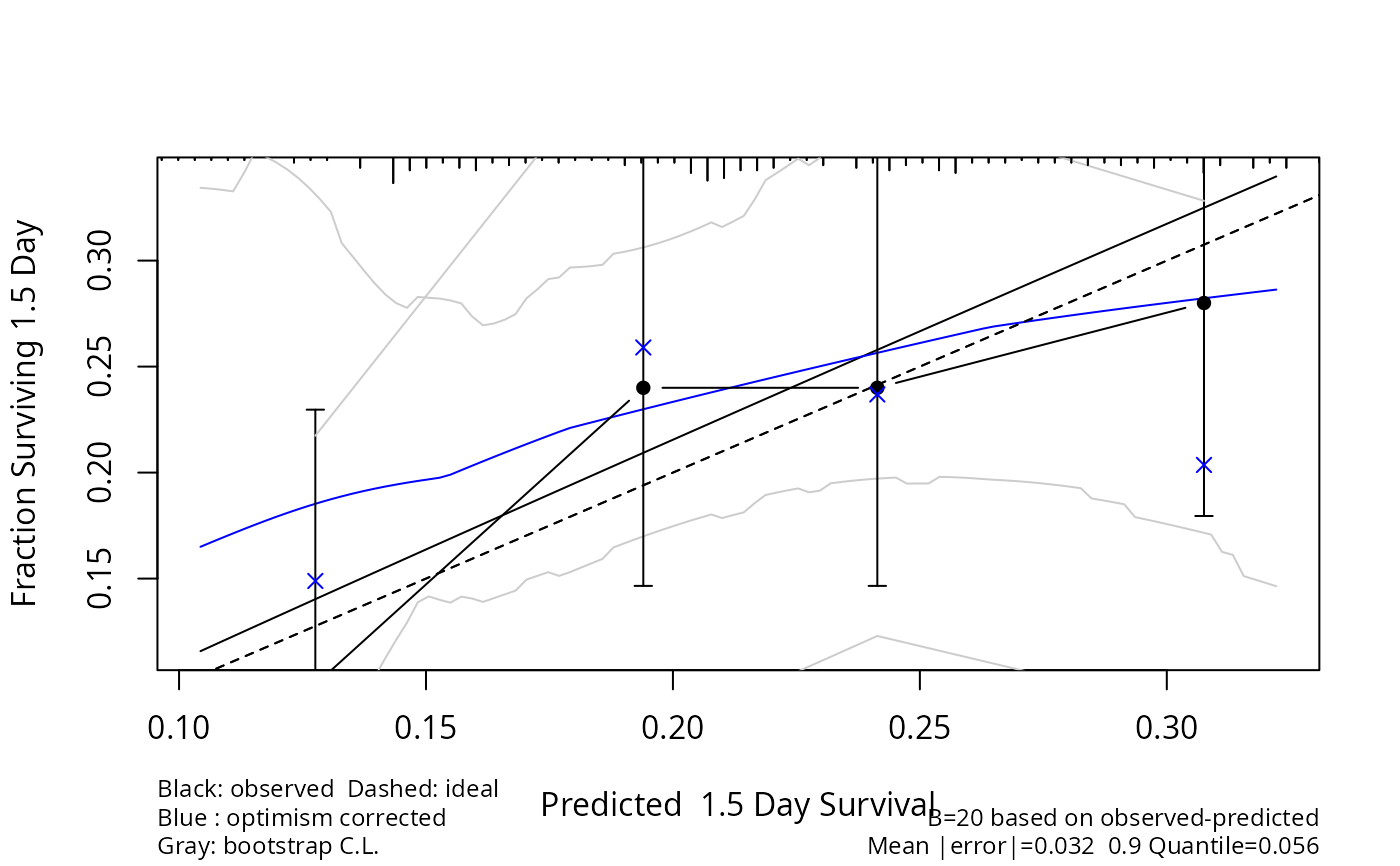
cal <- calibrate(f, u=1.5, B=20) # cmethod='hare'
plot(cal)
}
#> Using Cox survival estimates at 1.5 Days
cal <- calibrate(f, u=1.5, cmethod='KM', m=50, B=20) # usually B=200 or 300
#> Using Cox survival estimates at 1.5 Days
plot(cal, add=pa)
 set.seed(1)
y <- sample(0:2, n, TRUE)
x1 <- runif(n)
x2 <- runif(n)
x3 <- runif(n)
x4 <- runif(n)
f <- lrm(y ~ x1 + x2 + x3 * x4, x=TRUE, y=TRUE)
cal <- calibrate(f, kint=2, predy=seq(.2, .8, length=60),
group=y)
# group= does k-sample validation: make resamples have same
# numbers of subjects in each level of y as original sample
plot(cal)
set.seed(1)
y <- sample(0:2, n, TRUE)
x1 <- runif(n)
x2 <- runif(n)
x3 <- runif(n)
x4 <- runif(n)
f <- lrm(y ~ x1 + x2 + x3 * x4, x=TRUE, y=TRUE)
cal <- calibrate(f, kint=2, predy=seq(.2, .8, length=60),
group=y)
# group= does k-sample validation: make resamples have same
# numbers of subjects in each level of y as original sample
plot(cal)
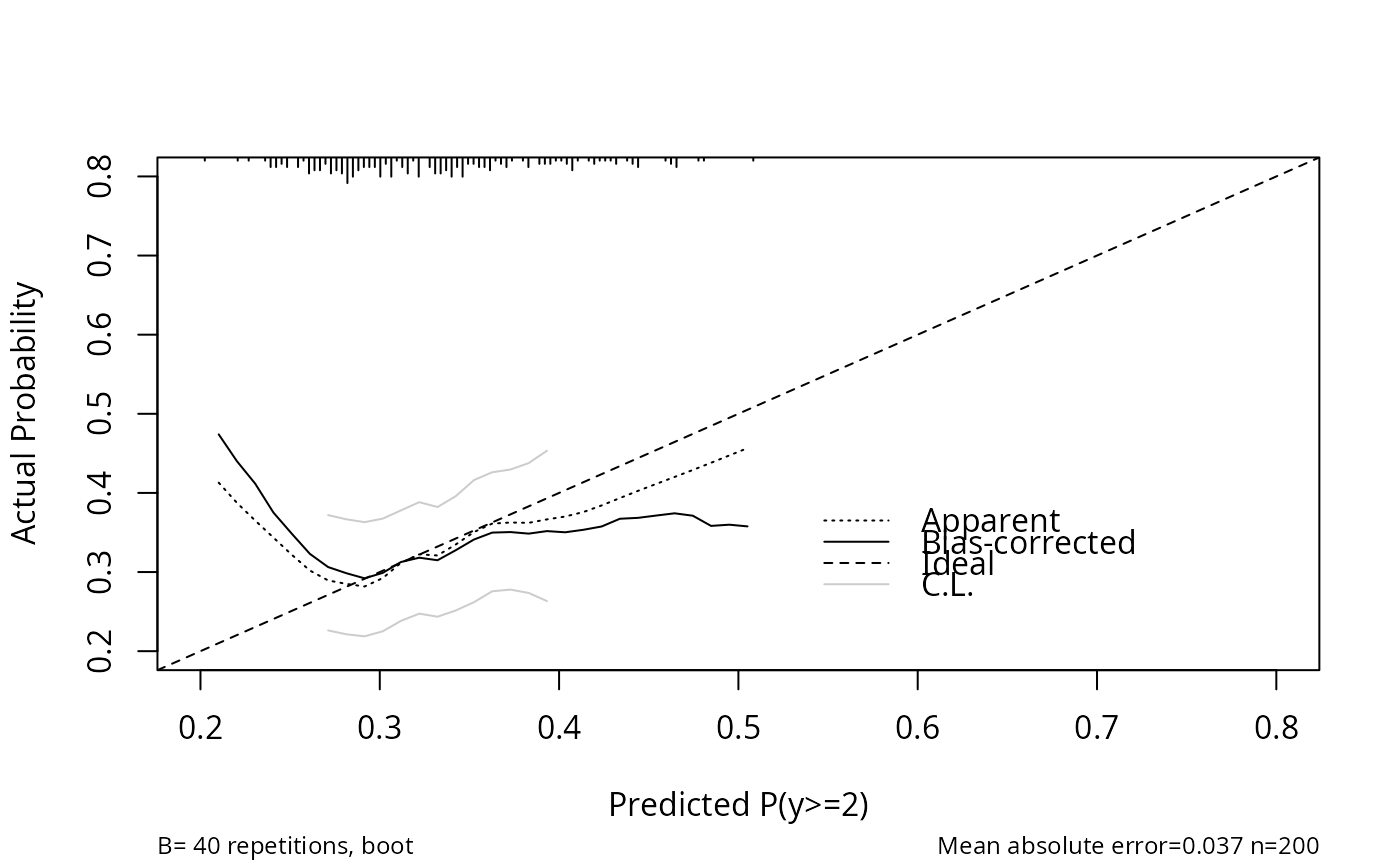 #>
#> n=200 Mean absolute error=0.037 Mean squared error=0.003
#> 0.9 Quantile of absolute error=0.09
#>
#See the example for the validate function for a method of validating
#continuation ratio ordinal logistic models. You can do the same
#thing for calibrate
#>
#> n=200 Mean absolute error=0.037 Mean squared error=0.003
#> 0.9 Quantile of absolute error=0.09
#>
#See the example for the validate function for a method of validating
#continuation ratio ordinal logistic models. You can do the same
#thing for calibrate