Cox Proportional Hazards Model and Extensions
cph.RdModification of Therneau's coxph function to fit the Cox model and
its extension, the Andersen-Gill model. The latter allows for interval
time-dependent covariables, time-dependent strata, and repeated events.
The Survival method for an object created by cph returns an S
function for computing estimates of the survival function.
The Quantile method for cph returns an S function for computing
quantiles of survival time (median, by default).
The Mean method returns a function for computing the mean survival
time. This function issues a warning if the last follow-up time is uncensored,
unless a restricted mean is explicitly requested.
Usage
cph(formula = formula(data), data=environment(formula),
weights, subset, na.action=na.delete,
method=c("efron","breslow","exact","model.frame","model.matrix"),
singular.ok=FALSE, robust=FALSE,
model=FALSE, x=FALSE, y=FALSE, se.fit=FALSE,
linear.predictors=TRUE, residuals=TRUE, nonames=FALSE,
eps=1e-4, init, iter.max=10, tol=1e-9, surv=FALSE, time.inc,
type=NULL, vartype=NULL, debug=FALSE, ...)
# S3 method for class 'cph'
Survival(object, ...)
# Evaluate result as g(times, lp, stratum=1, type=c("step","polygon"))
# S3 method for class 'cph'
Quantile(object, ...)
# Evaluate like h(q, lp, stratum=1, type=c("step","polygon"))
# S3 method for class 'cph'
Mean(object, method=c("exact","approximate"), type=c("step","polygon"),
n=75, tmax, ...)
# E.g. m(lp, stratum=1, type=c("step","polygon"), tmax, \dots)Arguments
- formula
an S formula object with a
Survobject on the left-hand side. Thetermscan specify any S model formula with up to third-order interactions. Thestratfunction may appear in the terms, as a main effect or an interacting factor. To stratify on both race and sex, you would include both termsstrat(race)andstrat(sex). Stratification factors may interact with non-stratification factors; not all stratification terms need interact with the same modeled factors.- object
an object created by
cphwithsurv=TRUE- data
name of an S data frame containing all needed variables. Omit this to use a data frame already in the S “search list”.
- weights
case weights
- subset
an expression defining a subset of the observations to use in the fit. The default is to use all observations. Specify for example
age>50 & sex="male"orc(1:100,200:300)respectively to use the observations satisfying a logical expression or those having row numbers in the given vector.- na.action
specifies an S function to handle missing data. The default is the function
na.delete, which causes observations with any variable missing to be deleted. The main difference betweenna.deleteand the S-supplied functionna.omitis thatna.deletemakes a list of the number of observations that are missing on each variable in the model. Thena.actionis usally specified by e.g.options(na.action="na.delete").- method
for
cph, specifies a particular fitting method,"model.frame"instead to return the model frame of the predictor and response variables satisfying any subset or missing value checks, or"model.matrix"to return the expanded design matrix. The default is"efron", to use Efron's likelihood for fitting the model.For
Mean.cph,methodis"exact"to use numerical integration of the survival function at any linear predictor value to obtain a mean survival time. Specifymethod="approximate"to use an approximate method that is slower whenMean.cphis executing but then is essentially instant thereafter. For the approximate method, the area is computed fornpoints equally spaced between the min and max observed linear predictor values. This calculation is done separately for each stratum. Then thenpairs (X beta, area) are saved in the generated S function, and when this function is evaluated, theapproxfunction is used to evaluate the mean for any given linear predictor values, using linear interpolation over thenX beta values.- singular.ok
If
TRUE, the program will automatically skip over columns of the X matrix that are linear combinations of earlier columns. In this case the coefficients for such columns will be NA, and the variance matrix will contain zeros. For ancillary calculations, such as the linear predictor, the missing coefficients are treated as zeros. The singularities will prevent many of the features of thermslibrary from working.- robust
if
TRUEa robust variance estimate is returned. Default isTRUEif the model includes acluster()operative,FALSEotherwise.- model
default is
FALSE(false). Set toTRUEto return the model frame as elementmodelof the fit object.- x
default is
FALSE. Set toTRUEto return the expanded design matrix as elementx(without intercept indicators) of the returned fit object.- y
default is
FALSE. Set toTRUEto return the vector of response values (Survobject) as elementyof the fit.- se.fit
default is
FALSE. Set toTRUEto compute the estimated standard errors of the estimate of X beta and store them in elementse.fitof the fit. The predictors are first centered to their means before computing the standard errors.- linear.predictors
set to
FALSEto omitlinear.predictorsvector from fit- residuals
set to
FALSEto omitresidualsvector from fit- nonames
set to
TRUEto not setnamesattribute forlinear.predictors,residuals,se.fit, and rows of design matrix- eps
convergence criterion - change in log likelihood.
- init
vector of initial parameter estimates. Defaults to all zeros. Special residuals can be obtained by setting some elements of
initto MLEs and others to zero and specifyingiter.max=1.- iter.max
maximum number of iterations to allow. Set to
0to obtain certain null-model residuals.- tol
tolerance for declaring singularity for matrix inversion (available only when survival5 or later package is in effect)
- surv
set to
TRUEto compute underlying survival estimates for each stratum, and to store these along with standard errors of log Lambda(t),maxtime(maximum observed survival or censoring time), andsurv.summaryin the returned object. Setsurv="summary"to only compute and storesurv.summary, not survival estimates at each unique uncensored failure time. If you specifyx=TRUEandy=TRUE, you can obtain predicted survival later, with accurate confidence intervals for any set of predictor values. The standard error information stored as a result ofsurv=TRUEare only accurate at the mean of all predictors. If the model has no covariables, these are of course OK. The main reason for usingsurvis to greatly speed up the computation of predicted survival probabilities as a function of the covariables, when accurate confidence intervals are not needed.- time.inc
time increment used in deriving
surv.summary. Survival, number at risk, and standard error will be stored fort=0, time.inc, 2 time.inc, ..., maxtime, wheremaxtimeis the maximum survival time over all strata.time.incis also used in constructing the time axis in thesurvplotfunction (see below). The default value fortime.incis 30 ifunits(ftime) = "Day"or nounitsattribute has been attached to the survival time variable. Ifunits(ftime)is a word other than"Day", the default fortime.incis 1 when it is omitted, unlessmaxtime<1, thenmaxtime/10is used astime.inc. Iftime.incis not given andmaxtime/ default time.inc> 25,time.incis increased.- type
(for
cph) applies ifsurvisTRUEor"summary". Iftypeis omitted, the method consistent withmethodis used. Seesurvfit.coxph(undersurvfit) orsurvfit.cphfor details and for the definitions of values oftypeFor
Survival, Quantile, Meanset to"polygon"to use linear interpolation instead of the usual step function. ForMean, the default ofstepwill yield the sample mean in the case of no censoring and no covariables, iftype="kaplan-meier"was specified tocph. Formethod="exact", the value oftypeis passed to the generated function, and it can be overridden when that function is actually invoked. Formethod="approximate",Mean.cphgenerates the function different ways according totype, and this cannot be changed when the function is actually invoked.- vartype
see
survfit.coxph- debug
set to
TRUEto print debugging information related to model matrix construction. You can also useoptions(debug=TRUE).- ...
other arguments passed to
coxph.fitfromcph. Ignored by other functions.- times
a scalar or vector of times at which to evaluate the survival estimates
- lp
a scalar or vector of linear predictors (including the centering constant) at which to evaluate the survival estimates
- stratum
a scalar stratum number or name (e.g.,
"sex=male") to use in getting survival probabilities- q
a scalar quantile or a vector of quantiles to compute
- n
the number of points at which to evaluate the mean survival time, for
method="approximate"inMean.cph.- tmax
For
Mean.cph, the default is to compute the overall mean (and produce a warning message if there is censoring at the end of follow-up). To compute a restricted mean life length, specify the truncation point astmax. Formethod="exact",tmaxis passed to the generated function and it may be overridden when that function is invoked. Formethod="approximate",tmaxmust be specified at the time thatMean.cphis run.
Value
For Survival, Quantile, or Mean, an S function is returned. Otherwise,
in addition to what is listed below, formula/design information and
the components
maxtime, time.inc, units, model, x, y, se.fit are stored, the last 5
depending on the settings of options by the same names.
The vectors or matrix stored if y=TRUE or x=TRUE have rows deleted according to subset and
to missing data, and have names or row names that come from the
data frame used as input data.
- n
table with one row per stratum containing number of censored and uncensored observations
- coef
vector of regression coefficients
- stats
vector containing the named elements
Obs,Events,Model L.R.,d.f.,P,Score,Score P,R2, Somers'Dxy,g-index, andgr, theg-index on the hazard ratio scale.R2is the Nagelkerke R-squared, with division by the maximum attainable R-squared.- var
variance/covariance matrix of coefficients
- linear.predictors
values of predicted X beta for observations used in fit, normalized to have overall mean zero, then having any offsets added
- resid
martingale residuals
- loglik
log likelihood at initial and final parameter values
- score
value of score statistic at initial values of parameters
- times
lists of times (if
surv="T")- surv
lists of underlying survival probability estimates
- std.err
lists of standard errors of estimate log-log survival
- surv.summary
a 3 dimensional array if
surv=TRUE. The first dimension is time ranging from 0 tomaxtimebytime.inc. The second dimension refers to strata. The third dimension contains the time-oriented matrix withSurvival, n.risk(number of subjects at risk), andstd.err(standard error of log-log survival).- center
centering constant, equal to overall mean of X beta.
Details
If there is any strata by covariable interaction in the model such that
the mean X beta varies greatly over strata, method="approximate" may
not yield very accurate estimates of the mean in Mean.cph.
For method="approximate" if you ask for an estimate of the mean for
a linear predictor value that was outside the range of linear predictors
stored with the fit, the mean for that observation will be NA.
Author
Frank Harrell
Department of Biostatistics, Vanderbilt University
fh@fharrell.com
See also
coxph, survival-internal,
Surv, residuals.cph,
cox.zph, survfit.cph,
survest.cph, survfit.coxph,
survplot, datadist,
rms, rms.trans, anova.rms,
summary.rms, Predict,
fastbw, validate, calibrate,
plot.Predict, ggplot.Predict,
specs.rms, lrm, which.influence,
na.delete,
na.detail.response, print.cph,
latex.cph, vif, ie.setup,
GiniMd, dxy.cens,
concordance
Examples
# Simulate data from a population model in which the log hazard
# function is linear in age and there is no age x sex interaction
require(survival)
require(ggplot2)
n <- 1000
set.seed(731)
age <- 50 + 12*rnorm(n)
label(age) <- "Age"
sex <- factor(sample(c('Male','Female'), n,
rep=TRUE, prob=c(.6, .4)))
cens <- 15*runif(n)
h <- .02*exp(.04*(age-50)+.8*(sex=='Female'))
dt <- -log(runif(n))/h
label(dt) <- 'Follow-up Time'
e <- ifelse(dt <= cens,1,0)
dt <- pmin(dt, cens)
units(dt) <- "Year"
dd <- datadist(age, sex)
options(datadist='dd')
S <- Surv(dt,e)
f <- cph(S ~ rcs(age,4) + sex, x=TRUE, y=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
cox.zph(f, "rank") # tests of PH
#> Error: object 'f' not found
anova(f)
#> Error: object 'f' not found
ggplot(Predict(f, age, sex)) # plot age effect, 2 curves for 2 sexes
#> Error: object 'f' not found
survplot(f, sex) # time on x-axis, curves for x2
#> Error: object 'f' not found
res <- resid(f, "scaledsch")
#> Error: object 'f' not found
time <- as.numeric(dimnames(res)[[1]])
#> Error: object 'res' not found
z <- loess(res[,4] ~ time, span=0.50) # residuals for sex
#> Error in eval(predvars, data, env): object 'res' not found
plot(time, fitted(z))
#> Error: object 'z' not found
lines(supsmu(time, res[,4]),lty=2)
#> Error: object 'res' not found
plot(cox.zph(f,"identity")) #Easier approach for last few lines
#> Error: object 'f' not found
# latex(f)
f <- cph(S ~ age + strat(sex), surv=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
g <- Survival(f) # g is a function
#> Error: object 'f' not found
g(seq(.1,1,by=.1), stratum="sex=Male", type="poly") #could use stratum=2
#> Error in g(seq(0.1, 1, by = 0.1), stratum = "sex=Male", type = "poly"): could not find function "g"
med <- Quantile(f)
#> Error: object 'f' not found
plot(Predict(f, age, fun=function(x) med(lp=x))) #plot median survival
#> Error: object 'f' not found
# Fit a model that is quadratic in age, interacting with sex as strata
# Compare standard errors of linear predictor values with those from
# coxph
# Use more stringent convergence criteria to match with coxph
f <- cph(S ~ pol(age,2)*strat(sex), x=TRUE, eps=1e-9, iter.max=20)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
coef(f)
#> Error: object 'f' not found
se <- predict(f, se.fit=TRUE)$se.fit
#> Error: object 'f' not found
require(lattice)
xyplot(se ~ age | sex, main='From cph')
#> Error in eval(i, data, env): object 'se' not found
a <- c(30,50,70)
comb <- data.frame(age=rep(a, each=2),
sex=rep(levels(sex), 3))
p <- predict(f, comb, se.fit=TRUE)
#> Error: object 'f' not found
comb$yhat <- p$linear.predictors
#> Error: object 'p' not found
comb$se <- p$se.fit
#> Error: object 'p' not found
z <- qnorm(.975)
comb$lower <- p$linear.predictors - z*p$se.fit
#> Error: object 'p' not found
comb$upper <- p$linear.predictors + z*p$se.fit
#> Error: object 'p' not found
comb
#> age sex
#> 1 30 Female
#> 2 30 Male
#> 3 50 Female
#> 4 50 Male
#> 5 70 Female
#> 6 70 Male
age2 <- age^2
f2 <- coxph(S ~ (age + age2)*strata(sex))
coef(f2)
#> age age2 age:strata(sex)Male
#> 0.100744 -0.000548 -0.020831
#> age2:strata(sex)Male
#> 0.000276
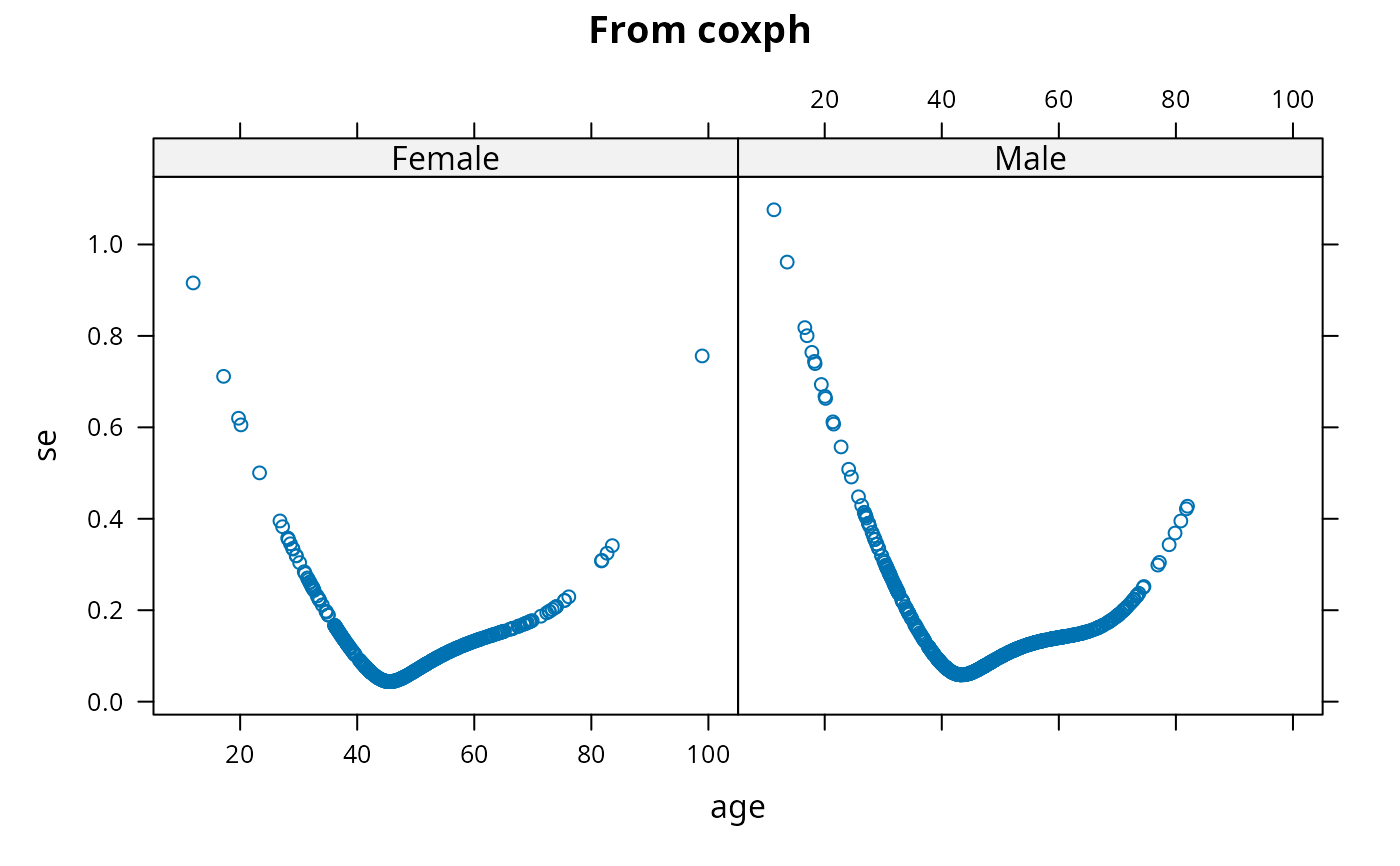
se <- predict(f2, se.fit=TRUE)$se.fit
xyplot(se ~ age | sex, main='From coxph')
 comb <- data.frame(age=rep(a, each=2), age2=rep(a, each=2)^2,
sex=rep(levels(sex), 3))
p <- predict(f2, newdata=comb, se.fit=TRUE)
comb$yhat <- p$fit
comb$se <- p$se.fit
comb$lower <- p$fit - z*p$se.fit
comb$upper <- p$fit + z*p$se.fit
comb
#> age age2 sex yhat se lower upper
#> 1 30 900 Female -1.0414 0.3084 -1.6458 -0.437
#> 2 30 900 Male -1.0665 0.3111 -1.6762 -0.457
#> 3 50 2500 Female 0.0964 0.0683 -0.0375 0.230
#> 4 50 2500 Male 0.0966 0.0979 -0.0953 0.289
#> 5 70 4900 Female 0.7957 0.1780 0.4469 1.145
#> 6 70 4900 Male 1.0423 0.1880 0.6738 1.411
# g <- cph(Surv(hospital.charges) ~ age, surv=TRUE)
# Cox model very useful for analyzing highly skewed data, censored or not
# m <- Mean(g)
# m(0) # Predicted mean charge for reference age
#Fit a time-dependent covariable representing the instantaneous effect
#of an intervening non-fatal event
rm(age)
set.seed(121)
dframe <- data.frame(failure.time=1:10, event=rep(0:1,5),
ie.time=c(NA,1.5,2.5,NA,3,4,NA,5,5,5),
age=sample(40:80,10,rep=TRUE))
z <- ie.setup(dframe$failure.time, dframe$event, dframe$ie.time)
S <- z$S
ie.status <- z$ie.status
attach(dframe[z$subs,]) # replicates all variables
f <- cph(S ~ age + ie.status, x=TRUE, y=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
#Must use x=TRUE,y=TRUE to get survival curves with time-dep. covariables
#Get estimated survival curve for a 50-year old who has an intervening
#non-fatal event at 5 days
new <- data.frame(S=Surv(c(0,5), c(5,999), c(FALSE,FALSE)), age=rep(50,2),
ie.status=c(0,1))
g <- survfit(f, new)
#> Error: object 'f' not found
plot(c(0,g$time), c(1,g$surv[,2]), type='s',
xlab='Days', ylab='Survival Prob.')
#> Error: object 'g' not found
# Not certain about what columns represent in g$surv for survival5
# but appears to be for different ie.status
#or:
#g <- survest(f, new)
#plot(g$time, g$surv, type='s', xlab='Days', ylab='Survival Prob.')
#Compare with estimates when there is no intervening event
new2 <- data.frame(S=Surv(c(0,5), c(5, 999), c(FALSE,FALSE)), age=rep(50,2),
ie.status=c(0,0))
g2 <- survfit(f, new2)
#> Error: object 'f' not found
lines(c(0,g2$time), c(1,g2$surv[,2]), type='s', lty=2)
#> Error: object 'g2' not found
#or:
#g2 <- survest(f, new2)
#lines(g2$time, g2$surv, type='s', lty=2)
detach("dframe[z$subs, ]")
options(datadist=NULL)
comb <- data.frame(age=rep(a, each=2), age2=rep(a, each=2)^2,
sex=rep(levels(sex), 3))
p <- predict(f2, newdata=comb, se.fit=TRUE)
comb$yhat <- p$fit
comb$se <- p$se.fit
comb$lower <- p$fit - z*p$se.fit
comb$upper <- p$fit + z*p$se.fit
comb
#> age age2 sex yhat se lower upper
#> 1 30 900 Female -1.0414 0.3084 -1.6458 -0.437
#> 2 30 900 Male -1.0665 0.3111 -1.6762 -0.457
#> 3 50 2500 Female 0.0964 0.0683 -0.0375 0.230
#> 4 50 2500 Male 0.0966 0.0979 -0.0953 0.289
#> 5 70 4900 Female 0.7957 0.1780 0.4469 1.145
#> 6 70 4900 Male 1.0423 0.1880 0.6738 1.411
# g <- cph(Surv(hospital.charges) ~ age, surv=TRUE)
# Cox model very useful for analyzing highly skewed data, censored or not
# m <- Mean(g)
# m(0) # Predicted mean charge for reference age
#Fit a time-dependent covariable representing the instantaneous effect
#of an intervening non-fatal event
rm(age)
set.seed(121)
dframe <- data.frame(failure.time=1:10, event=rep(0:1,5),
ie.time=c(NA,1.5,2.5,NA,3,4,NA,5,5,5),
age=sample(40:80,10,rep=TRUE))
z <- ie.setup(dframe$failure.time, dframe$event, dframe$ie.time)
S <- z$S
ie.status <- z$ie.status
attach(dframe[z$subs,]) # replicates all variables
f <- cph(S ~ age + ie.status, x=TRUE, y=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
#Must use x=TRUE,y=TRUE to get survival curves with time-dep. covariables
#Get estimated survival curve for a 50-year old who has an intervening
#non-fatal event at 5 days
new <- data.frame(S=Surv(c(0,5), c(5,999), c(FALSE,FALSE)), age=rep(50,2),
ie.status=c(0,1))
g <- survfit(f, new)
#> Error: object 'f' not found
plot(c(0,g$time), c(1,g$surv[,2]), type='s',
xlab='Days', ylab='Survival Prob.')
#> Error: object 'g' not found
# Not certain about what columns represent in g$surv for survival5
# but appears to be for different ie.status
#or:
#g <- survest(f, new)
#plot(g$time, g$surv, type='s', xlab='Days', ylab='Survival Prob.')
#Compare with estimates when there is no intervening event
new2 <- data.frame(S=Surv(c(0,5), c(5, 999), c(FALSE,FALSE)), age=rep(50,2),
ie.status=c(0,0))
g2 <- survfit(f, new2)
#> Error: object 'f' not found
lines(c(0,g2$time), c(1,g2$surv[,2]), type='s', lty=2)
#> Error: object 'g2' not found
#or:
#g2 <- survest(f, new2)
#lines(g2$time, g2$surv, type='s', lty=2)
detach("dframe[z$subs, ]")
options(datadist=NULL)