Weighted Statistical Estimates
wtd.stats.RdThese functions compute various weighted versions of standard
estimators. In most cases the weights vector is a vector the same
length of x, containing frequency counts that in effect expand x
by these counts. weights can also be sampling weights, in which
setting normwt to TRUE will often be appropriate. This results in
making weights sum to the length of the non-missing elements in
x. normwt=TRUE thus reflects the fact that the true sample size is
the length of the x vector and not the sum of the original values of
weights (which would be appropriate had normwt=FALSE). When weights
is all ones, the estimates are all identical to unweighted estimates
(unless one of the non-default quantile estimation options is
specified to wtd.quantile). When missing data have already been
deleted for, x, weights, and (in the case of wtd.loess.noiter) y,
specifying na.rm=FALSE will save computation time. Omitting the
weights argument or specifying NULL or a zero-length vector will
result in the usual unweighted estimates.
wtd.mean, wtd.var, and wtd.quantile compute
weighted means, variances, and quantiles, respectively. wtd.Ecdf
computes a weighted empirical distribution function. wtd.table
computes a weighted frequency table (although only one stratification
variable is supported at present). wtd.rank computes weighted
ranks, using mid–ranks for ties. This can be used to obtain Wilcoxon
tests and rank correlation coefficients. wtd.loess.noiter is a
weighted version of loess.smooth when no iterations for outlier
rejection are desired. This results in especially good smoothing when
y is binary. wtd.quantile removes any observations with
zero weight at the beginning. Previously, these were changing the
quantile estimates.
num.denom.setup is a utility function that allows one to deal with
observations containing numbers of events and numbers of trials, by
outputting two observations when the number of events and non-events
(trials - events) exceed zero. A vector of subscripts is generated
that will do the proper duplications of observations, and a new binary
variable y is created along with usual cell frequencies (weights)
for each of the y=0, y=1 cells per observation.
Usage
wtd.mean(x, weights=NULL, normwt="ignored", na.rm=TRUE)
wtd.var(x, weights=NULL, normwt=FALSE, na.rm=TRUE,
method=c('unbiased', 'ML'))
wtd.quantile(x, weights=NULL, probs=c(0, .25, .5, .75, 1),
type=c('quantile','(i-1)/(n-1)','i/(n+1)','i/n'),
normwt=FALSE, na.rm=TRUE)
wtd.Ecdf(x, weights=NULL,
type=c('i/n','(i-1)/(n-1)','i/(n+1)'),
normwt=FALSE, na.rm=TRUE)
wtd.table(x, weights=NULL, type=c('list','table'),
normwt=FALSE, na.rm=TRUE)
wtd.rank(x, weights=NULL, normwt=FALSE, na.rm=TRUE)
wtd.loess.noiter(x, y, weights=rep(1,n),
span=2/3, degree=1, cell=.13333,
type=c('all','ordered all','evaluate'),
evaluation=100, na.rm=TRUE)
num.denom.setup(num, denom)Arguments
- x
a numeric vector (may be a character or
categoryorfactorvector forwtd.table)- num
vector of numerator frequencies
- denom
vector of denominators (numbers of trials)
- weights
a numeric vector of weights
- normwt
specify
normwt=TRUEto makeweightssum tolength(x)after deletion ofNAs. Ifweightsare frequency weights, thennormwtshould beFALSE, and ifweightsare normalization (aka reliability) weights, thennormwtshould beTRUE. In the case of the former, no check is made thatweightsare valid frequencies.- na.rm
set to
FALSEto suppress checking for NAs- method
determines the estimator type; if
'unbiased'(the default) then the usual unbiased estimate (using Bessel's correction) is returned, if'ML'then it is the maximum likelihood estimate for a Gaussian distribution. In the case of the latter, thenormwtargument has no effect. Usesstats:cov.wtfor both methods.- probs
a vector of quantiles to compute. Default is 0 (min), .25, .5, .75, 1 (max).
- type
For
wtd.quantile,typedefaults toquantileto use the same interpolated order statistic method asquantile. Settypeto"(i-1)/(n-1)","i/(n+1)", or"i/n"to use the inverse of the empirical distribution function, using, respectively, (wt - 1)/T, wt/(T+1), or wt/T, where wt is the cumulative weight and T is the total weight (usually total sample size). These three values oftypeare the possibilities forwtd.Ecdf. Forwtd.tablethe defaulttypeis"list", meaning that the function is to return a list containing two vectors:xis the sorted unique values ofxandsum.of.weightsis the sum of weights for thatx. This is the default so that you don't have to convert thenamesattribute of the result that can be obtained withtype="table"to a numeric variable whenxwas originally numeric.type="table"forwtd.tableresults in an object that is the same structure as those returned fromtable. Forwtd.loess.noiterthe defaulttypeis"all", indicating that the function is to return a list containing all the original values ofx(including duplicates and without sorting) and the smoothedyvalues corresponding to them. Settype="ordered all"to sort byx, andtype="evaluate"to evaluate the smooth only atevaluationequally spaced points between the observed limits ofx.- y
a numeric vector the same length as
x- span, degree, cell, evaluation
see
loess.smooth. The default is linear (degree=1) and 100 points to evaluation (iftype="evaluate").
Value
wtd.mean and wtd.var return scalars. wtd.quantile returns a
vector the same length as probs. wtd.Ecdf returns a list whose
elements x and Ecdf correspond to unique sorted values of x.
If the first CDF estimate is greater than zero, a point (min(x),0) is
placed at the beginning of the estimates.
See above for wtd.table. wtd.rank returns a vector the same
length as x (after removal of NAs, depending on na.rm). See above
for wtd.loess.noiter.
Details
The functions correctly combine weights of observations having
duplicate values of x before computing estimates.
When normwt=FALSE the weighted variance will not equal the
unweighted variance even if the weights are identical. That is because
of the subtraction of 1 from the sum of the weights in the denominator
of the variance formula. If you want the weighted variance to equal the
unweighted variance when weights do not vary, use normwt=TRUE.
The articles by Gatz and Smith discuss alternative approaches, to arrive
at estimators of the standard error of a weighted mean.
wtd.rank does not handle NAs as elegantly as rank if
weights is specified.
Author
Frank Harrell
Department of Biostatistics
Vanderbilt University School of Medicine
fh@fharrell.com
Benjamin Tyner
btyner@gmail.com
References
Research Triangle Institute (1995): SUDAAN User's Manual, Release 6.40, pp. 8-16 to 8-17.
Gatz DF, Smith L (1995): The standard error of a weighted mean concentration–I. Bootstrapping vs other methods. Atmospheric Env 11:1185-1193.
Gatz DF, Smith L (1995): The standard error of a weighted mean concentration–II. Estimating confidence intervals. Atmospheric Env 29:1195-1200.
https://en.wikipedia.org/wiki/Weighted_arithmetic_mean
Examples
set.seed(1)
x <- runif(500)
wts <- sample(1:6, 500, TRUE)
std.dev <- sqrt(wtd.var(x, wts))
wtd.quantile(x, wts)
#> 0% 25% 50% 75% 100%
#> 0.00184 0.26024 0.46155 0.73964 0.99608
death <- sample(0:1, 500, TRUE)
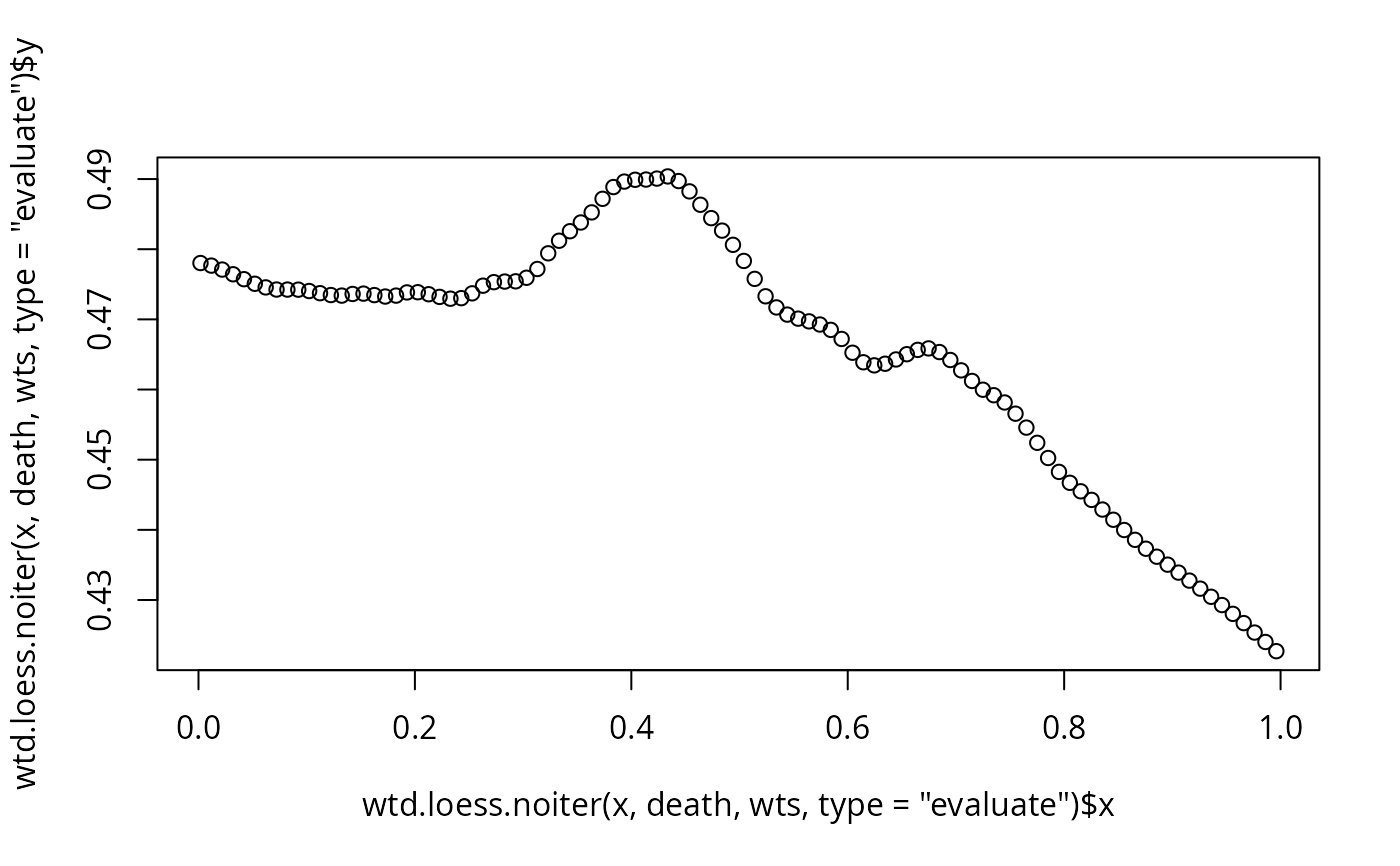
plot(wtd.loess.noiter(x, death, wts, type='evaluate'))
 describe(~x, weights=wts)
#> x
#>
#> 2 Variables 500 Observations
#> --------------------------------------------------------------------------------
#> x
#> n missing distinct Info Mean .05 .10 .25
#> 1780 0 500 1 0.4949 0.07068 0.12169 0.26024
#> .50 .75 .90 .95
#> 0.46155 0.73964 0.91037 0.95373
#>
#> lowest : 0.00183686 0.00193283 0.0111495 0.0130776 0.0133903
#> highest: 0.991839 0.991906 0.992684 0.993749 0.996077
#> --------------------------------------------------------------------------------
#> (weights)
#> n missing distinct Info Mean
#> 1780 0 6 0.949 4.385
#>
#> Value 1 2 3 4 5 6
#> Frequency 79 174 219 328 470 510
#> Proportion 0.044 0.098 0.123 0.184 0.264 0.287
#> --------------------------------------------------------------------------------
# describe uses wtd.mean, wtd.quantile, wtd.table
xg <- cut2(x,g=4)
table(xg)
#> xg
#> [0.00184,0.258) [0.25817,0.476) [0.47635,0.740) [0.73964,0.996]
#> 125 125 125 125
wtd.table(xg, wts, type='table')
#> [0.00184,0.258) [0.25817,0.476) [0.47635,0.740) [0.73964,0.996]
#> 444 467 421 448
# Here is a method for getting stratified weighted means
y <- runif(500)
g <- function(y) wtd.mean(y[,1],y[,2])
summarize(cbind(y, wts), llist(xg), g, stat.name='y')
#> xg y
#> 1 [0.00184,0.258) 0.444
#> 2 [0.25817,0.476) 0.500
#> 3 [0.47635,0.740) 0.512
#> 4 [0.73964,0.996] 0.496
# Empirically determine how methods used by wtd.quantile match with
# methods used by quantile, when all weights are unity
set.seed(1)
u <- eval(formals(wtd.quantile)$type)
v <- as.character(1:9)
r <- matrix(0, nrow=length(u), ncol=9, dimnames=list(u,v))
for(n in c(8, 13, 22, 29))
{
x <- rnorm(n)
for(i in 1:5) {
probs <- sort( runif(9))
for(wtype in u) {
wq <- wtd.quantile(x, type=wtype, weights=rep(1,length(x)), probs=probs)
for(qtype in 1:9) {
rq <- quantile(x, type=qtype, probs=probs)
r[wtype, qtype] <- max(r[wtype,qtype], max(abs(wq-rq)))
}
}
}
}
r
#> 1 2 3 4 5 6 7 8 9
#> quantile 0.573 0.573 0.755 5.02e-01 0.373 6.42e-01 0.00e+00 0.472 0.442
#> (i-1)/(n-1) 0.573 0.573 0.755 5.02e-01 0.373 6.42e-01 8.88e-16 0.472 0.442
#> i/(n+1) 0.709 0.709 0.857 7.20e-01 0.321 4.44e-16 6.42e-01 0.214 0.241
#> i/n 0.777 0.777 0.409 4.44e-16 0.428 7.20e-01 5.02e-01 0.546 0.517
# Restructure data to generate a dichotomous response variable
# from records containing numbers of events and numbers of trials
num <- c(10,NA,20,0,15) # data are 10/12 NA/999 20/20 0/25 15/35
denom <- c(12,999,20,25,35)
w <- num.denom.setup(num, denom)
w
#> $subs
#> [1] 1 3 5 1 4 5
#>
#> $weights
#> [1] 10 20 15 2 25 20
#>
#> $y
#> [1] 1 1 1 0 0 0
#>
# attach(my.data.frame[w$subs,])
describe(~x, weights=wts)
#> x
#>
#> 2 Variables 500 Observations
#> --------------------------------------------------------------------------------
#> x
#> n missing distinct Info Mean .05 .10 .25
#> 1780 0 500 1 0.4949 0.07068 0.12169 0.26024
#> .50 .75 .90 .95
#> 0.46155 0.73964 0.91037 0.95373
#>
#> lowest : 0.00183686 0.00193283 0.0111495 0.0130776 0.0133903
#> highest: 0.991839 0.991906 0.992684 0.993749 0.996077
#> --------------------------------------------------------------------------------
#> (weights)
#> n missing distinct Info Mean
#> 1780 0 6 0.949 4.385
#>
#> Value 1 2 3 4 5 6
#> Frequency 79 174 219 328 470 510
#> Proportion 0.044 0.098 0.123 0.184 0.264 0.287
#> --------------------------------------------------------------------------------
# describe uses wtd.mean, wtd.quantile, wtd.table
xg <- cut2(x,g=4)
table(xg)
#> xg
#> [0.00184,0.258) [0.25817,0.476) [0.47635,0.740) [0.73964,0.996]
#> 125 125 125 125
wtd.table(xg, wts, type='table')
#> [0.00184,0.258) [0.25817,0.476) [0.47635,0.740) [0.73964,0.996]
#> 444 467 421 448
# Here is a method for getting stratified weighted means
y <- runif(500)
g <- function(y) wtd.mean(y[,1],y[,2])
summarize(cbind(y, wts), llist(xg), g, stat.name='y')
#> xg y
#> 1 [0.00184,0.258) 0.444
#> 2 [0.25817,0.476) 0.500
#> 3 [0.47635,0.740) 0.512
#> 4 [0.73964,0.996] 0.496
# Empirically determine how methods used by wtd.quantile match with
# methods used by quantile, when all weights are unity
set.seed(1)
u <- eval(formals(wtd.quantile)$type)
v <- as.character(1:9)
r <- matrix(0, nrow=length(u), ncol=9, dimnames=list(u,v))
for(n in c(8, 13, 22, 29))
{
x <- rnorm(n)
for(i in 1:5) {
probs <- sort( runif(9))
for(wtype in u) {
wq <- wtd.quantile(x, type=wtype, weights=rep(1,length(x)), probs=probs)
for(qtype in 1:9) {
rq <- quantile(x, type=qtype, probs=probs)
r[wtype, qtype] <- max(r[wtype,qtype], max(abs(wq-rq)))
}
}
}
}
r
#> 1 2 3 4 5 6 7 8 9
#> quantile 0.573 0.573 0.755 5.02e-01 0.373 6.42e-01 0.00e+00 0.472 0.442
#> (i-1)/(n-1) 0.573 0.573 0.755 5.02e-01 0.373 6.42e-01 8.88e-16 0.472 0.442
#> i/(n+1) 0.709 0.709 0.857 7.20e-01 0.321 4.44e-16 6.42e-01 0.214 0.241
#> i/n 0.777 0.777 0.409 4.44e-16 0.428 7.20e-01 5.02e-01 0.546 0.517
# Restructure data to generate a dichotomous response variable
# from records containing numbers of events and numbers of trials
num <- c(10,NA,20,0,15) # data are 10/12 NA/999 20/20 0/25 15/35
denom <- c(12,999,20,25,35)
w <- num.denom.setup(num, denom)
w
#> $subs
#> [1] 1 3 5 1 4 5
#>
#> $weights
#> [1] 10 20 15 2 25 20
#>
#> $y
#> [1] 1 1 1 0 0 0
#>
# attach(my.data.frame[w$subs,])