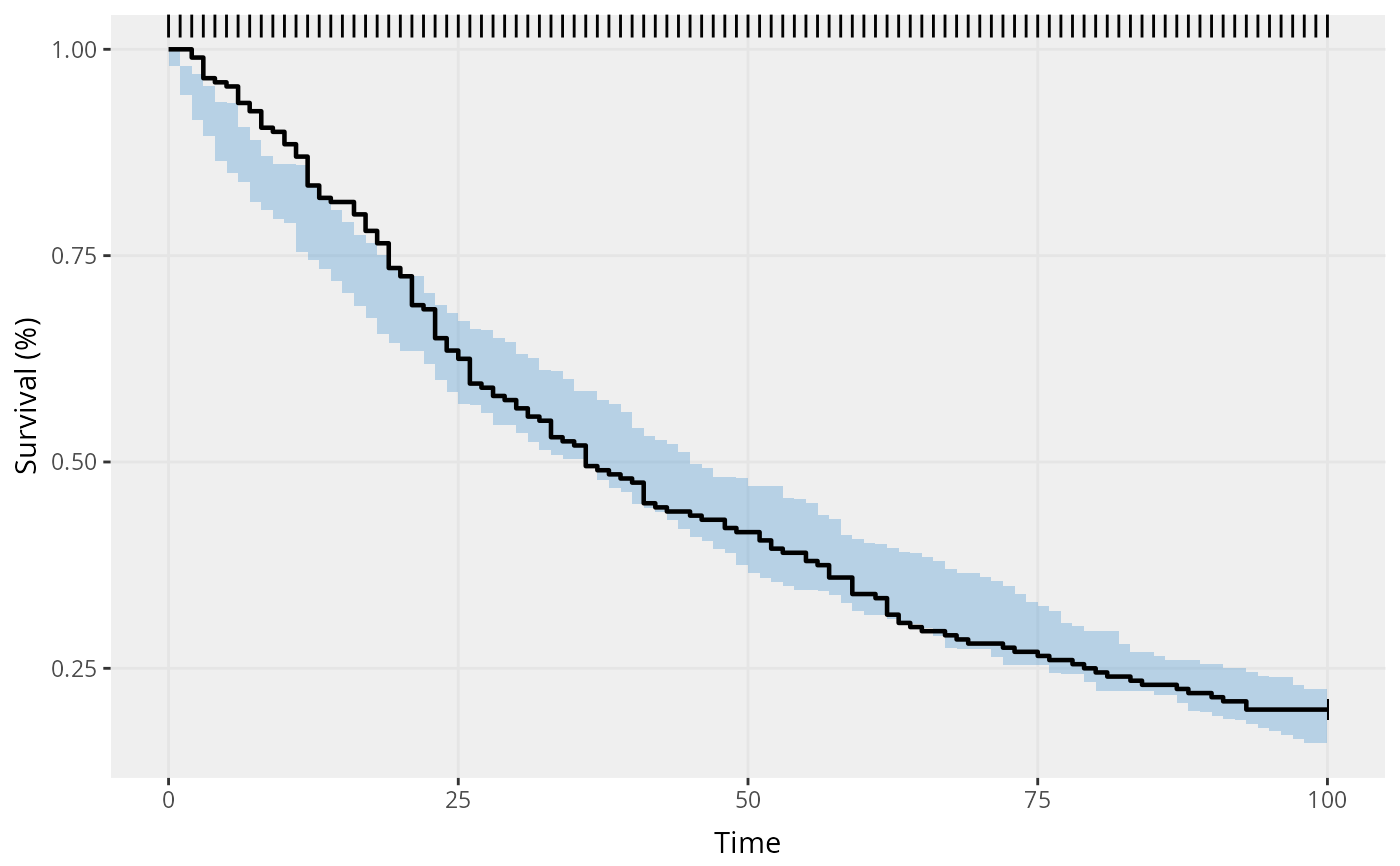
This function can be used for either single time-to-event (TTE) or repeated time-to-event (RTTE) data.
Usage
vpc_tte(
sim = NULL,
obs = NULL,
psn_folder = NULL,
rtte = FALSE,
rtte_calc_diff = TRUE,
rtte_conditional = TRUE,
events = NULL,
bins = FALSE,
n_bins = 10,
software = "auto",
obs_cols = NULL,
sim_cols = NULL,
kmmc = NULL,
reverse_prob = FALSE,
stratify = NULL,
stratify_color = NULL,
ci = c(0.05, 0.95),
xlab = "Time",
ylab = "Survival (%)",
show = NULL,
as_percentage = TRUE,
title = NULL,
smooth = FALSE,
vpc_theme = NULL,
facet = "wrap",
labeller = NULL,
verbose = FALSE,
vpcdb = FALSE
)Arguments
- sim
this is usually a data.frame with observed data, containing the independent and dependent variable, a column indicating the individual, and possibly covariates. E.g. load in from NONMEM using read_table_nm. However it can also be an object like a nlmixr or xpose object
- obs
a data.frame with observed data, containing the independent and dependent variable, a column indicating the individual, and possibly covariates. E.g. load in from NONMEM using read_table_nm
- psn_folder
instead of specifying "sim" and "obs", specify a PsN-generated VPC-folder
- rtte
repeated time-to-event data? Default is FALSE (treat as single-event TTE)
- rtte_calc_diff
recalculate time (T/F)? When simulating in NONMEM, you will probably need to set this to TRUE to recalculate the TIME to relative times between events (unless you output the time difference between events and specify that as independent variable to the vpc_tte() function.
- rtte_conditional
`TRUE` (default) or `FALSE`. Compute the probability for each event newly (`TRUE`), or calculate the absolute probability (`FALSE`, i.e. the "probability of a 1st, 2nd, 3rd event etc" rather than the "probability of an event happening").
- events
numeric vector describing which events to show a VPC for when repeated TTE data, e.g. c(1:4). Default is NULL, which shows all events.
- bins
either "density", "time", or "data", "none", or one of the approaches available in classInterval() such as "jenks" (default) or "pretty", or a numeric vector specifying the bin separators.
- n_bins
when using the "auto" binning method, what number of bins to aim for
- software
name of software platform using (e.g. nonmem, phoenix)
- obs_cols
list for mapping observation data columns, e.g. `list(dv = "DV", id = "ID", idv = "TIME", pred="PRED")`
- sim_cols
list for mapping simulation data columns, e.g. `list(dv = "DV", id = "ID", idv = "TIME", pred="PRED")`
- kmmc
either NULL (for regular TTE vpc, default), or a variable name for a KMMC plot (e.g. "WT")
- reverse_prob
reverse the probability scale (i.e. plot 1-probability)
- stratify
character vector of stratification variables. Only 1 or 2 stratification variables can be supplied.
- stratify_color
character vector of stratification variables. Only 1 stratification variable can be supplied, cannot be used in conjunction with `stratify`.
- ci
confidence interval to plot. Default is (0.05, 0.95)
- xlab
label for x axis
- ylab
label for y axis
- show
what to show in VPC (obs_dv, obs_ci, pi, pi_as_area, pi_ci, obs_median, sim_median, sim_median_ci)
- as_percentage
Show y-scale from 0-100 percent? TRUE by default, if FALSE then scale from 0-1.
- title
title
- smooth
"smooth" the VPC (connect bin midpoints) or show bins as rectangular boxes. Default is TRUE.
- vpc_theme
theme to be used in VPC. Expects list of class vpc_theme created with function vpc_theme()
- facet
either "wrap", "columns", or "rows"
- labeller
ggplot2 labeller function to be passed to underlying ggplot object
- verbose
show debugging information (TRUE or FALSE)
- vpcdb
Boolean whether to return the underlying vpcdb rather than the plot
Examples
## See vpc-docs.ronkeizer.com for more documentation and examples.
## Example for repeated) time-to-event data
## with NONMEM-like data (e.g. simulated using a dense grid)
data(rtte_obs_nm)
data(rtte_sim_nm)
# treat RTTE as TTE, no stratification
vpc_tte(sim = rtte_sim_nm[rtte_sim_nm$sim <= 20,],
obs = rtte_obs_nm,
rtte = FALSE,
sim_cols=list(dv = "dv", idv = "t"), obs_cols=list(idv = "t"))
#> Warning: No software packages matched for filtering values, not filtering.
#> Object class: other, data.frame
#> Available filters: phoenix, nonmem
#> Warning: No software packages matched for filtering values, not filtering.
#> Object class: other, data.frame
#> Available filters: phoenix, nonmem