Plot Survival Curves and Hazard Functions
survplot.RdPlot estimated survival curves, and for parametric survival models, plot
hazard functions. There is an option to print the number of subjects
at risk at the start of each time interval for certain models. Curves are automatically
labeled at the points of maximum separation (using the labcurve
function), and there are many other options for labeling that can be
specified with the label.curves parameter. For example, different
plotting symbols can be placed at constant x-increments and a legend
linking the symbols with category labels can automatically positioned on
the most empty portion of the plot.
If the fit is from psm and ggplot=TRUE is specified, a ggplot2 graphic
will instead be produced using the survplot.orm function.
For the case of a two stratum analysis by npsurv,
survdiffplot plots the difference in two Kaplan-Meier estimates
along with approximate confidence bands for the differences, with a
reference line at zero. The number of subjects at risk is optionally
plotted. This number is taken as the minimum of the number of subjects
at risk over the two strata. When conf='diffbands',
survdiffplot instead does not make a new plot but adds a shaded
polygon to an existing plot, showing the midpoint of two survival
estimates plus or minus 1/2 the width of the confidence interval for the
difference of two Kaplan-Meier estimates.
survplotp creates an interactive plotly graphic with
shaded confidence bands for fits other than from orms.
In the two strata case, it draws the 1/2
confidence bands for the difference in two probabilities centered at the
midpoint of the probability estimates, so that where the two curves
touch this band there is no significant difference (no multiplicity
adjustment is made). For the two strata case, the two individual
confidence bands have entries in the legend but are not displayed until
the user clicks on the legend.
When code was from running npsurv on a
multi-state/competing risk Surv object, survplot plots
cumulative incidence curves properly accounting for competing risks.
You must specify exactly one state/event cause to plot using the
state argument. survplot will not plot multiple states on
one graph. This can be accomplished using multiple calls with different
values of state and specifying add=TRUE for all but the
first call.
Usage
survplot(fit, ...)
survplotp(fit, ...)
# S3 method for class 'rms'
survplot(fit, ..., xlim,
ylim=if(loglog) c(-5, 1.5) else if
(what == "survival" & missing(fun)) c(0, 1),
xlab, ylab, time.inc,
what=c("survival","hazard"),
type=c("tsiatis","kaplan-meier"),
conf.type=c("log","log-log","plain","none"),
conf.int=FALSE, conf=c("bands","bars"), mylim=NULL,
add=FALSE, label.curves=TRUE,
abbrev.label=FALSE, levels.only=FALSE,
lty, lwd=par("lwd"),
col=1, col.fill=gray(seq(.95, .75, length=5)),
adj.subtitle=TRUE, loglog=FALSE, fun,
n.risk=FALSE, logt=FALSE, dots=FALSE, dotsize=.003,
grid=NULL, srt.n.risk=0, sep.n.risk=0.056, adj.n.risk=1,
y.n.risk, cex.n.risk=.6, cex.xlab=par('cex.lab'),
cex.ylab=cex.xlab, pr=FALSE, ggplot=FALSE)
# S3 method for class 'npsurv'
survplot(fit, xlim,
ylim, xlab, ylab, time.inc, state=NULL,
conf=c("bands","bars","diffbands","none"), mylim=NULL,
add=FALSE, label.curves=TRUE, abbrev.label=FALSE,
levels.only=FALSE, lty,lwd=par('lwd'),
col=1, col.fill=gray(seq(.95, .75, length=5)),
loglog=FALSE, fun, n.risk=FALSE, aehaz=FALSE, times=NULL,
logt=FALSE, dots=FALSE, dotsize=.003, grid=NULL,
srt.n.risk=0, sep.n.risk=.056, adj.n.risk=1,
y.n.risk, cex.n.risk=.6, cex.xlab=par('cex.lab'), cex.ylab=cex.xlab,
pr=FALSE, ...)
# S3 method for class 'npsurv'
survplotp(fit, xlim, ylim, xlab, ylab, time.inc, state=NULL,
conf=c("bands", "none"), mylim=NULL, abbrev.label=FALSE,
col=colorspace::rainbow_hcl, levels.only=TRUE,
loglog=FALSE, fun=function(y) y, aehaz=FALSE, times=NULL,
logt=FALSE, pr=FALSE, ...)
survdiffplot(fit, order=1:2, fun=function(y) y,
xlim, ylim, xlab, ylab="Difference in Survival Probability",
time.inc, conf.int, conf=c("shaded", "bands","diffbands","none"),
add=FALSE, lty=1, lwd=par('lwd'), col=1,
n.risk=FALSE, grid=NULL,
srt.n.risk=0, adj.n.risk=1,
y.n.risk, cex.n.risk=.6, cex.xlab=par('cex.lab'),
cex.ylab=cex.xlab, convert=function(f) f)Arguments
- fit
result of fit (
cph,psm,npsurv,survest.psm). Forsurvdiffplot,fitmust be the result ofnpsurv.- ...
list of factors with names used in model. For fits from
npsurvthese arguments do not appear - all strata are plotted. Otherwise the first factor listed is the factor used to determine different survival curves. Any other factors are used to specify single constants to be adjusted to, when defaults given to fitting routine (throughlimits) are not used. The value given to factors is the original coding of data given to fit, except that for categorical or strata factors the text string levels may be specified. The form of values given to the first factor are none (omit the equal sign to use default range or list of all values if variable is discrete),"text"if factor is categorical,c(value1, value2, ...), or a function which returns a vector, such asseq(low,high,by=increment). Only the first factor may have the values omitted. In this case theLow effect,Adjust to, andHigh effectvalues will be used fromdatadistif the variable is continuous. For variables not defined todatadist, you must specify non-missing constant settings (or a vector of settings for the one displayed variable). Note that sincenpsurvobjects do not use the variable list in..., you can specify any extra arguments tolabcurveby adding them at the end of the list of arguments. Forsurvplotp... (e.g.,height,width) is passed toplotly::plot_ly.- xlim
a vector of two numbers specifiying the x-axis range for follow-up time. Default is
(0,maxtime)wheremaxtimewas thepretty()d version of the maximum follow-up time in any stratum, stored infit$maxtime. Iflogt=TRUE, default is(1, log(maxtime)).- ylim
y-axis limits. Default is
c(0,1)for survival, andc(-5,1.5)ifloglog=TRUE. Iffunorloglog=TRUEare given andylimis not, the limits will be computed from the data. Forwhat="hazard", default limits are computed from the first hazard function plotted.- xlab
x-axis label. Default is
unitsattribute of failure time variable given toSurv.- ylab
y-axis label. Default is
"Survival Probability"or"log(-log Survival Probability)". Iffunis given, the default is"". Forwhat="hazard", the default is"Hazard Function". For a multi-state/competing risk application the default is"Cumulative Incidence".- time.inc
time increment for labeling the x-axis and printing numbers at risk. If not specified, the value of
time.incstored with the model fit will be used.- state
the state/event cause to use in plotting if the fit was for a multi-state/competing risk
Survobject- type
specifies type of estimates,
"tsiatis"(the default) or"kaplan-meier"."tsiatis"here corresponds to the Breslow estimator. This is ignored if survival estimates stored withsurv=TRUEare being used. For fits fromnpsurv, this argument is also ignored, since it is specified as an argument tonpsurv.- conf.type
specifies the basis for confidence limits. This argument is ignored for fits from
npsurv.- conf.int
Default is
FALSE. Specify e.g..95to plot 0.95 confidence bands. For fits from parametric survival models, or Cox models withx=TRUEandy=TRUEspecified to the fit, the exact asymptotic formulas will be used to compute standard errors, and confidence limits are based onlog(-log S(t))ifloglog=TRUE. Ifx=TRUEandy=TRUEwere not specified tocphbutsurv=TRUEwas, the standard errors stored for the underlying survival curve(s) will be used. These agree with the former if predictions are requested at the mean value of X beta or if there are only stratification factors in the model. This argument is ignored for fits fromnpsurv, which must have previously specified confidence interval specifications. Forsurvdiffplotifconf.intis not specified, the level used in the call tonpsurvwill be used.- conf
"bars"for confidence bars at eachtime.inctime point. If the fit was fromcph(..., surv=TRUE), thetime.incused will be that stored with the fit. Useconf="bands"(the default) for bands using standard errors at each failure time. Fornpsurvobjects only,confmay also be"none", indicating that confidence interval information stored with thenpsurvresult should be ignored. Fornpsurvandsurvdiffplot,confmay be"diffbands"whereby a shaded region is drawn for comparing two curves. The polygon is centered at the midpoint of the two survival estimates and the height of the polygon is 1/2 the width of the approximateconf.intpointwise confidence region. Survival curves not overlapping the shaded area are approximately significantly different at the1 - conf.intlevel.- mylim
used to curtail computed
ylim. Whenylimis not given by the user, the computed limits are expanded to force inclusion of the values specified inmylim.- what
defaults to
"survival"to plot survival estimates. Set to"hazard"or an abbreviation to plot the hazard function (forpsmfits only). Confidence intervals are not available forwhat="hazard".- add
set to
TRUEto add curves to an existing plot.- label.curves
default is
TRUEto uselabcurveto label curves where they are farthest apart. Setlabel.curvesto alistto specify options tolabcurve, e.g.,label.curves=list(method="arrow", cex=.8). These option names may be abbreviated in the usual way arguments are abbreviated. Use for examplelabel.curves=list(keys=1:5)to draw symbols (as inpch=1:5- seepoints) on the curves and automatically position a legend in the most empty part of the plot. Setlabel.curves=FALSEto suppress drawing curve labels. Thecol,lty,lwd, andtypeparameters are automatically passed tolabcurve, although you can override them here. To distinguish curves by line types and still havelabcurveconstruct a legend, use for examplelabel.curves=list(keys="lines"). The negative value for the plotting symbol will suppress a plotting symbol from being drawn either on the curves or in the legend.- abbrev.label
set to
TRUEtoabbreviate()curve labels that are plotted- levels.only
set to
TRUEto removevariablename=from the start of curve labels.- lty
vector of line types to use for different factor levels. Default is
c(1,3,4,5,6,7,...).- lwd
vector of line widths to use for different factor levels. Default is current
parsetting forlwd.- col
color for curve, default is
1. Specify a vector to assign different colors to different curves. Forsurvplotp,colis a vector of colors corresponding to strata, or a function that will be called to generate such colors.- col.fill
a vector of colors to used in filling confidence bands
- adj.subtitle
set to
FALSEto suppress plotting subtitle with levels of adjustment factors not plotted. Defaults toTRUE. This argument is ignored fornpsurv.- loglog
set to
TRUEto plotlog(-log Survival)instead ofSurvival- fun
specifies any function to translate estimates and confidence limits before plotting. If the fit is a multi-state object the default for
funisfunction(y) 1 - yto draw cumulative incidence curves.- logt
set to
TRUEto plotlog(t)instead ofton the x-axis- n.risk
set to
TRUEto add number of subjects at risk for each curve, using thesurv.summarycreated bycphor using the failure times used in fitting the model ify=TRUEwas specified to the fit or if the fit was fromnpsurv. The numbers are placed at the bottom of the graph unlessy.n.riskis given. If the fit is fromsurvest.psm,n.riskdoes not apply.- srt.n.risk
angle of rotation for leftmost number of subjects at risk (since this number may run into the second or into the y-axis). Default is
0.- adj.n.risk
justification for leftmost number at risk. Default is
1for right justification. Use0for left justification,.5for centered.- sep.n.risk
multiple of upper y limit - lower y limit for separating lines of text containing number of subjects at risk. Default is
.056*(ylim[2]-ylim[1]).- y.n.risk
When
n.risk=TRUE, the default is to place numbers of patients at risk above the x-axis. You can specify a y-coordinate for the bottom line of the numbers usingy.n.risk. Specifyy.n.risk='auto'to place the numbers below the x-axis at a distance of 1/3 of the range ofylim.- cex.n.risk
character size for number of subjects at risk (when
n.riskisTRUE)- cex.xlab
cexfor x-axis label- cex.ylab
cexfor y-axis label- dots
set to
TRUEto plot a grid of dots. Will be plotted at everytime.inc(seecph) and at survival increments of .1 (ifd>.4), .05 (if.2 < d <= .4), or .025 (ifd <= .2), wheredis the range of survival displayed.- dotsize
size of dots in inches
- grid
defaults to
NULL(not drawing grid lines). Set toTRUEto plotgray(.8)grid lines, or specify any color.- pr
set to
TRUEto print survival curve coordinates used in the plots- ggplot
set to
TRUEto usesurvplot.ormto draw the curves instead, for apsmfit- aehaz
set to
TRUEto add number of events and exponential distribution hazard rate estimates in curve labels. For competing risk data the number of events is for the cause of interest, and the hazard rate is the number of events divided by the sum of all failure and censoring times.- times
a numeric vector of times at which to compute cumulative incidence probability estimates to add to curve labels
- order
an integer vector of length two specifying the order of groups when computing survival differences. The default of
1:2indicates that the second group is subtracted from the first. Specifyorder=2:1to instead subtract the first from the second. A subtitle indicates what was done.- convert
a function to convert the output of
summary.survfitmsto pick off the data needed for a single state
Value
list with components adjust (text string specifying adjustment levels)
and curve.labels (vector of text strings corresponding to levels
of factor used to distinguish curves). For npsurv, the returned
value is the vector of strata labels, or NULL if there are no strata.
Side Effects
plots. If par()$mar[4] < 4, issues par(mar=) to increment mar[4] by 2
if n.risk=TRUE and add=FALSE. The user may want to reset par(mar) in
this case to not leave such a wide right margin for plots. You usually
would issue par(mar=c(5,4,4,2)+.1).
Details
survplot will not work for Cox models with time-dependent covariables.
Use survest or survfit for that purpose.
There is a set a system option mgp.axis.labels to allow x
and y-axes to have differing mgp graphical parameters (see par).
This is important when labels for y-axis tick marks are to be written
horizontally (par(las=1)), as a larger gap between the labels and
the tick marks are needed. You can set the axis-specific 2nd
component of mgp using mgp.axis.labels(c(xvalue,yvalue)).
References
Boers M (2004): Null bar and null zone are better than the error bar to compare group means in graphs. J Clin Epi 57:712-715.
Examples
# Simulate data from a population model in which the log hazard
# function is linear in age and there is no age x sex interaction
require(survival)
n <- 1000
set.seed(731)
age <- 50 + 12*rnorm(n)
label(age) <- "Age"
sex <- factor(sample(c('male','female'), n, TRUE))
cens <- 15*runif(n)
h <- .02*exp(.04*(age-50)+.8*(sex=='female'))
dt <- -log(runif(n))/h
label(dt) <- 'Follow-up Time'
e <- ifelse(dt <= cens,1,0)
dt <- pmin(dt, cens)
units(dt) <- "Year"
dd <- datadist(age, sex)
options(datadist='dd')
S <- Surv(dt,e)
# When age is in the model by itself and we predict at the mean age,
# approximate confidence intervals are ok
f <- cph(S ~ age, surv=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
survplot(f, age=mean(age), conf.int=.95)
#> Error: object 'f' not found
g <- cph(S ~ age, x=TRUE, y=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
survplot(g, age=mean(age), conf.int=.95, add=TRUE, col='red', conf='bars')
#> Error: object 'g' not found
# Repeat for an age far from the mean; not ok
survplot(f, age=75, conf.int=.95)
#> Error: object 'f' not found
survplot(g, age=75, conf.int=.95, add=TRUE, col='red', conf='bars')
#> Error: object 'g' not found
#Plot stratified survival curves by sex, adj for quadratic age effect
# with age x sex interaction (2 d.f. interaction)
f <- cph(S ~ pol(age,2)*strat(sex), x=TRUE, y=TRUE)
#> Error in Design(data, formula, specials = c("strat", "strata")): dataset dd not found for options(datadist=)
#or f <- psm(S ~ pol(age,2)*sex)
Predict(f, sex, age=c(30,50,70))
#> Error: object 'f' not found
survplot(f, sex, n.risk=TRUE, levels.only=TRUE) #Adjust age to median
#> Error: object 'f' not found
survplot(f, sex, logt=TRUE, loglog=TRUE) #Check for Weibull-ness (linearity)
#> Error: object 'f' not found
survplot(f, sex=c("male","female"), age=50)
#> Error: object 'f' not found
#Would have worked without datadist
#or with an incomplete datadist
survplot(f, sex, label.curves=list(keys=c(2,0), point.inc=2))
#> Error: object 'f' not found
#Identify curves with symbols
survplot(f, sex, label.curves=list(keys=c('m','f')))
#> Error: object 'f' not found
#Identify curves with single letters
#Plots by quintiles of age, adjusting sex to male
options(digits=3)
survplot(f, age=quantile(age,(1:4)/5), sex="male")
#> Error: object 'f' not found
#Plot survival Kaplan-Meier survival estimates for males
f <- npsurv(S ~ 1, subset=sex=="male")
survplot(f)
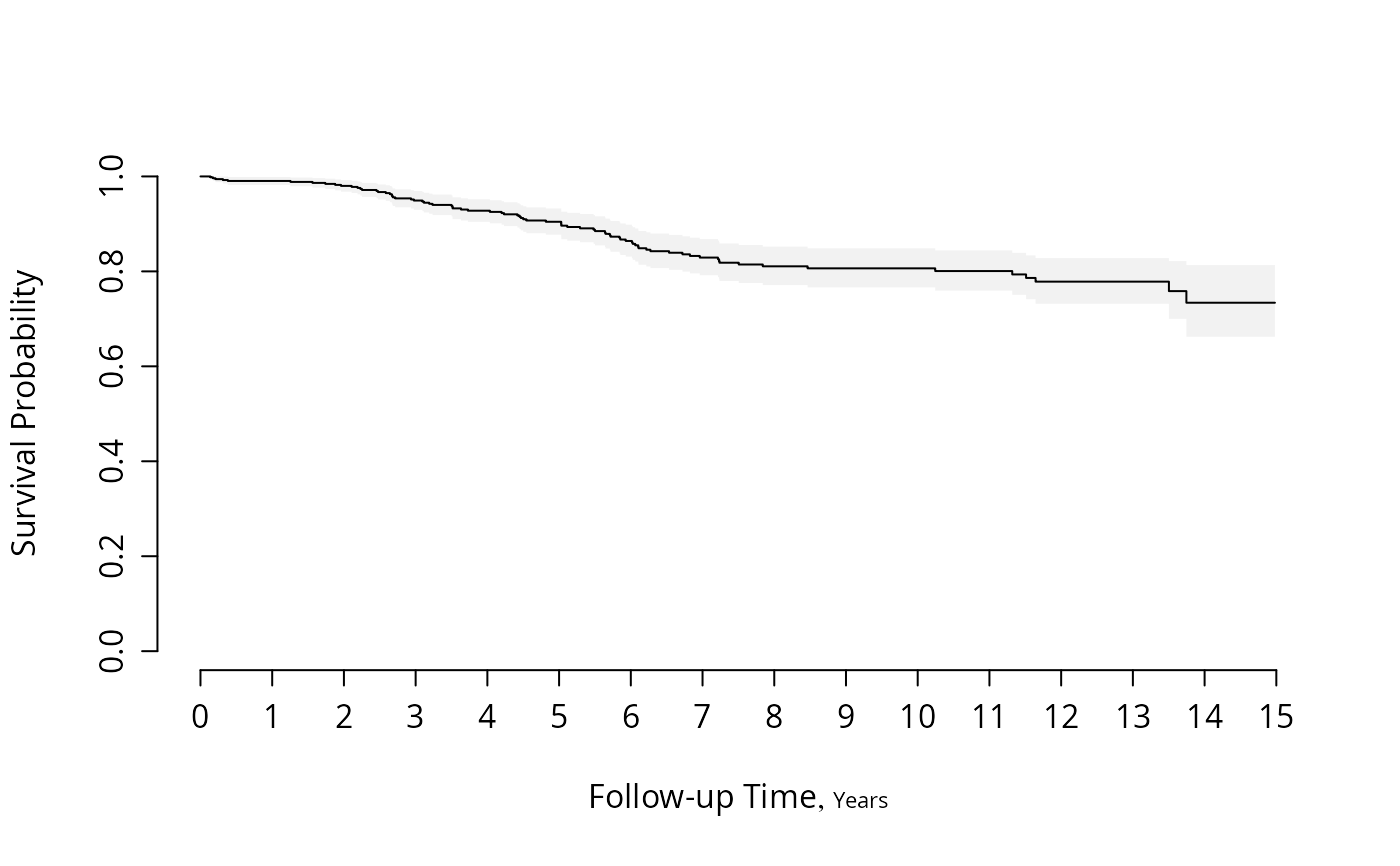 #Plot survival for both sexes and show exponential hazard estimates
f <- npsurv(S ~ sex)
survplot(f, aehaz=TRUE)
#Plot survival for both sexes and show exponential hazard estimates
f <- npsurv(S ~ sex)
survplot(f, aehaz=TRUE)
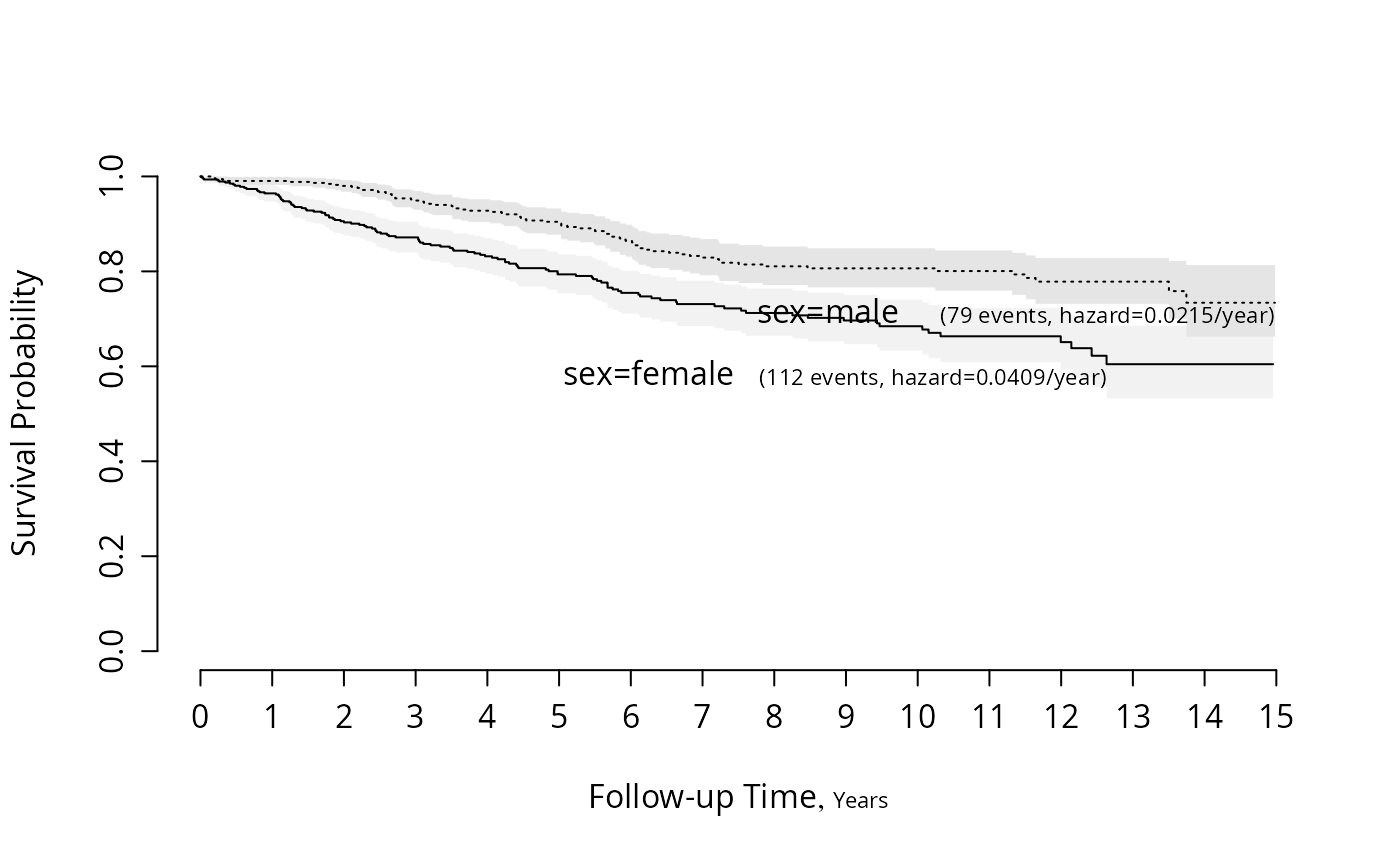 #Check for log-normal and log-logistic fits
survplot(f, fun=qnorm, ylab="Inverse Normal Transform")
#Check for log-normal and log-logistic fits
survplot(f, fun=qnorm, ylab="Inverse Normal Transform")
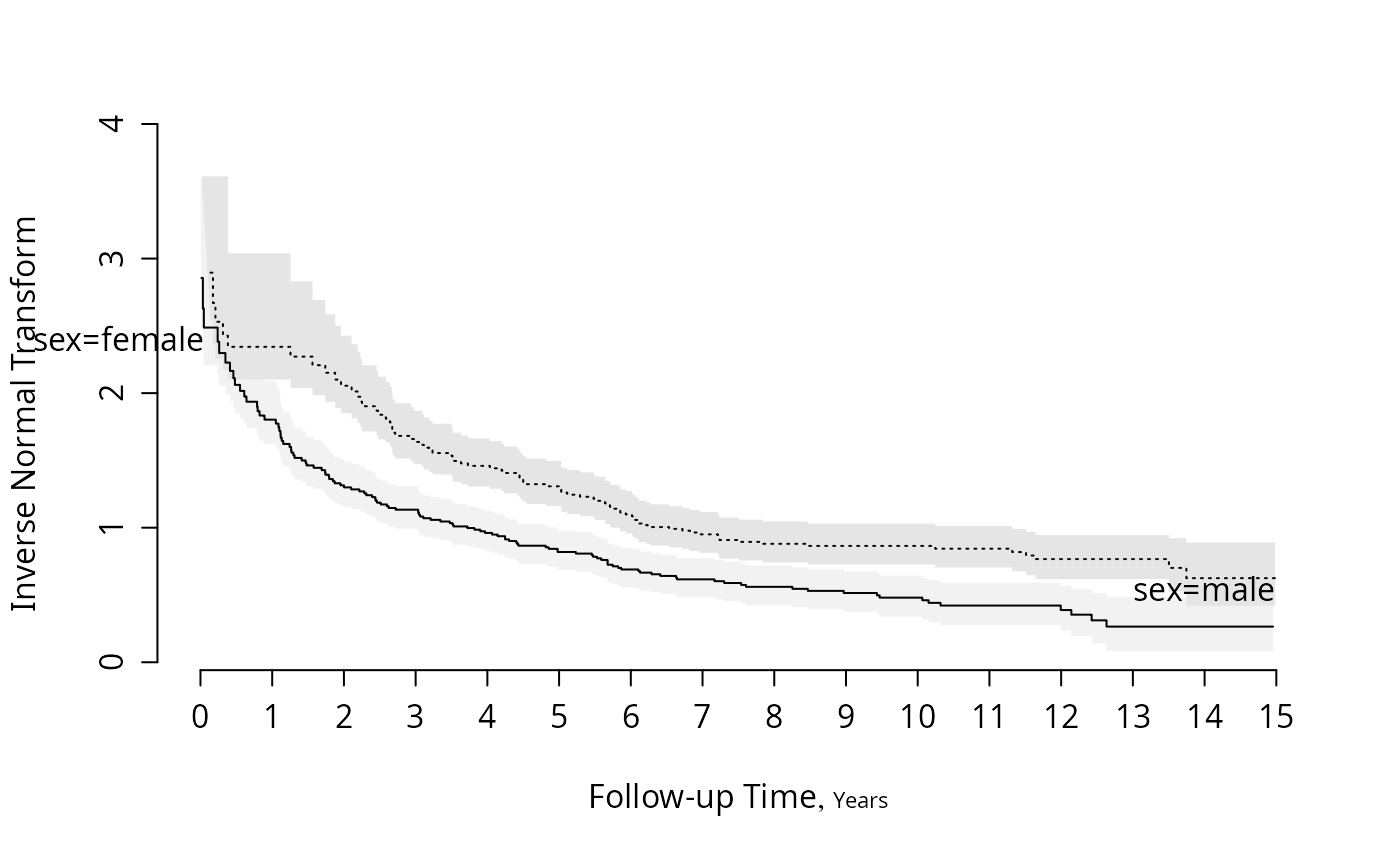 survplot(f, fun=function(y)log(y/(1-y)), ylab="Logit S(t)")
survplot(f, fun=function(y)log(y/(1-y)), ylab="Logit S(t)")
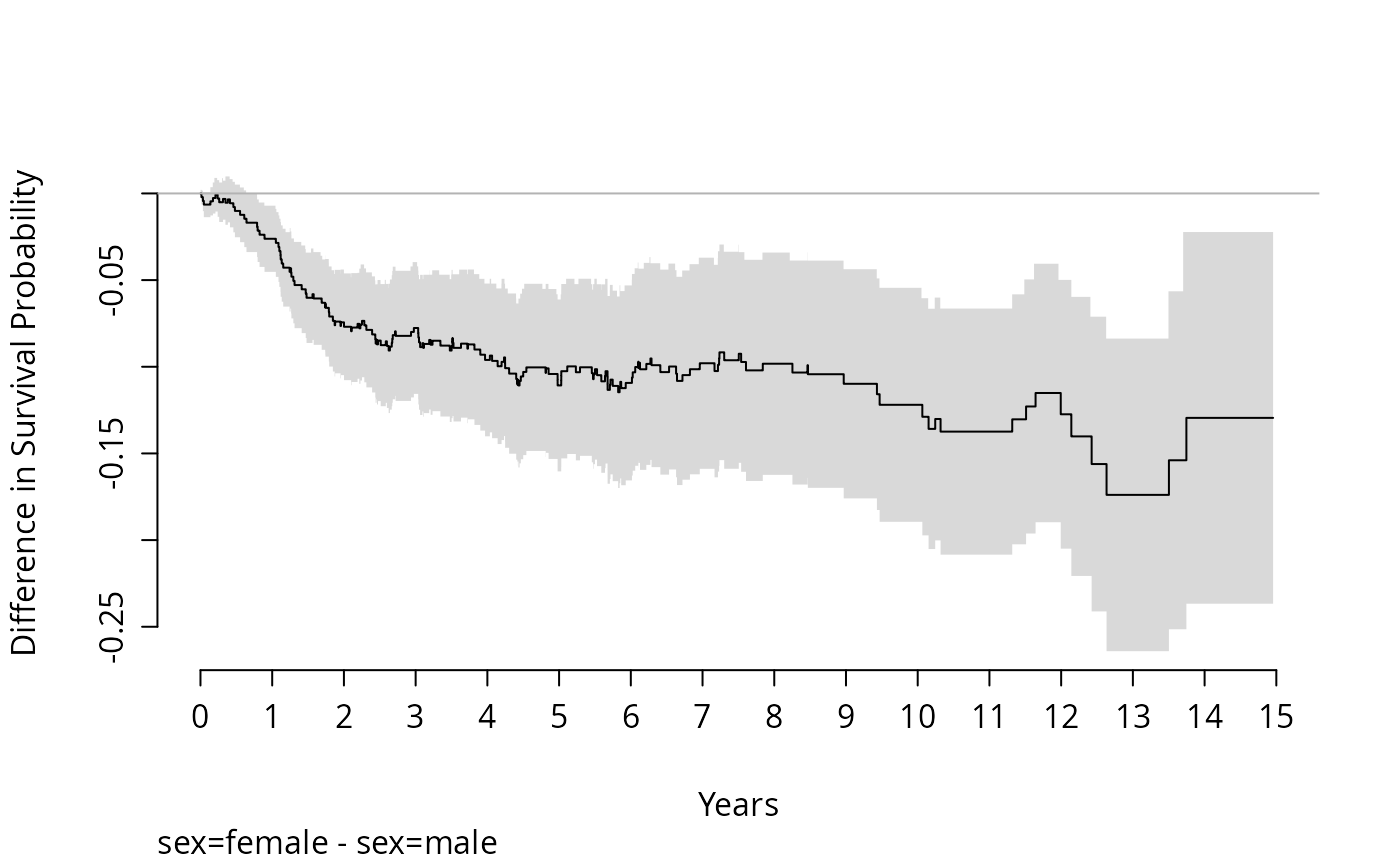
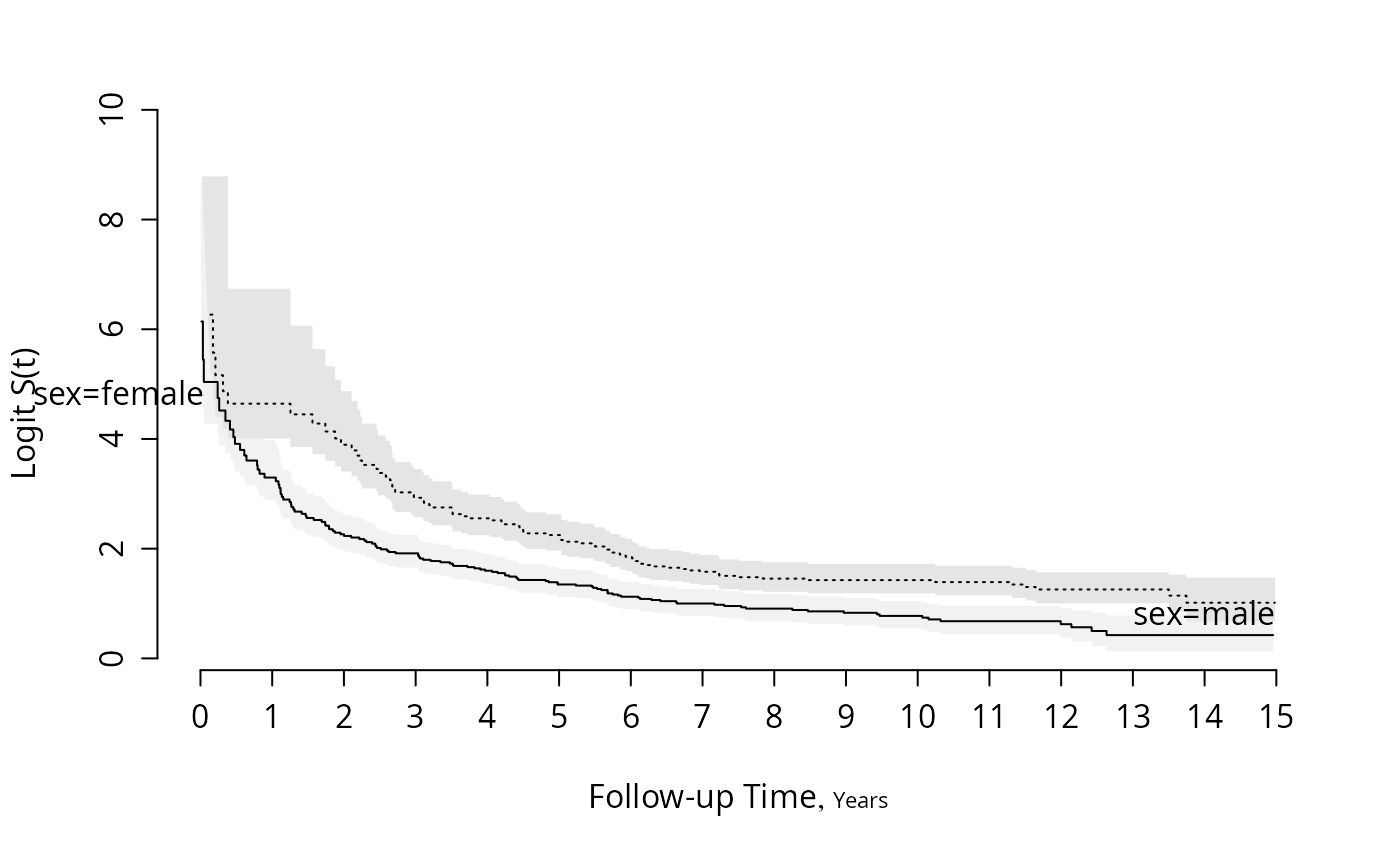 #Plot the difference between sexes
survdiffplot(f)
#Plot the difference between sexes
survdiffplot(f)
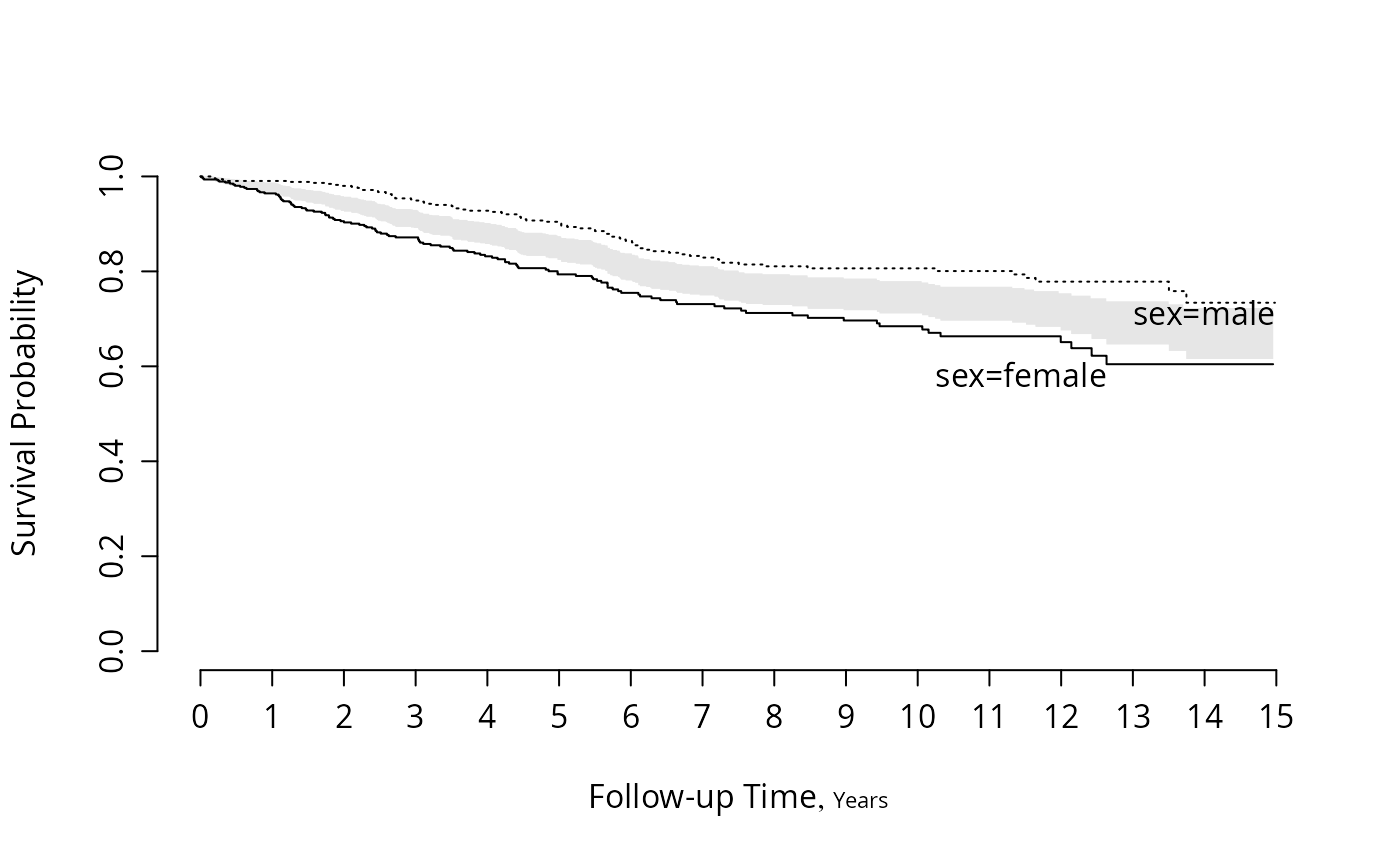
 #Similar but show half-width of confidence intervals centered
#at average of two survival estimates
#See Boers (2004)
survplot(f, conf='diffbands')
#Similar but show half-width of confidence intervals centered
#at average of two survival estimates
#See Boers (2004)
survplot(f, conf='diffbands')
 options(datadist=NULL)
if (FALSE) { # \dontrun{
#
# Time to progression/death for patients with monoclonal gammopathy
# Competing risk curves (cumulative incidence)
# status variable must be a factor with first level denoting right censoring
m <- upData(mgus1, stop = stop / 365.25, units=c(stop='years'),
labels=c(stop='Follow-up Time'), subset=start == 0)
f <- npsurv(Surv(stop, event) ~ 1, data=m)
# Use survplot for enhanced displays of cumulative incidence curves for
# competing risks
survplot(f, state='pcm', n.risk=TRUE, xlim=c(0, 20), ylim=c(0, .5), col=2)
survplot(f, state='death', aehaz=TRUE, col=3,
label.curves=list(keys='lines'))
f <- npsurv(Surv(stop, event) ~ sex, data=m)
survplot(f, state='death', aehaz=TRUE, n.risk=TRUE, conf='diffbands',
label.curves=list(keys='lines'))
# Plot survival curves estimated from an ordinal semiparametric model
f <- orm(Ocens(y, ifelse(y <= cens, y, Inf)) ~ age)
survplot(f, age=c(30, 50))
} # }
options(datadist=NULL)
if (FALSE) { # \dontrun{
#
# Time to progression/death for patients with monoclonal gammopathy
# Competing risk curves (cumulative incidence)
# status variable must be a factor with first level denoting right censoring
m <- upData(mgus1, stop = stop / 365.25, units=c(stop='years'),
labels=c(stop='Follow-up Time'), subset=start == 0)
f <- npsurv(Surv(stop, event) ~ 1, data=m)
# Use survplot for enhanced displays of cumulative incidence curves for
# competing risks
survplot(f, state='pcm', n.risk=TRUE, xlim=c(0, 20), ylim=c(0, .5), col=2)
survplot(f, state='death', aehaz=TRUE, col=3,
label.curves=list(keys='lines'))
f <- npsurv(Surv(stop, event) ~ sex, data=m)
survplot(f, state='death', aehaz=TRUE, n.risk=TRUE, conf='diffbands',
label.curves=list(keys='lines'))
# Plot survival curves estimated from an ordinal semiparametric model
f <- orm(Ocens(y, ifelse(y <= cens, y, Inf)) ~ age)
survplot(f, age=c(30, 50))
} # }