The Relaxed Lasso
Trevor Hastie
Balasubramanian Narasimhan
Rob Tibshirani
July 02, 2025
relax.RmdIntroduction
In this vignette, we describe how the glmnet package can
be used to fit the relaxed lasso.
The idea of the relaxed lasso is to take a glmnet fitted
object, and then for each lambda, refit the variables in the active set
without any penalization. This gives the “relaxed” fit. (We note that
there have been other definitions of a relaxed fit, but this is the one
we prefer.) This could of course be done for elastic net fits as well as
lasso. However, if the number of variables gets too close to the sample
size
,
the relaxed path will be truncated. Furthermore, for binomial and other
nonlinear generalized linear models (GLMs) convergence can be an issue
with our current implementation if the number of variables is too large,
and perversely if the relaxed fit is too strong.
Suppose the glmnet fitted linear predictor at
is
and the relaxed version is
.
We also allow for shrinkage between the two:
is an additional tuning parameter which can be selected by cross-validation (CV). The debiasing will potentially improve prediction performance, and CV will typically select a model with a smaller number of variables.
This procedure is very competitive with forward-stepwise and
best-subset regression, and has a considerable speed advantage when the
number of variables is large. This is especially true for best-subset,
but even so for forward stepwise. The latter has to plod through the
variables one-at-a-time, while glmnet will just plunge in
and find a good active set.
Further details on this form of relaxed fitting can be found in Hastie, Tibshirani, and Tibshirani (2017); more information on glmnet and elastic-net model in general is given in Friedman, Hastie, and Tibshirani (2010), Simon et al. (2011), Tibshirani et al. (2012), and Simon, Friedman, and Hastie (2013).
Simple relaxed fitting
We demonstrate the most basic relaxed lasso fit as a first example.
We load some pre-generated data and fit the relaxed lasso on it by
calling glmnet with relax = TRUE:
## Loading required package: Matrix## Loaded glmnet 4.1-9
data(QuickStartExample)
x <- QuickStartExample$x
y <- QuickStartExample$y
fit <- glmnet(x, y, relax = TRUE)
print(fit)##
## Call: glmnet(x = x, y = y, relax = TRUE)
## Relaxed
##
## Df %Dev %Dev R Lambda
## 1 0 0.00 0.00 1.63100
## 2 2 5.53 58.90 1.48600
## 3 2 14.59 58.90 1.35400
## 4 2 22.11 58.90 1.23400
## 5 2 28.36 58.90 1.12400
## 6 2 33.54 58.90 1.02400
## 7 4 39.04 76.56 0.93320
## 8 5 45.60 80.59 0.85030
## 9 5 51.54 80.59 0.77470
## 10 6 57.35 87.99 0.70590
....In addition to the three columns usually printed for
glmnet objects (Df, %Dev and
Lambda), there is an extra column %Dev R
(R stands for “relaxed”) which is the percent deviance
explained by the relaxed fit. This is always higher than its neighboring
column, which is the percent deviance exaplined for the penalized fit
(on the training data). Notice that when the Df stays the
same, the %Dev R does not change, since this typically
means the active set is the same. (The code is also smart enough to only
fit such models once, so in the truncated display shown, 9 lasso models
are fit, but only 4 relaxed fits are computed).
The fit object is of class "relaxed", which inherits
from class "glmnet". Hence, the usual plot
method for "glmnet" objects can be used. The code below
demonstrates some additional flexibility that "relaxed"
objects have for plotting.
par(mfrow = c(1, 3), mar=c(4,4,5.5,1))
plot(fit, main = "gamma = 1")
plot(fit, gamma = 0.5, main = "gamma = 0.5")
plot(fit, gamma = 0, main = "gamma = 0")
gamma = 1 is the traditional glmnet fit
(also relax = FALSE, the default), gamma = 0
is the unpenalized fit, and gamma = 0.5 is a mixture of the
two (at the coefficient level, and hence also the linear
predictors).
We can also select gamma using cv.glmnet,
which by default uses the 5 values
c(0, 0.25, 0.5, 0.75, 1). This returns an object of class
"cv.relaxed".
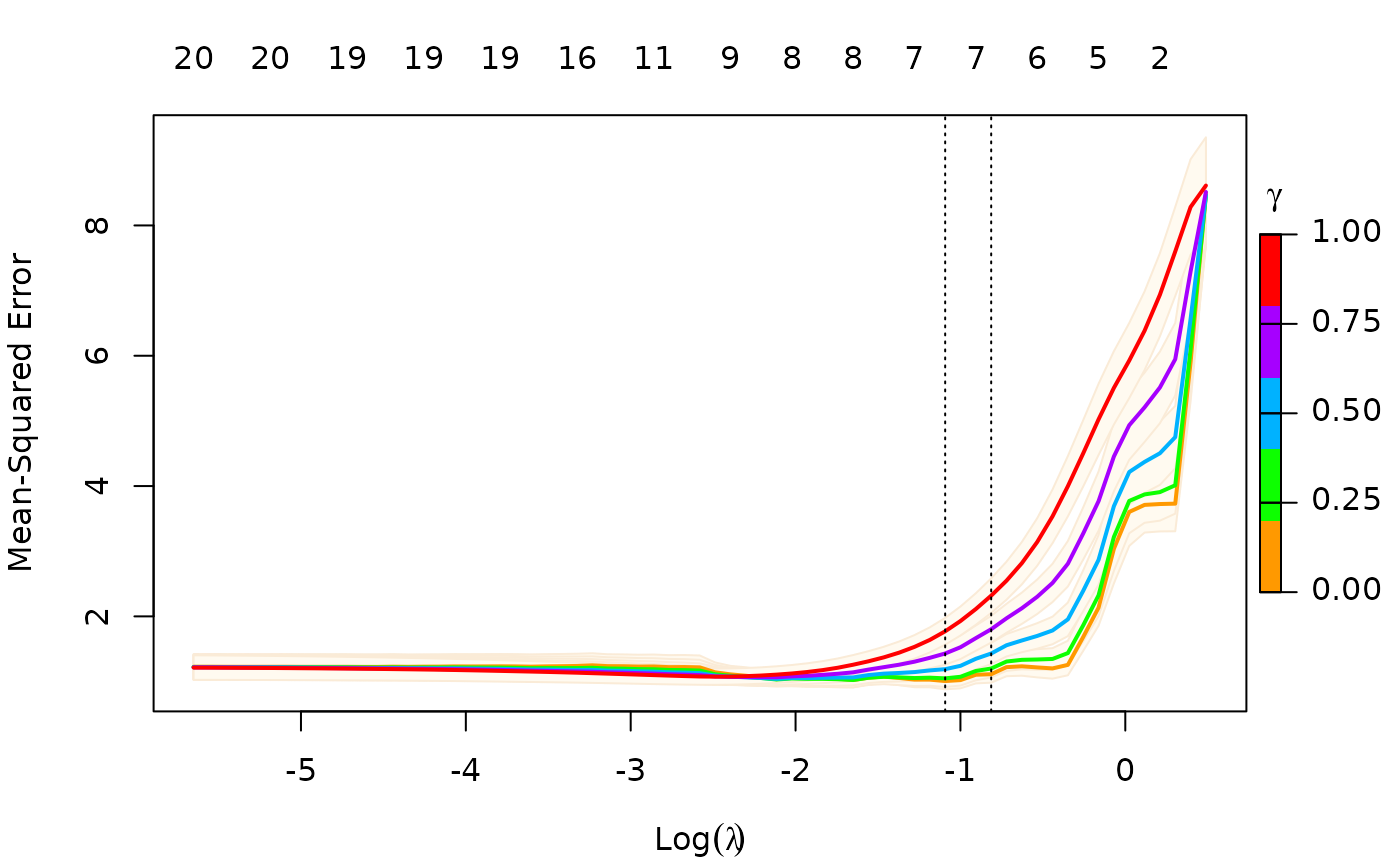
To remove the shading of the standard error bands, pass
se.bands = FALSE:
plot(cfit, se.bands = FALSE)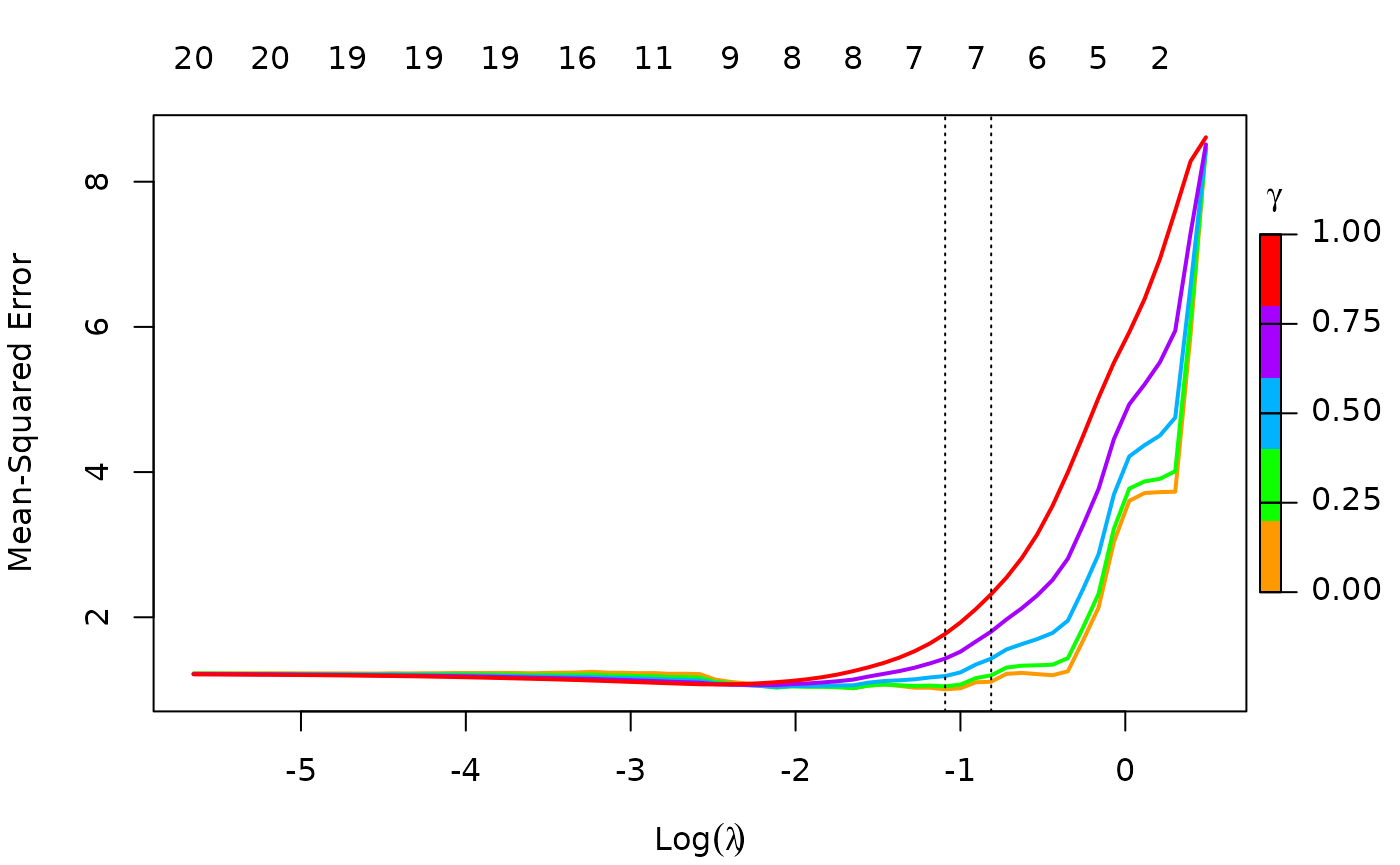
As with regular "cv.glmnet" objects, you can make
predictions from a relaxed CV object. Just as the s option
(for lambda) admits two special strings
"lambda.1se" and "lambda.min" for special
values of lambda, the gamma option admits two
special strings "gamma.1se" and "gamma.min"
for special values of gamma. For example, the code below
makes predictions for newx at the lambda and
gamma values that has the smallest CV error:
predict(cfit, newx = x[1:5, ], s = "lambda.min", gamma = "gamma.min")## lambda.min
## [1,] -1.6281344
## [2,] 2.6698376
## [3,] 0.3511622
## [4,] 1.9185110
## [5,] 1.5664349Printing class "cv.relaxed" objects gives some basic
information on the cross-validation:
print(cfit)##
## Call: cv.glmnet(x = x, y = y, relax = TRUE)
##
## Measure: Mean-Squared Error
##
## Gamma Index Lambda Index Measure SE Nonzero
## min 0 1 0.3354 18 1.007 0.1269 7
## 1se 0 1 0.4433 15 1.112 0.1298 7More details on relaxed fitting
While we only demonstrate relaxed fits for the default Gaussian
family, any of the families fit by glmnet can also
be fit with the relaxed option.
Although glmnet has a relax option, you can
also fit relaxed lasso models by post-processing a glmnet
object with the relax.glmnet function.
fit <- glmnet(x,y)
fitr <- relax.glmnet(fit, x = x, y = y)This will rarely need to be done; one use case is if the original fit
took a long time, and the user wants to avoid refitting it. Note that
the arguments are named in the call in order for them to be passed
correctly via the ... argument in
relax.glmnet.
As mentioned, a "relaxed" object inherits from class
"glmnet". Apart from the class modification, it has an
additional component named relaxed which is itself a
glmnet object, but with the relaxed coefficients. The
default behavior of extractor functions like predict and
coef, as well as plot will be to present
results from the glmnet fit, unless a value of
gamma is given different from the default value
gamma = 1 (see the plots above). The print
method gives additional info on the relaxed fit.
Likewise, a cv.relaxed object inherits from class
cv.glmnet. Here the predict method by default
uses the optimal relaxed fit; if predictions from the CV-optimal
original glmnet fit are desired, one can directly
use predict.cv.glmnet. Similarly, use print to
print information for cross-validation on the relaxed fit, and
print.cv.glmnet for information on the cross-validation for
the original glmnet fit.
print(cfit)##
## Call: cv.glmnet(x = x, y = y, relax = TRUE)
##
## Measure: Mean-Squared Error
##
## Gamma Index Lambda Index Measure SE Nonzero
## min 0 1 0.3354 18 1.007 0.1269 7
## 1se 0 1 0.4433 15 1.112 0.1298 7
print.cv.glmnet(cfit)##
## Call: cv.glmnet(x = x, y = y, relax = TRUE)
##
## Measure: Mean-Squared Error
##
## Lambda Index Measure SE Nonzero
## min 0.08307 33 1.075 0.1251 9
## 1se 0.15933 26 1.175 0.1374 8Possible convergence issues for relaxed fits
glmnet itself is used to fit the relaxed fits by using a
single value of zero for lambda. However, for nonlinear
models such as family = "binomial",
family = "multinomial" and family="poisson",
there can be convergence issues. This is because glmnet
does not do step size optimization, rather relying on the pathwise fit
to stay in the “quadratic” zone of the log-likelihood. We have an
optional path = TRUE option for relax.glmnet,
which actually fits a regurized path toward the lambda = 0
solution, and thus avoids the issue. The default is
path = FALSE since this option adds to the computing
time.
Application to forward stepwise regression
One use case for a relaxed fit is as a faster version of forward
stepwise regression. With a large number p of variables,
forward stepwise regression can be tedious. On the other hand, because
the lasso solves a convex problem, it can plunge in and identify good
candidate sets of variables over 100 values of lambda, even
though p could be in the tens of thousands. In a case like
this, one can have cv.glmnet do the selection of
variables.
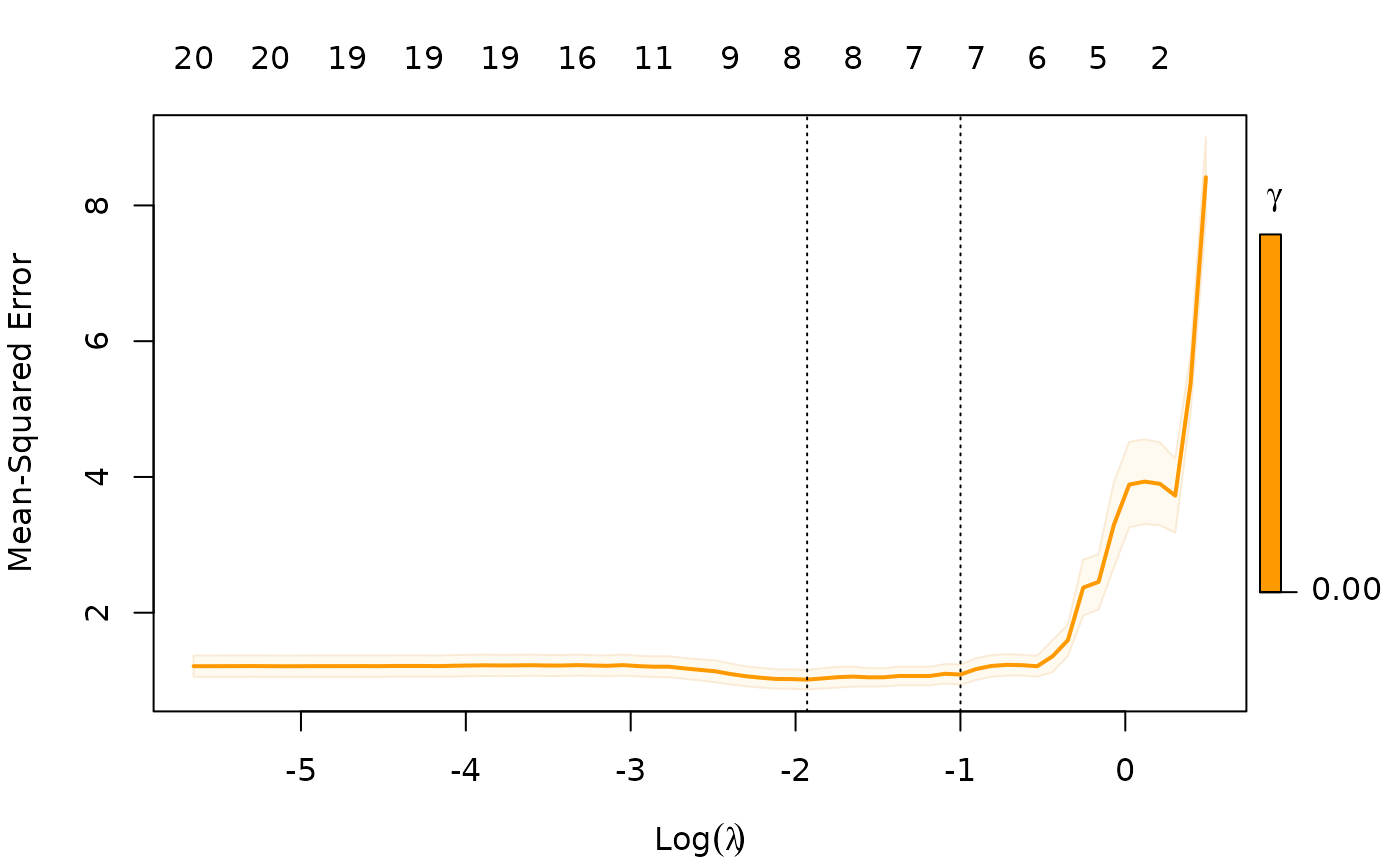
Notice that we only allow gamma = 0, so in this case we
are not considering the blended fits.