Motivation
While glmmTMB is primarily designed for maximum
likelihood estimation (or restricted ML), there are certain situations
where it is convenient to be able to add priors for particular
parameters or sets of parameters, e.g.:
- to mitigate complete separation (technically, when there is some linear combination of parameters that divides zero from non-zero responses in a count or binomial model; in practice, typically when one treatment combination has all-zero responses)
- to mitigate singular fits in random effects, i.e. when there is insufficient data to estimate a variance parameter so that single variances collapse to zero or covariance matrices are estimated with less than full rank (Chung et al. 2013)
- to apply a ridge penalty to a set of parameters (corresponding to an independent Gaussian prior on each parameter)
- more generally, to regularize models that would otherwise be numerically unstable
- for models that will be used with the
tmbstanpackage as part of a fully Bayesian analysis (see below)
See Banner, Irvine, and Rodhouse (2020) and Sarma and Kay (2020) for some opinions/discussion of priors.
When priors are specified, glmmTMB will fit a
maximum a posteriori (MAP) estimate. In other words, unlike
most Bayesian estimate procedures that use Markov chain Monte Carlo to
sample the entire parameter space and compute (typically) posterior mean
or median value of the parameters, glmmTMB will find the
mode of the posterior distribution or the most likely
value. The MAP estimate is theoretically less useful than the posterior
mean or median, but is often a useful approximation.
One can apply tmbstan to a fitted glmmTMB
model that specifies priors (see the MCMC
vignette in order to get samples from the posterior distribution as
in a more typical Bayesian analysis.
Load packages
library(glmmTMB)
library(lme4)
library(blme)
library(broom.mixed)
library(purrr)
library(dplyr)
library(ggplot2)
theme_set(theme_bw())
OkIt <- unname(palette.colors(n = 8, palette = "Okabe-Ito"))[-1]Culcita example: near-complete separation
From Bolker (2015), an example where we can regularize nearly complete separation: see the more complete description here.
For comparison, we’ll fit (1) unpenalized/prior-free
glmer and glmmTMB models; (2)
blme::bglmer(), which adds a prior to a glmer
model; (3) glmmTMB with priors.
We read the data and drop one observation that is identified as having an extremely large residual:
cdat <- readRDS(system.file("vignette_data", "culcita.rds", package = "glmmTMB"))
cdatx <- cdat[-20,]Fit glmer, glmmTMB without priors, as well
as a bglmer model with regularizing priors (mean 0, SD 3,
expressed as a 4
4 diagonal covariance matrix with diagonal elements (variances) equal to
9:
form <- predation~ttt + (1 | block)
cmod_glmer <- glmer(form, data = cdatx, family = binomial)
cmod_glmmTMB <- glmmTMB(form, data = cdatx, family = binomial)
cmod_bglmer <- bglmer(form,
data = cdatx, family = binomial,
fixef.prior = normal(cov = diag(9, 4))
)Specify the same priors for glmmTMB: note that we have
to specify regularizing priors for the intercept and the remaining
fixed-effect priors separately
cprior <- data.frame(prior = rep("normal(0,3)", 2),
class = rep("fixef", 2),
coef = c("(Intercept)", ""))
print(cprior)## prior class coef
## 1 normal(0,3) fixef (Intercept)
## 2 normal(0,3) fixef
cmod_glmmTMB_p <- update(cmod_glmmTMB, priors = cprior)Check (approximate) equality of estimated coefficients:
## comment out for now ...
## stopifnot(all.equal(coef(summary(cmod_bglmer)),
## coef(summary(cmod_glmmTMB_p))$cond,
## tolerance = 5e-2))Pack the models into a list and get the coefficients:
cmods <- ls(pattern = "cmod_[bg].*")
cmod_list <- mget(cmods) |> setNames(gsub("cmod_", "", cmods))
cres <- (purrr::map_dfr(cmod_list,
~ tidy(., conf.int = TRUE, effects = "fixed"),
.id = "model"
)
|> select(model, term, estimate, lwr = conf.low, upr = conf.high)
|> mutate(across(
model,
~ factor(., levels = c(
"glmer", "glmmTMB",
"glmmTMB_p", "bglmer"
))
))
)
ggplot(cres, aes(x = estimate, y = term, colour = model)) +
geom_pointrange(aes(xmin = lwr, xmax = upr),
position = position_dodge(width = 0.5)
) +
scale_colour_manual(values = OkIt)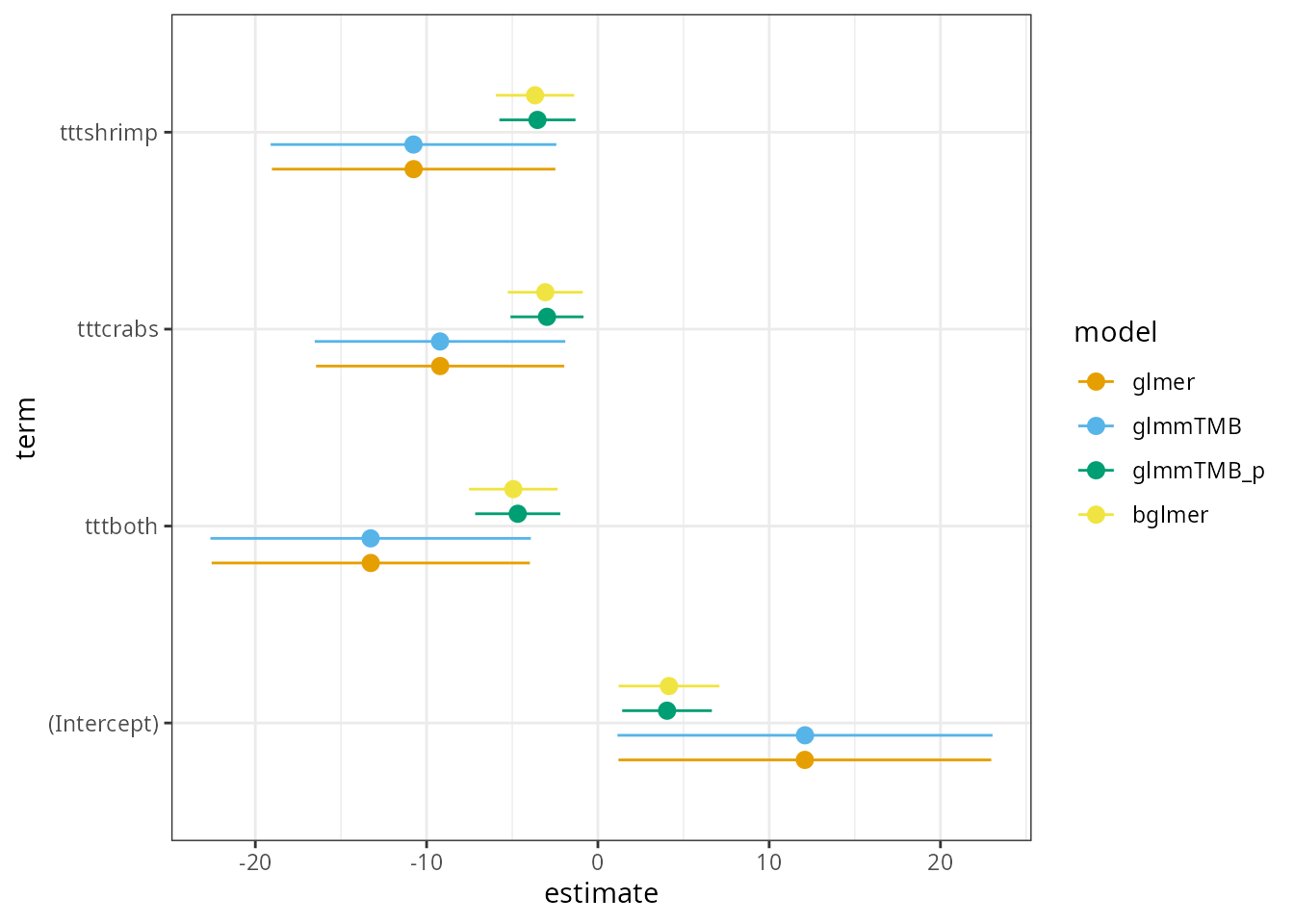
Gopher tortoise example: mitigate singular fit
Also from Bolker (2015):
gdat <- readRDS(system.file("vignette_data", "gophertortoise.rds", package = "glmmTMB"))
form <- shells~prev + offset(log(Area)) + factor(year) + (1 | Site)
gmod_glmer <- glmer(form, family = poisson, data = gdat)## boundary (singular) fit: see help('isSingular')
gmod_bglmer <- bglmer(form, family = poisson, data = gdat)
## cov.prior = gamma(shape = 2.5, rate = 0, common.scale = TRUE, posterior.scale = "sd"))
gmod_glmmTMB <- glmmTMB(form, family = poisson, data = gdat) ## 1e-5
## bglmer default corresponds to gamma(Inf, 2.5)
gprior <- data.frame(prior = "gamma(1e8, 2.5)",
class = "ranef",
coef = "")
gmod_glmmTMB_p <- update(gmod_glmmTMB, priors = gprior)
vc1 <- c(VarCorr(gmod_glmmTMB_p)$cond$Site)
vc2 <- c(VarCorr(gmod_bglmer)$Site)
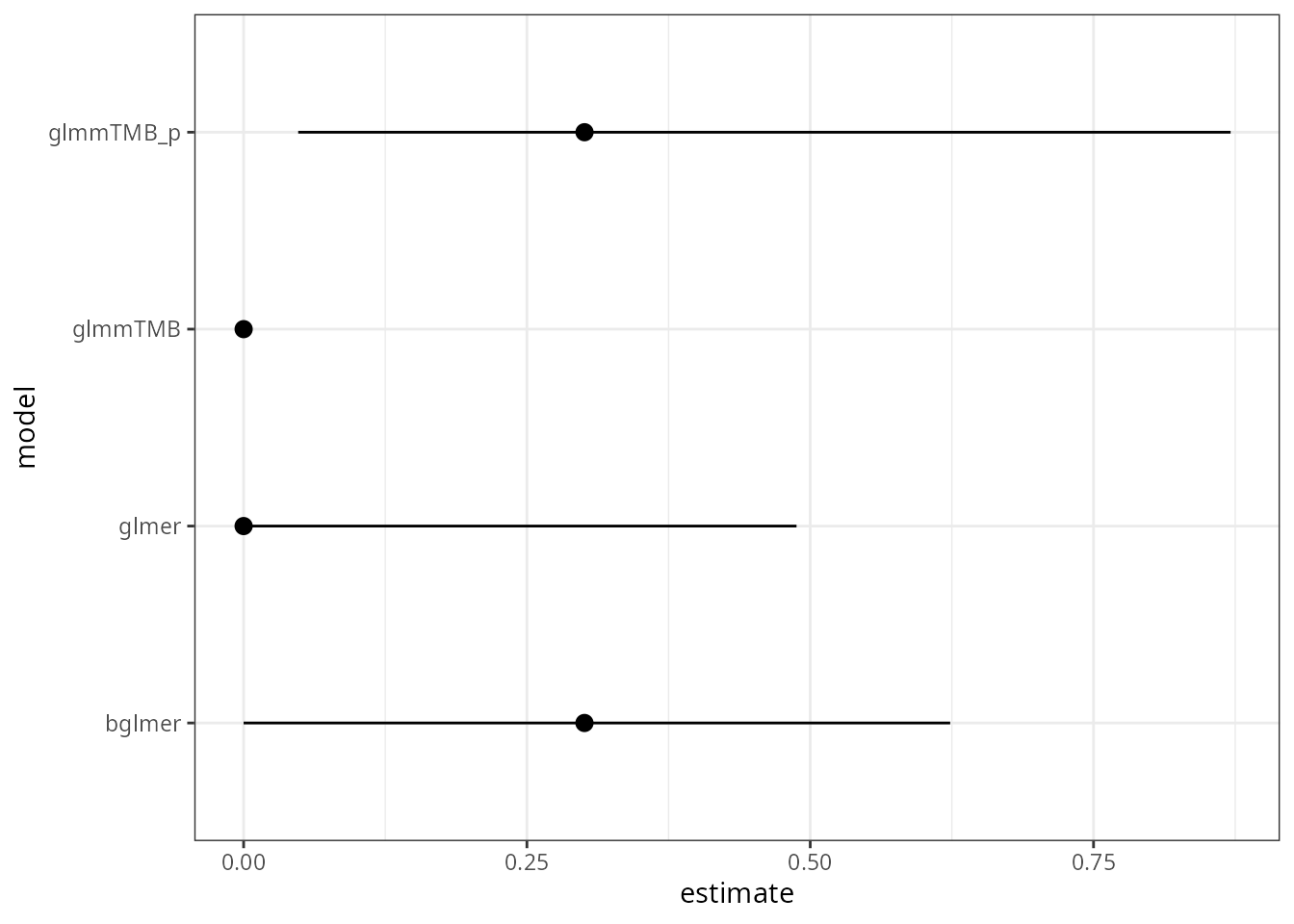
stopifnot(all.equal(vc1, vc2, tolerance = 5e-4))Pack the models into a list and get the coefficients:
The code for extracting CIs is currently a little bit ugly (because
profile confidence intervals aren’t quite working for
glmmTMB objects with broom.mixed::tidy(), and
because profile CIs can be fussy in any case)
blme defaults: Wishart(dim + 2.5), or gamma(2.5). For
dim = 1 (scalar), Wishart(n) corresponds to chi-squared(n), or
gamma(shape = n/2, scale = n/2). Chung et al propose
gamma(2, Inf); not sure why blme uses
gamma(2.5) instead? or if specified via Wishart, shape =
3.5 → gamma shape of 1.75?
TO DO/FIX ME
- try to get internal structure of priors fixed before release,
otherwise
up2datemight get annoying … - document synonyms
- why is
bglmerprofile CI failing (inbroom.mixed, but not externally?) - figure out/document
blmedefault priors - add tests!
- document that gamma is applied on exp() scale
- move prior info to a separate man page?
- implement elementwise priors
- start with specifying by number, do lookup by name later
- allow multivariate (joint) priors on parameter vectors rather than
iid priors?
- esp for correlation matrices: LKJ, Wishart etc. (from Mikael Jagan here)
- add beta priors for zi, corr, etc. ?
- number of prior parameters (save annoying C++ code); can specify via
_coror_sdon the R side (will pick out sd-specific or cor-specific elements) - start and end indices in vector
- number of prior parameters (save annoying C++ code); can specify via
- test!
- safety checks (e.g. error at end of switch statements in C++)
Development issues
It seems useful to use the API/user interface from
brms
- downside:
brmshas lots of downstream dependencies thatglmmTMBdoesn’t- might be able to copy the relevant code (the full file is 2210 lines (!), but this includes documentation and a lot of code we don’t need …
rd <- \(x) tools::package_dependencies("brms", recursive = TRUE)[[x]] ## rd <- \(x) packrat:::recursivePackageDependencies(x, ignores = "", lib.loc = .libPaths()[1]) ## not sure why packrat and tools get different answers, but difference ## doesn't matter much brms_dep <- rd("brms") glmmTMB_dep <- rd("glmmTMB") length(setdiff(brms_dep, glmmTMB_dep)) - at its simplest, this is just a front-end for a data frame
## requires brms to evaluate, wanted to avoid putting it in Suggests: ...
bprior <- c(prior_string("normal(0,10)", class = "b"),
prior(normal(1,2), class = b, coef = treat),
prior_(~cauchy(0,2), class = ~sd,
group = ~subject, coef = ~Intercept))
str(bprior)## Classes 'brmsprior' and 'data.frame': 3 obs. of 10 variables:
## $ prior : chr "normal(0,10)" "normal(1, 2)" "cauchy(0, 2)"
## $ class : chr "b" "b" "sd"
## $ coef : chr "" "treat" "Intercept"
## $ group : chr "" "" "subject"
## $ resp : chr "" "" ""
## $ dpar : chr "" "" ""
## $ nlpar : chr "" "" ""
## $ lb : chr NA NA NA
## $ ub : chr NA NA NA
## $ source: chr "user" "user" "user"We probably only need to pay attention to the columns
prior, class, coef,
group. For our purposes, prior is the name and
parameters; class will be the name of the parameter vector;
coef will specify an index within the vector (could be a
number or name?)
TMB-side data structure:
-
vector of prior codes
- we need a new
enum,.valid_priors: seemake-enumin the Makefile
- we need a new
list of parameter vectors? or
prior_p1,prior_p2,prior_p3(do any prior families have more than two parameters? What about non-scalar parameters, e.g. Wishart priors … ???)vector of parameter codes (another
enum?) (beta,theta,thetaf…b?)each index (corresponding to
coef) is scalar, either NA (prior over all elements) or integer (a specific element)new loop after loglik loop to add (negative log-)prior components: loop over prior spec
add
theta_corr,theta_sdas enum options (synonyms:ranef_corr,ranef_sd) to specify penalizing only SD vector or only corr vector from a particular element?-
‘coef’ picks out elements
- fixed effect: find numeric index in
colnames(X)of corresponding component - random effect: find indices (start and stop?) in corresponding
thetavector -
ranef_corr,ranef_sd: find indices … (depends on RE structure)
- fixed effect: find numeric index in