Influence Index Plot
infIndexPlot.RdProvides index plots of influence and related diagnostics for a regression model.
Usage
infIndexPlot(model, ...)
influenceIndexPlot(model, ...)
# S3 method for class 'lm'
infIndexPlot(model, vars=c("Cook", "Studentized", "Bonf", "hat"),
id=TRUE, grid=TRUE, main="Diagnostic Plots", ...)
# S3 method for class 'influence.merMod'
infIndexPlot(model,
vars = c("dfbeta", "dfbetas", "var.cov.comps",
"cookd"), id = TRUE, grid = TRUE, main = "Diagnostic Plots", ...)
# S3 method for class 'influence.lme'
infIndexPlot(model,
vars = c("dfbeta", "dfbetas", "var.cov.comps",
"cookd"), id = TRUE, grid = TRUE, main = "Diagnostic Plots", ...)Arguments
- model
A regression object of class
lm,glm, orlmerMod, or an influence object for almer,glmer, orlmeobject (seeinfluence.mixed.models). The"lmerMod"method calls the"lm"method and can take the same arguments.- vars
All the quantities listed in this argument are plotted. Use
"Cook"for Cook's distances,"Studentized"for Studentized residuals,"Bonf"for Bonferroni p-values for an outlier test, and and"hat"for hat-values (or leverages) for a linear or generalized linear model, or"dfbeta","dfbetas","var.cov.comps", and"cookd"for an influence object derived from a mixed model. Capitalization is optional. All but"dfbeta"and"dfbetas"may be abbreviated by the first one or more letters.- main
main title for graph
- id
a list of named values controlling point labelling. The default,
TRUE, is equivalent toid=list(method="y", n=2, cex=1, col=carPalette()[1], location="lr");FALSEsuppresses point labelling. SeeshowLabelsfor details.- grid
If TRUE, the default, a light-gray background grid is put on the graph.
- ...
Arguments passed to
plot
References
Cook, R. D. and Weisberg, S. (1999) Applied Regression, Including Computing and Graphics. Wiley.
Fox, J. (2016) Applied Regression Analysis and Generalized Linear Models, Third Edition. Sage. Fox, J. and Weisberg, S. (2019) An R Companion to Applied Regression, Third Edition, Sage.
Weisberg, S. (2014) Applied Linear Regression, Fourth Edition, Wiley.
Author
Sanford Weisberg sandy@umn.edu and John Fox
Examples
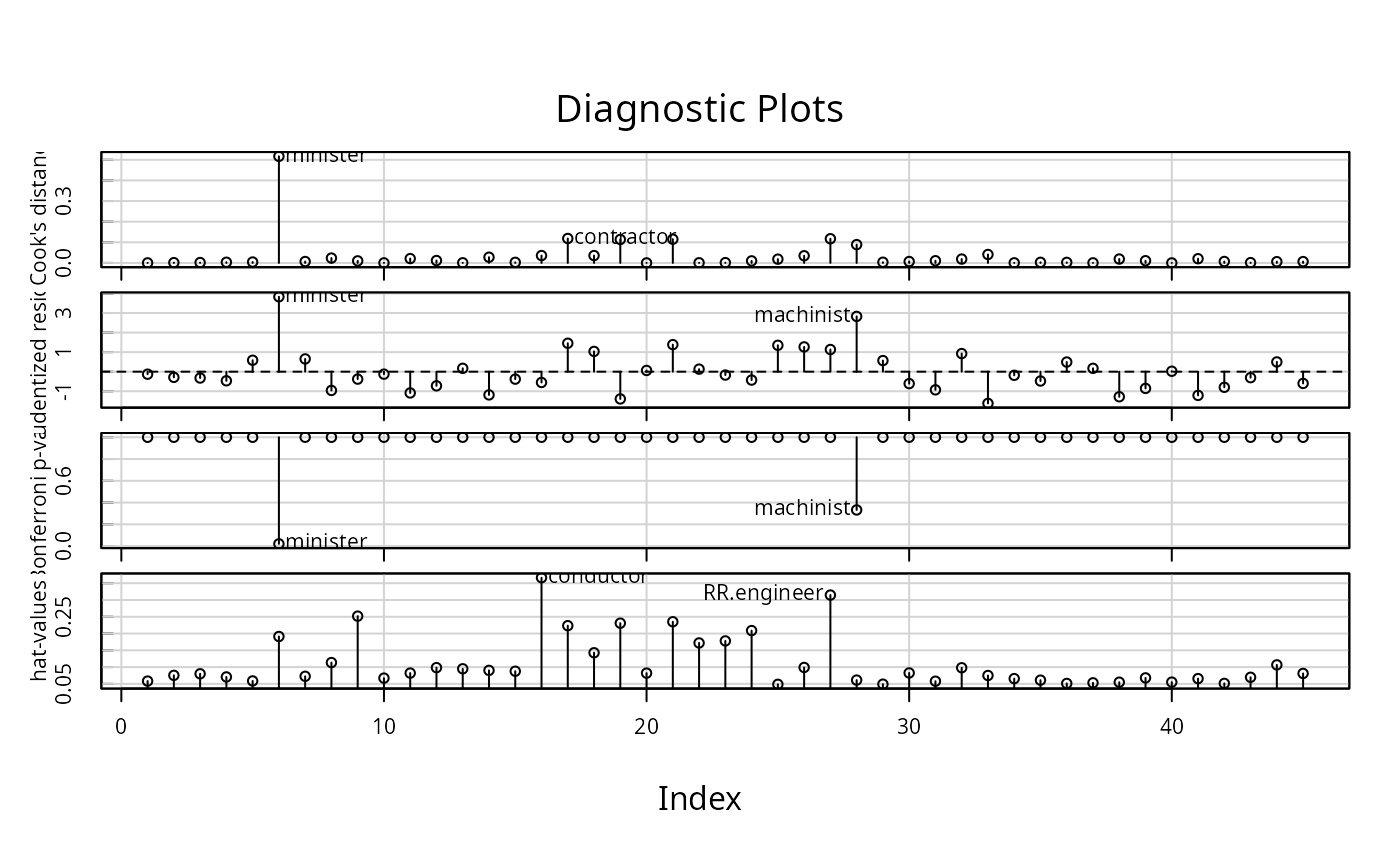
influenceIndexPlot(lm(prestige ~ income + education + type, Duncan))
 if (FALSE) # a little slow
if (require(lme4)){
print(fm1 <- lmer(Reaction ~ Days + (Days | Subject),
sleepstudy)) # from ?lmer
infIndexPlot(influence(fm1, "Subject"))
infIndexPlot(influence(fm1))
}
if (require(lme4)){
gm1 <- glmer(cbind(incidence, size - incidence) ~ period + (1 | herd),
data = cbpp, family = binomial) # from ?glmer
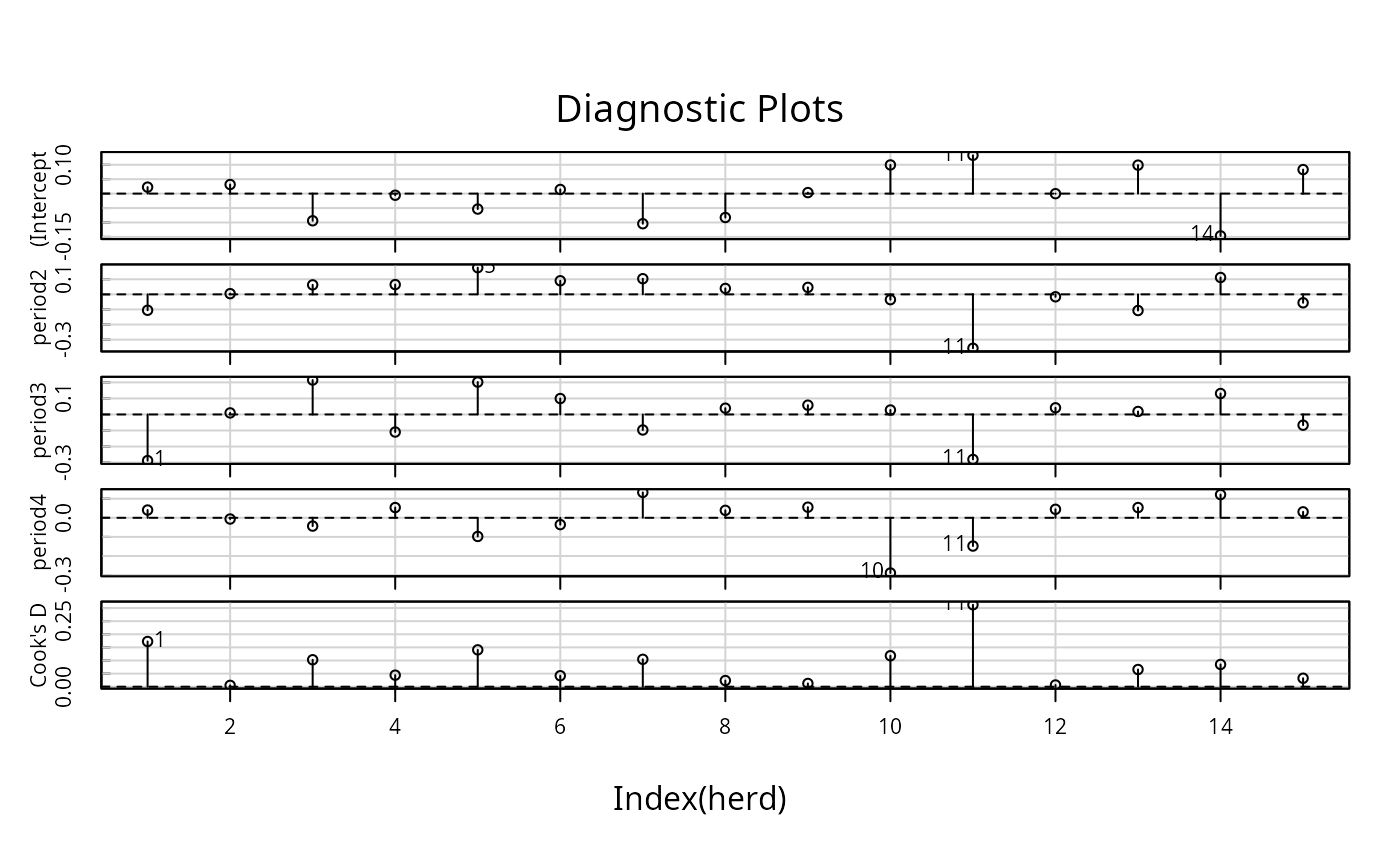
infIndexPlot(influence(gm1, "herd", maxfun=100))
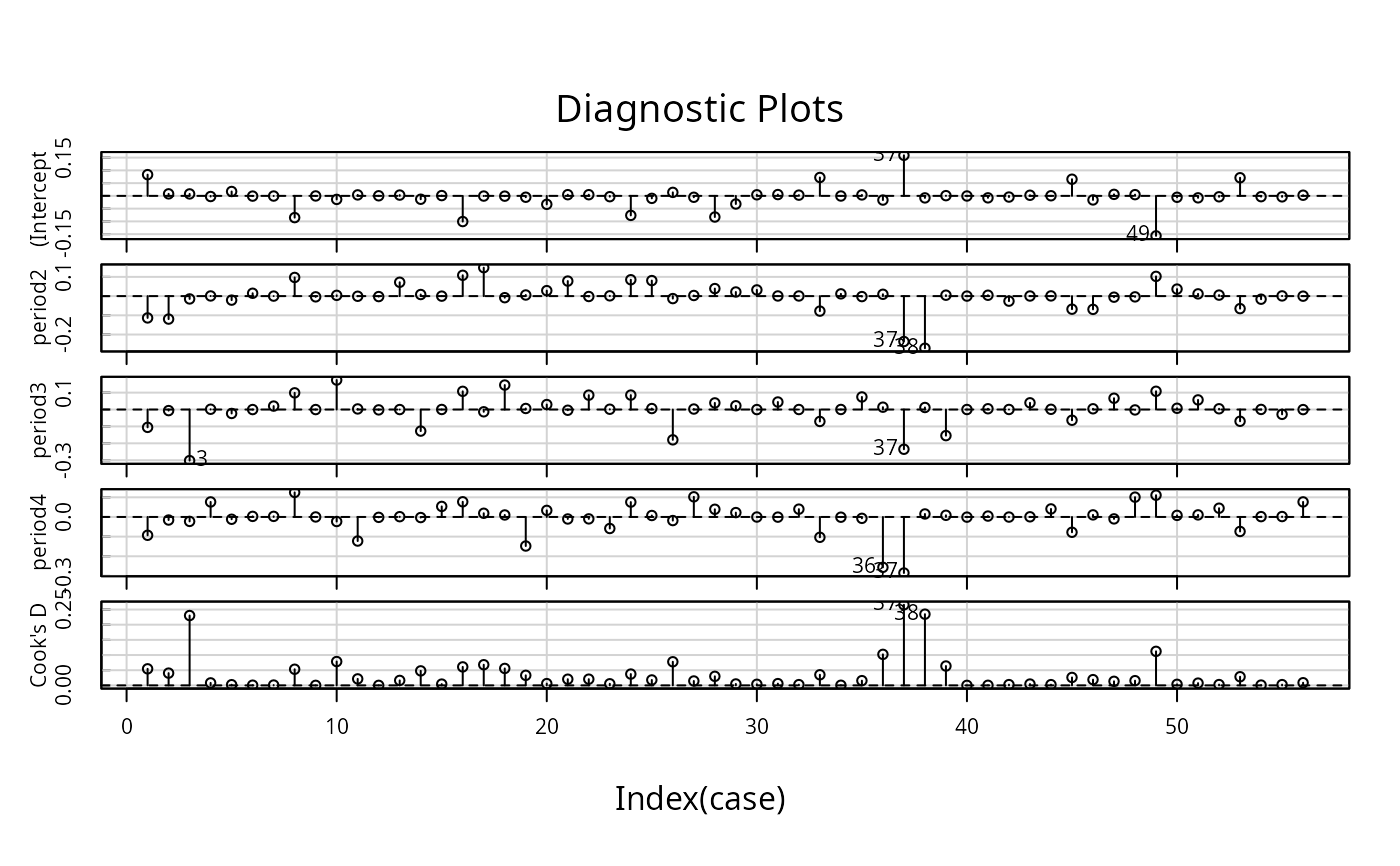
infIndexPlot(influence(gm1, maxfun=100))
gm1.11 <- update(gm1, subset = herd != 11) # check deleting herd 11
compareCoefs(gm1, gm1.11)
}
#> Loading required package: lme4
#> Loading required package: Matrix
#>
#> Attaching package: ‘lme4’
#> The following object is masked from ‘package:rio’:
#>
#> factorize
if (FALSE) # a little slow
if (require(lme4)){
print(fm1 <- lmer(Reaction ~ Days + (Days | Subject),
sleepstudy)) # from ?lmer
infIndexPlot(influence(fm1, "Subject"))
infIndexPlot(influence(fm1))
}
if (require(lme4)){
gm1 <- glmer(cbind(incidence, size - incidence) ~ period + (1 | herd),
data = cbpp, family = binomial) # from ?glmer
infIndexPlot(influence(gm1, "herd", maxfun=100))
infIndexPlot(influence(gm1, maxfun=100))
gm1.11 <- update(gm1, subset = herd != 11) # check deleting herd 11
compareCoefs(gm1, gm1.11)
}
#> Loading required package: lme4
#> Loading required package: Matrix
#>
#> Attaching package: ‘lme4’
#> The following object is masked from ‘package:rio’:
#>
#> factorize
 #> Calls:
#> 1: glmer(formula = cbind(incidence, size - incidence) ~ period + (1 | herd),
#> data = cbpp, family = binomial)
#> 2: glmer(formula = cbind(incidence, size - incidence) ~ period + (1 | herd),
#> data = cbpp, family = binomial, subset = herd != 11)
#>
#> Model 1 Model 2
#> (Intercept) -1.398 -1.271
#> SE 0.231 0.240
#>
#> period2 -0.992 -1.364
#> SE 0.303 0.343
#>
#> period3 -1.128 -1.399
#> SE 0.323 0.354
#>
#> period4 -1.580 -1.710
#> SE 0.422 0.453
#>
# \dontrun{}
#> Calls:
#> 1: glmer(formula = cbind(incidence, size - incidence) ~ period + (1 | herd),
#> data = cbpp, family = binomial)
#> 2: glmer(formula = cbind(incidence, size - incidence) ~ period + (1 | herd),
#> data = cbpp, family = binomial, subset = herd != 11)
#>
#> Model 1 Model 2
#> (Intercept) -1.398 -1.271
#> SE 0.231 0.240
#>
#> period2 -0.992 -1.364
#> SE 0.303 0.343
#>
#> period3 -1.128 -1.399
#> SE 0.323 0.354
#>
#> period4 -1.580 -1.710
#> SE 0.422 0.453
#>
# \dontrun{}