Influence Diagnostics for Mixed-Effects Models
influence-mixed-models.RdThese functions compute deletion influence diagnostics for linear mixed-effects models fit by lme in the nlme package.
The main function is a method for the influence generic function. Other functions are provided for
computing dfbeta, dfbetas, cooks.distance, and influence on variance-covariance components based
on the object computed by influence.lme.
Usage
# S3 method for class 'lme'
influence(model, groups, data, ncores=1, ...)
# S3 method for class 'influence.lme'
cooks.distance(model, ...)
# S3 method for class 'influence.lme'
dfbeta(model, which = c("fixed", "var.cov"), ...)
# S3 method for class 'influence.lme'
dfbetas(model, ...)Arguments
- model
in the case
influence, a model of class"lme"; in the case ofcooks.distance,dfbeta, ordfbetas, an object returned byinfluence.lme.- groups
a character vector containing the name of a grouping factor or names of grouping factors; if more than one name is supplied, then groups are defined by all combinations of levels of the grouping factors that appear in the data. If omitted, then each individual row of the data matrix is treated as a "group" to be deleted in turn.
- data
an optional data frame with the data to which
modelwas fit;influence.lmecan access the data unlesskeep.data=FALSEwas specified in the call tolme, so it's usually unnecessary to supply thedataargument.- ncores
number of cores for parallel computation of diagnostics; if
1(the default), the computation isn't parallelized; ifInf, all of the available physical cores (not necessarily logical cores — seedetectCores) on the computer will be used.- which
if
"fixed.effects"(the default), return influence on the fixed effects; if"var.cov", return influence on the variance-covariance components.- ...
ignored.
Details
influence.lme starts with the estimated variance-covariance components from model and then refits
the model omitting each group in turn.
The other functions are methods for the dfbeta, dfbetas, and cooks.distance generics, to be applied to the
"influence.lme" object produced by the influence function; the dfbeta methods can also return
influence on the variance-covariance components.
Value
influence.lme returns an object of class
"influence.lme",
which contains the following elements:
"fixed.effects"the estimated fixed effects for the model.
"fixed.effects[-groups]"a matrix with columns corresponding to the fixed-effects coefficients and rows corresponding to groups, giving the estimated fixed effects with each group deleted in turn; groups is formed from the name(s) of the grouping factor(s).
"var.cov.comps"the estimated variance-covariance parameters for the model.
"var.cov.comps[-groups]"a matrix with the estimated covariance parameters (in columns) with each group deleted in turn.
"vcov"The estimated covariance matrix of the fixed-effects coefficients.
"vcov[-groups]"a list each of whose elements is the estimated covariance matrix of the fixed-effects coefficients with one group deleted.
"groups"a character vector giving the names of the grouping factors.
"deleted"the possibly composite grouping factor, each of whose elements is deleted in turn.
For plotting "influence.lme" objects, see infIndexPlot.
References
Fox, J. and Weisberg, S. (2019) An R Companion to Applied Regression, Third Edition, Sage.
Author
J. Fox jfox@mcmaster.ca
Examples
if (require(nlme)){
print(fm1 <- lme(distance ~ age, data = Orthodont))
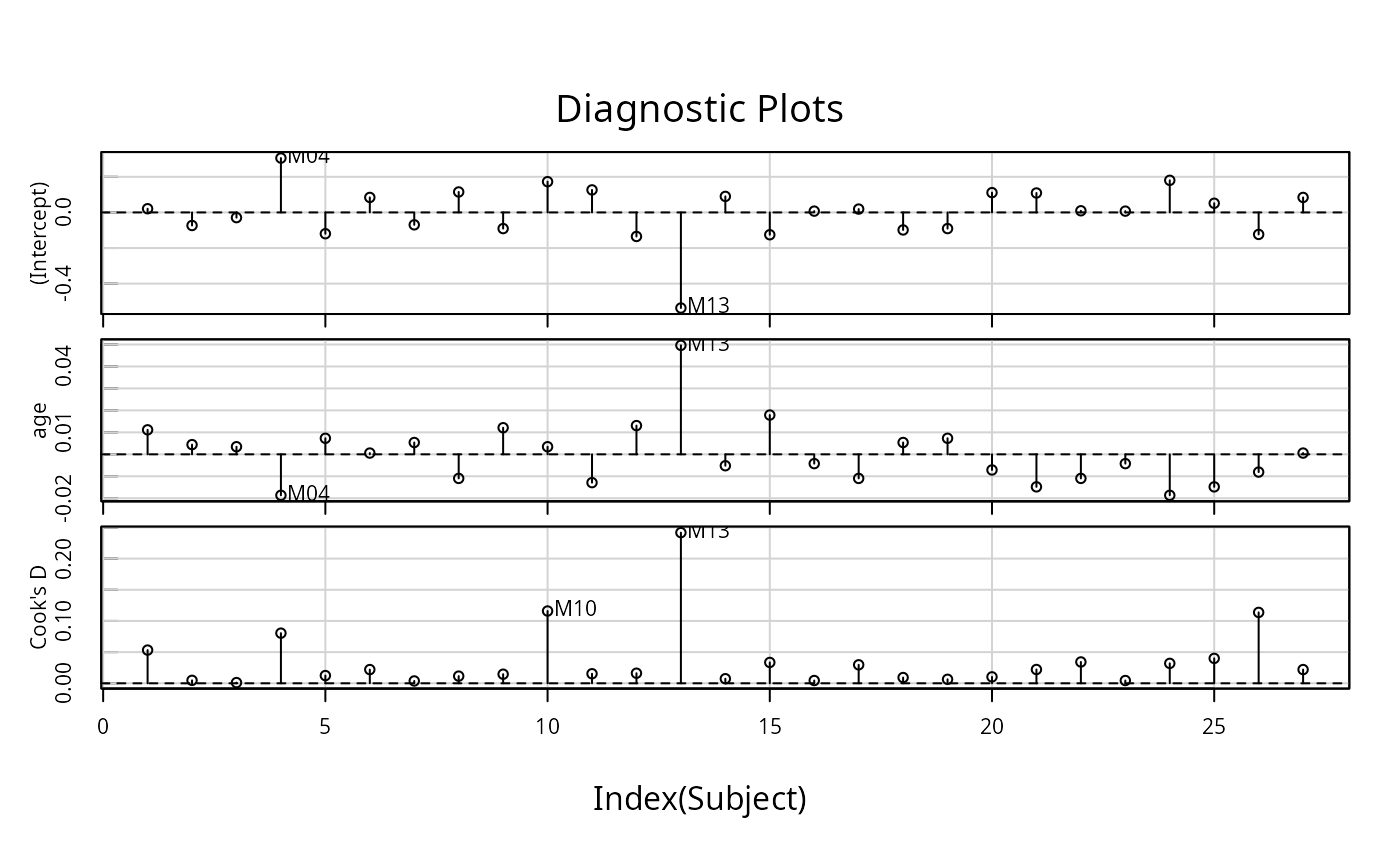
infIndexPlot(influence(fm1, "Subject"))
# a little slow:
if (FALSE) { # \dontrun{
infIndexPlot(influence(fm1))
} # }
}
#> Loading required package: nlme
#>
#> Attaching package: ‘nlme’
#> The following object is masked from ‘package:lme4’:
#>
#> lmList
#> Linear mixed-effects model fit by REML
#> Data: Orthodont
#> Log-restricted-likelihood: -221.3183
#> Fixed: distance ~ age
#> (Intercept) age
#> 16.7611111 0.6601852
#>
#> Random effects:
#> Formula: ~age | Subject
#> Structure: General positive-definite
#> StdDev Corr
#> (Intercept) 2.3270339 (Intr)
#> age 0.2264276 -0.609
#> Residual 1.3100399
#>
#> Number of Observations: 108
#> Number of Groups: 27