For each subject cardiac output is measured repeatedly (three to six times) by impedance cardiography (IC) and radionuclide ventriculography (RV).
Format
A data frame with 120 observations on the following 4 variables.
metha factor with levels
ICRVitema numeric vector giving the item number.
repla numeric vector with replicate number.
ythe measuremnts of cardiac output.
Source
The dataset is adapted from table 4 in: JM Bland and DG Altman: Measuring agreement in method comparison studies. Statistical Methods in Medical Research, 8:136-160, 1999. Originally supplied to Bland & Altman by Dr LS Bowling, see: Bowling LS, Sageman WS, O'Connor SM, Cole R, Amundson DE. Lack of agreement between measurement of ejection fraction by impedance cardiography versus radionuclide ventriculography. Critical Care Medicine 1993; 21: 1523-27.
Details
It is not entirely clear from the source whether the replicates are exchangeable within (method,item) or whether they represent pairs of measurements. From the description it looks as if replicates are linked between methods, but in the paper they are treated as if they were not.
Examples
data(cardiac)
cardiac <- Meth(cardiac)
#> The following variables from the dataframe
#> "cardiac" are used as the Meth variables:
#> meth: meth
#> item: item
#> repl: repl
#> y: y
#> #Replicates
#> Method 3 4 5 6 #Items #Obs: 120 Values: min med max
#> IC 1 3 3 5 12 60 2.32 4.610 7.40
#> RV 1 3 3 5 12 60 2.85 5.105 7.89
summary(cardiac)
#> #Replicates
#> Method 3 4 5 6 #Items #Obs: 120 Values: min med max
#> IC 1 3 3 5 12 60 2.32 4.610 7.40
#> RV 1 3 3 5 12 60 2.85 5.105 7.89
# Visually check exchangeability
plot( cardiac )
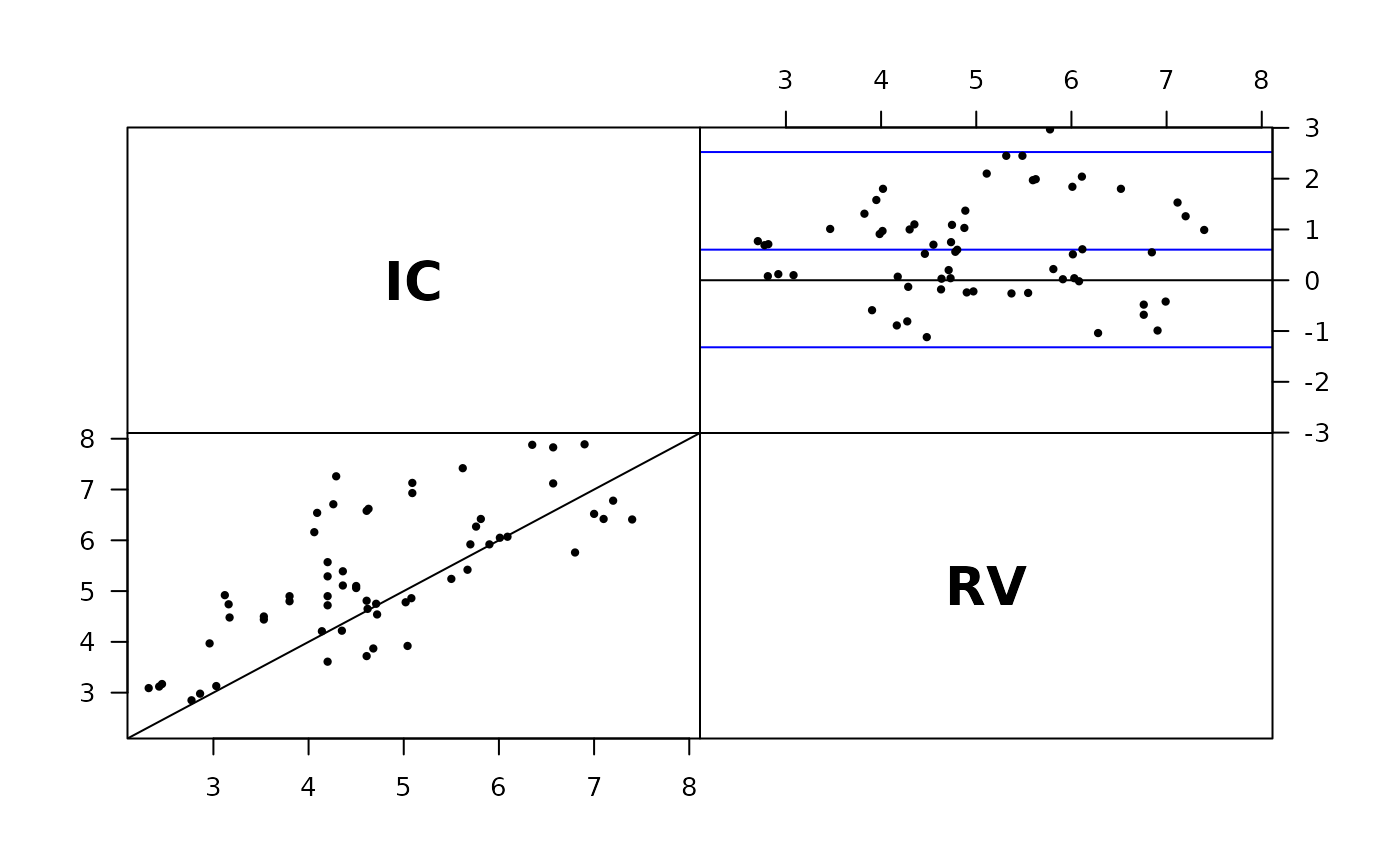 plot( perm.repl( cardiac ) )
plot( perm.repl( cardiac ) )
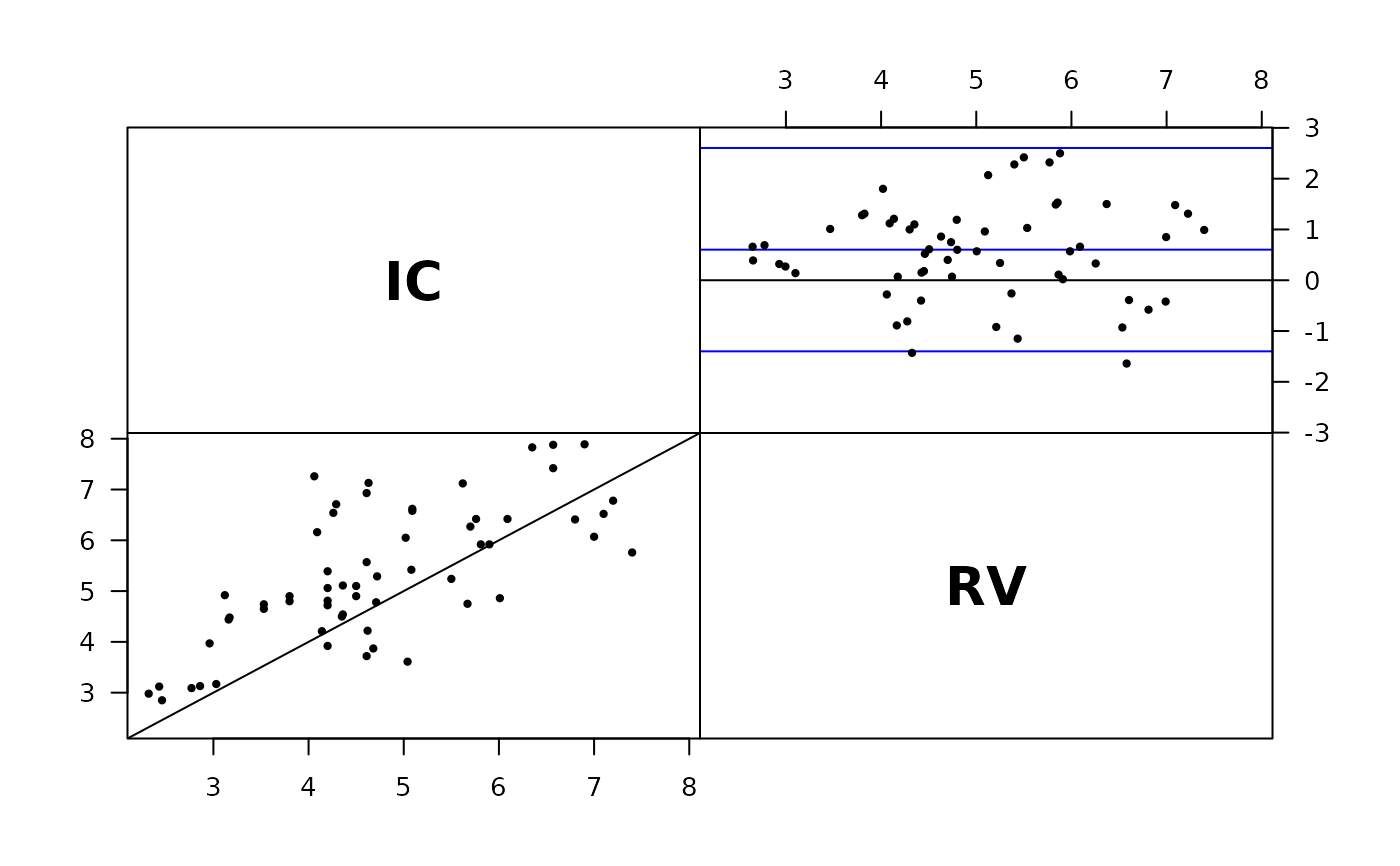 BA.est(cardiac)
#>
#> Conversion between methods:
#> alpha beta sd.pred LoA-lo LoA-up
#> To: From:
#> IC IC 0.000 1.000 0.449 -0.898 0.898
#> RV -0.705 1.000 1.022 -2.748 1.339
#> RV IC 0.705 1.000 1.022 -1.339 2.748
#> RV 0.000 1.000 0.374 -0.749 0.749
#>
#> Variance components (sd):
#> IxR MxI res
#> IC 0.193 0.661 0.317
#> RV 0.193 0.661 0.265
# Run MCmcmc using BRugs for an insufficient amount of iterations
if (FALSE) card.mi.ir <- MCmcmc( cardiac,
beta=FALSE, random=c("mi","ir"),
n.iter=100, trace=T )
print( card.mi.ir ) # \dontrun{}
#> Error: object 'card.mi.ir' not found
BA.est(cardiac)
#>
#> Conversion between methods:
#> alpha beta sd.pred LoA-lo LoA-up
#> To: From:
#> IC IC 0.000 1.000 0.449 -0.898 0.898
#> RV -0.705 1.000 1.022 -2.748 1.339
#> RV IC 0.705 1.000 1.022 -1.339 2.748
#> RV 0.000 1.000 0.374 -0.749 0.749
#>
#> Variance components (sd):
#> IxR MxI res
#> IC 0.193 0.661 0.317
#> RV 0.193 0.661 0.265
# Run MCmcmc using BRugs for an insufficient amount of iterations
if (FALSE) card.mi.ir <- MCmcmc( cardiac,
beta=FALSE, random=c("mi","ir"),
n.iter=100, trace=T )
print( card.mi.ir ) # \dontrun{}
#> Error: object 'card.mi.ir' not found