Creates a dataframe with columns meth, item, (repl) and y.
Arguments
- data
A data frame
- meth
Vector of methods, numeric, character or factor. Can also be a number or character referring to a column in
data.- item
Vector of items, numeric, character or factor. Can also be a number or character referring to a column in
data.- repl
Vector of replicates, numeric, character or factor. Can also be a number or character referring to a column in
data.- y
Vector of measurements. Can also be a character or numerical vector pointing to columns in
datawhich contains the measurements by different methods or a dataframe with columns representing measurements by different methods. In this case the argumentmethis ignored, and the names of the columns are taken as method names.Logical: Should a summary result be printed?
- keep.vars
Logical. Should the remaining variables from the dataframe
databe transferred to theMethobject.
Value
The Meth function returns a Meth object which is a dataframe with columns meth, item, (repl) and y. summary.Meth returns a table classified by method and no. of replicate measurements, extended with columns of the total number of items, total number of observations and the range of the measurements.
Details
In order to perform analyses of method comparisons it is convenient to have a
dataframe with classifying factors, meth, item, and possibly
repl and the response variable y. This function creates such a
dataframe, and gives it a class, Meth, for which there is a number of
methods: summary - tabulation, plot - plotting and a couple of
analysis methods.
If there are replicates in the values of item it is assumed
that those observations represent replicate measurements and different
replicate numbers are given to those.
Examples
data(fat)
# Different ways of selecting columns and generating replicate numbers
Sub1 <- Meth(fat,meth=2,item=1,repl=3,y=4,print=TRUE)
#> The following variables from the dataframe
#> "fat" are used as the Meth variables:
#> meth: Obs
#> item: Id
#> repl: Rep
#> y: Sub
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
Sub2 <- Meth(fat,2,1,3,4,print=TRUE)
#> The following variables from the dataframe
#> "fat" are used as the Meth variables:
#> meth: Obs
#> item: Id
#> repl: Rep
#> y: Sub
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
Sub3 <- Meth(fat,meth="Obs",item="Id",repl="Rep",y="Sub",print=TRUE)
#> The following variables from the dataframe
#> "fat" are used as the Meth variables:
#> meth: Obs
#> item: Id
#> repl: Rep
#> y: Sub
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
summary( Sub3 )
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
plot( Sub3 )
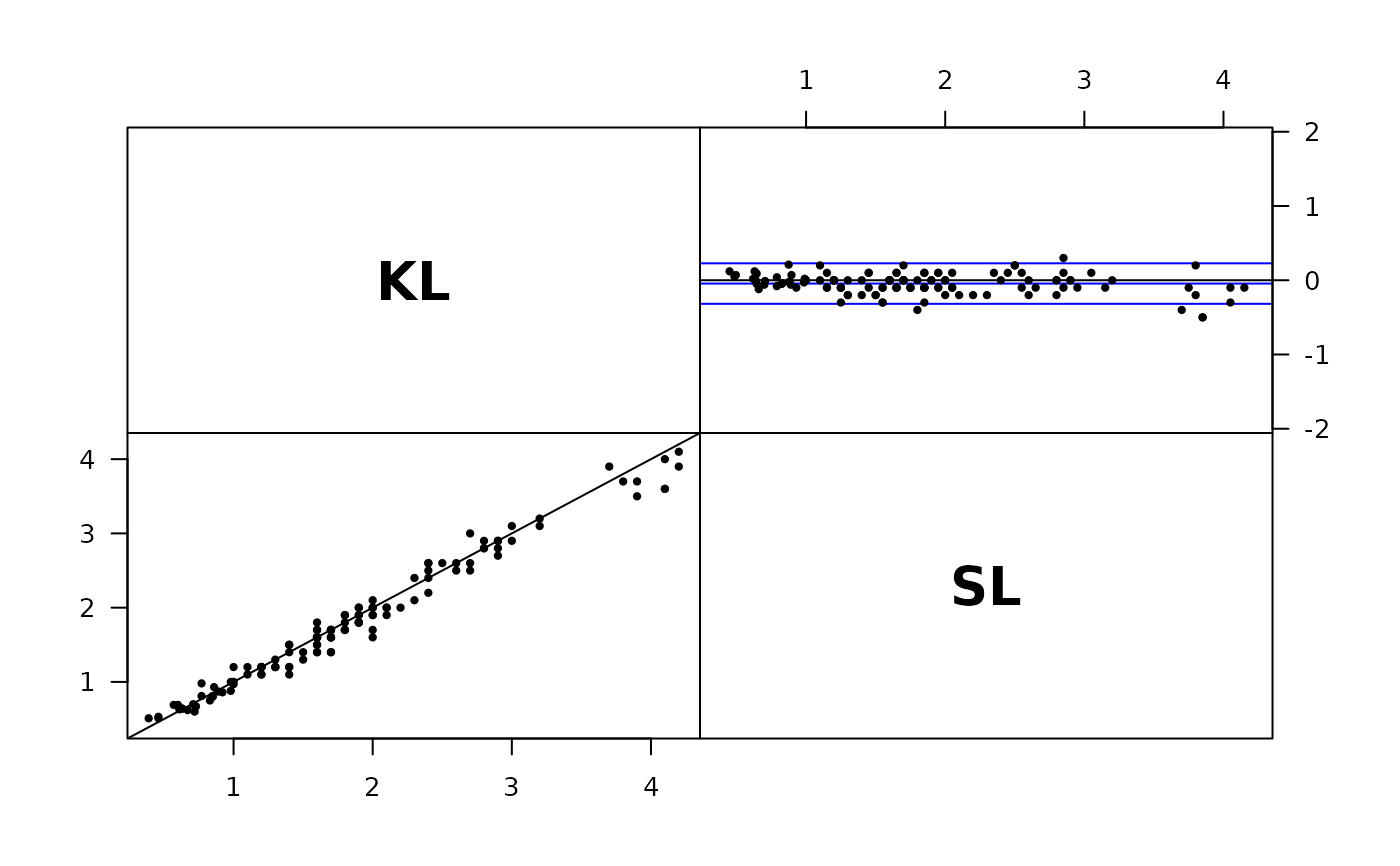 # Use observation in different columns as methods
data( CardOutput )
head( CardOutput )
#> Age Diag VO2 Svo2 Scvo2 TCO FCO Sex
#> 1 66 sepsis 207 63 62 4.9 5.3 M
#> 2 73 sepsis 300 75 78 7.4 10.2 M
#> 3 53 sepsis 289 69 70 8.1 8.7 F
#> 4 28 cardiogenic 150 70 71 3.1 3.7 F
#> 5 72 cardiogenic 350 66 68 5.7 7.6 M
#> 6 71 sepsis 150 70 72 3.1 3.7 M
sv <- Meth( CardOutput, y=c("Svo2","Scvo2") )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 #Items #Obs: 30 Values: min med max
#> Scvo2 15 15 15 61 71 84
#> Svo2 15 15 15 62 70 82
# Note that replicates are generated if a non-unique item-id is used
sv <- Meth( CardOutput, y=c("Svo2","Scvo2"), item="Age" )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> item: Age
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 2 3 #Items #Obs: 30 Values: min med max
#> Scvo2 8 2 1 11 15 61 71 84
#> Svo2 8 2 1 11 15 62 70 82
str( sv )
#> Classes ‘Meth’ and 'data.frame': 30 obs. of 9 variables:
#> $ meth: Factor w/ 2 levels "Scvo2","Svo2": 2 2 2 2 2 2 2 2 2 2 ...
#> $ item: Factor w/ 11 levels "24","28","53",..: 7 10 3 2 9 8 8 3 5 11 ...
#> $ repl: Factor w/ 3 levels "1","2","3": 1 1 1 1 1 1 2 2 1 1 ...
#> $ y : num 63 75 69 70 66 70 75 74 63 64 ...
#> $ Diag: Factor w/ 3 levels "sepsis","cardiogenic",..: 1 1 1 2 2 1 1 1 1 2 ...
#> $ VO2 : num 207 300 289 150 350 150 305 140 167 318 ...
#> $ TCO : num 4.9 7.4 8.1 3.1 5.7 3.1 6.3 3.1 4.1 6.1 ...
#> $ FCO : num 5.3 10.2 8.7 3.7 7.6 3.7 9.6 3.7 3.4 6.8 ...
#> $ Sex : Factor w/ 2 levels "F","M": 2 2 1 1 2 2 1 2 1 1 ...
# A summary is not created if the the first argument (data=) is not used:
sv <- Meth( y=CardOutput[,c("Svo2","Scvo2")], item=CardOutput$VO2 )
#>
#> NOTE: Replication numbers generated in the order of the data
summary(sv)
#> #Replicates
#> Method 1 2 #Items #Obs: 30 Values: min med max
#> Scvo2 13 1 14 15 61 71 84
#> Svo2 13 1 14 15 62 70 82
# Sample items
ssv <- sample.Meth( sv, how="item", N=8 )
# More than two methods
data( sbp )
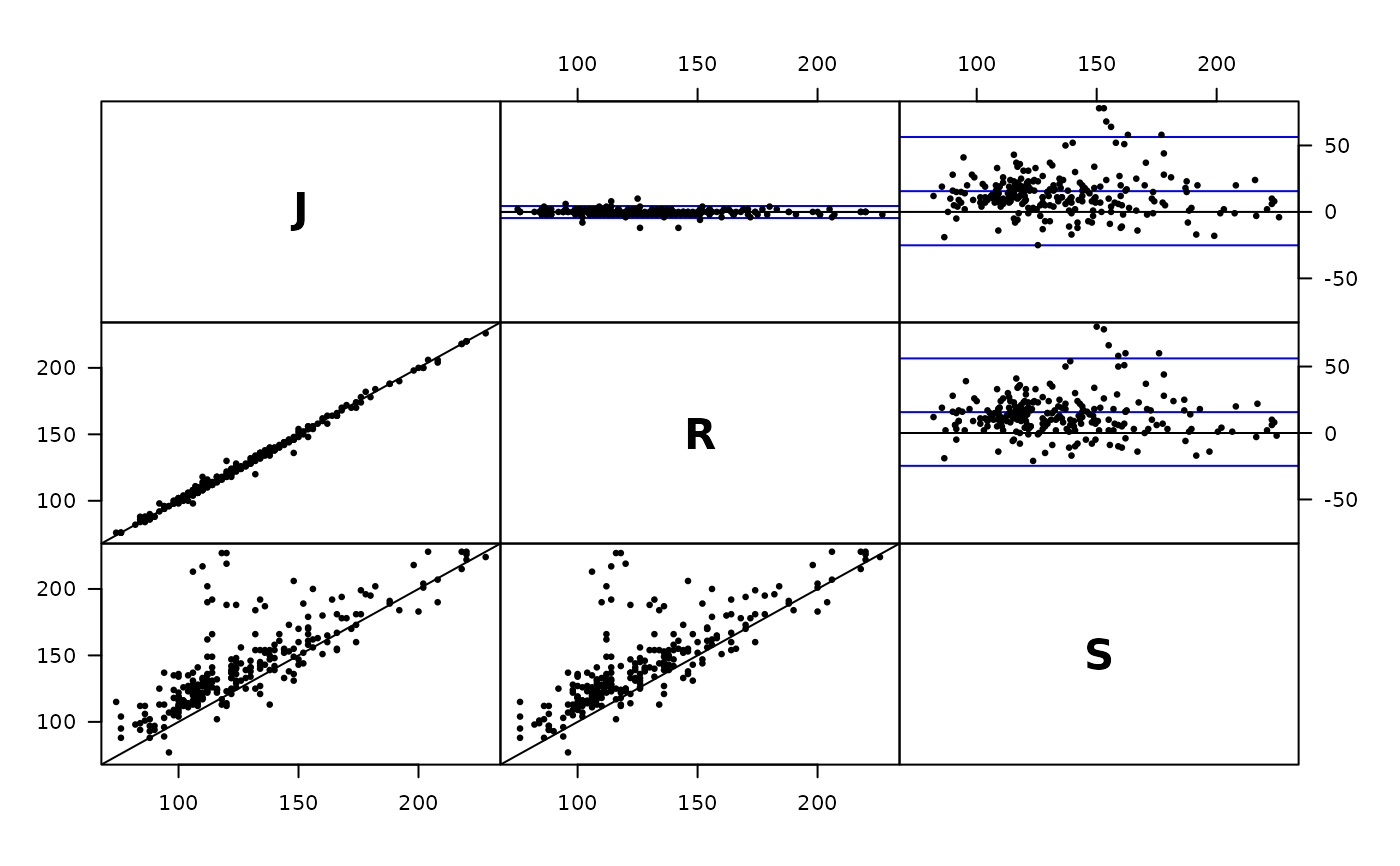
plot( Meth( sbp ) )
#> The following variables from the dataframe
#> "sbp" are used as the Meth variables:
#> meth: meth
#> item: item
#> repl: repl
#> y: y
#> #Replicates
#> Method 3 #Items #Obs: 765 Values: min med max
#> J 85 85 255 74 120 228
#> R 85 85 255 76 120 226
#> S 85 85 255 77 135 228
# Use observation in different columns as methods
data( CardOutput )
head( CardOutput )
#> Age Diag VO2 Svo2 Scvo2 TCO FCO Sex
#> 1 66 sepsis 207 63 62 4.9 5.3 M
#> 2 73 sepsis 300 75 78 7.4 10.2 M
#> 3 53 sepsis 289 69 70 8.1 8.7 F
#> 4 28 cardiogenic 150 70 71 3.1 3.7 F
#> 5 72 cardiogenic 350 66 68 5.7 7.6 M
#> 6 71 sepsis 150 70 72 3.1 3.7 M
sv <- Meth( CardOutput, y=c("Svo2","Scvo2") )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 #Items #Obs: 30 Values: min med max
#> Scvo2 15 15 15 61 71 84
#> Svo2 15 15 15 62 70 82
# Note that replicates are generated if a non-unique item-id is used
sv <- Meth( CardOutput, y=c("Svo2","Scvo2"), item="Age" )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> item: Age
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 2 3 #Items #Obs: 30 Values: min med max
#> Scvo2 8 2 1 11 15 61 71 84
#> Svo2 8 2 1 11 15 62 70 82
str( sv )
#> Classes ‘Meth’ and 'data.frame': 30 obs. of 9 variables:
#> $ meth: Factor w/ 2 levels "Scvo2","Svo2": 2 2 2 2 2 2 2 2 2 2 ...
#> $ item: Factor w/ 11 levels "24","28","53",..: 7 10 3 2 9 8 8 3 5 11 ...
#> $ repl: Factor w/ 3 levels "1","2","3": 1 1 1 1 1 1 2 2 1 1 ...
#> $ y : num 63 75 69 70 66 70 75 74 63 64 ...
#> $ Diag: Factor w/ 3 levels "sepsis","cardiogenic",..: 1 1 1 2 2 1 1 1 1 2 ...
#> $ VO2 : num 207 300 289 150 350 150 305 140 167 318 ...
#> $ TCO : num 4.9 7.4 8.1 3.1 5.7 3.1 6.3 3.1 4.1 6.1 ...
#> $ FCO : num 5.3 10.2 8.7 3.7 7.6 3.7 9.6 3.7 3.4 6.8 ...
#> $ Sex : Factor w/ 2 levels "F","M": 2 2 1 1 2 2 1 2 1 1 ...
# A summary is not created if the the first argument (data=) is not used:
sv <- Meth( y=CardOutput[,c("Svo2","Scvo2")], item=CardOutput$VO2 )
#>
#> NOTE: Replication numbers generated in the order of the data
summary(sv)
#> #Replicates
#> Method 1 2 #Items #Obs: 30 Values: min med max
#> Scvo2 13 1 14 15 61 71 84
#> Svo2 13 1 14 15 62 70 82
# Sample items
ssv <- sample.Meth( sv, how="item", N=8 )
# More than two methods
data( sbp )
plot( Meth( sbp ) )
#> The following variables from the dataframe
#> "sbp" are used as the Meth variables:
#> meth: meth
#> item: item
#> repl: repl
#> y: y
#> #Replicates
#> Method 3 #Items #Obs: 765 Values: min med max
#> J 85 85 255 74 120 228
#> R 85 85 255 76 120 226
#> S 85 85 255 77 135 228
 # Creating non-unique replicate numbers per (meth,item) creates a warning:
data( hba1c )
hb1 <- with( hba1c,
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 6 7 8 #Items #Obs: 835 Values: min med max
#> BR.V2 0 19 19 38 285 5.3 8.0 12.6
#> BR.VC 19 0 19 38 266 5.3 8.1 12.1
#> Tosoh 0 20 18 38 284 5.0 7.9 12.1
#>
#> WARNING: You have chosen a replicate variable which is not unique
#> within each (meth,item).
hb2 <- with( subset(hba1c,type=="Cap"),
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 3 4 #Items #Obs: 437 Values: min med max
#> BR.V2 0 38 38 152 5.3 8.0 12.6
#> BR.VC 19 19 38 133 5.3 8.2 12.1
#> Tosoh 0 38 38 152 5.0 7.8 11.8
# Creating non-unique replicate numbers per (meth,item) creates a warning:
data( hba1c )
hb1 <- with( hba1c,
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 6 7 8 #Items #Obs: 835 Values: min med max
#> BR.V2 0 19 19 38 285 5.3 8.0 12.6
#> BR.VC 19 0 19 38 266 5.3 8.1 12.1
#> Tosoh 0 20 18 38 284 5.0 7.9 12.1
#>
#> WARNING: You have chosen a replicate variable which is not unique
#> within each (meth,item).
hb2 <- with( subset(hba1c,type=="Cap"),
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 3 4 #Items #Obs: 437 Values: min med max
#> BR.V2 0 38 38 152 5.3 8.0 12.6
#> BR.VC 19 19 38 133 5.3 8.2 12.1
#> Tosoh 0 38 38 152 5.0 7.8 11.8