Replicate numbers are generated within (item,method) in a dataframe representing a method comparison study. The function assumes that observations are in the correct order within each (item,method), i.e. if replicate observations are non-exchangeable within method, linked observations are assumed to be in the same order within each (item,method).
perm.repl(data)Arguments
- data
A
Methobject or a data frame with columnsmeth,itemandy.
Value
make.repl returns a dataframe with a column, repl
added or replaced, whereas has.repl returns a logical indicating
wheter a combination of (meth,item) wioth more that one valid
\(y\)- value.
perm.repl returns a dataframe of class Meth where the
rows (i.e. replicates) are randomly permuted within
(meth,item), and subsequently ordered by
(meth,item,repl).
Details
make.repl just adds replicate numbers in the order of the data.frame
rows. perm.repl is designed to explore the effect of permuting the
replicates within (item,method). If replicates are truly exchangeable within
methods, the inference should be independent of this permutation.
See also
perm.repl
Examples
data(ox)
xx <- subset( ox, item<4 )[,-3]
cbind( xx, make.repl(xx) )
#> meth item y meth item y repl
#> 1 CO 1 78.0 CO 1 78.0 1
#> 2 CO 1 76.4 CO 1 76.4 2
#> 3 CO 1 77.2 CO 1 77.2 3
#> 4 CO 2 68.7 CO 2 68.7 1
#> 5 CO 2 67.6 CO 2 67.6 2
#> 6 CO 2 68.3 CO 2 68.3 3
#> 7 CO 3 82.9 CO 3 82.9 1
#> 8 CO 3 80.1 CO 3 80.1 2
#> 9 CO 3 80.7 CO 3 80.7 3
#> 184 pulse 1 71.0 pulse 1 71.0 1
#> 185 pulse 1 72.0 pulse 1 72.0 2
#> 186 pulse 1 73.0 pulse 1 73.0 3
#> 187 pulse 2 68.0 pulse 2 68.0 1
#> 188 pulse 2 67.0 pulse 2 67.0 2
#> 189 pulse 2 68.0 pulse 2 68.0 3
#> 190 pulse 3 82.0 pulse 3 82.0 1
#> 191 pulse 3 77.0 pulse 3 77.0 2
#> 192 pulse 3 77.0 pulse 3 77.0 3
cbind( make.repl(xx), perm.repl(xx) )
#> meth item y repl meth item repl y
#> 1 CO 1 78.0 1 CO 1 1 76.4
#> 2 CO 1 76.4 2 CO 1 2 77.2
#> 3 CO 1 77.2 3 CO 1 3 78.0
#> 4 CO 2 68.7 1 CO 2 1 68.7
#> 5 CO 2 67.6 2 CO 2 2 67.6
#> 6 CO 2 68.3 3 CO 2 3 68.3
#> 7 CO 3 82.9 1 CO 3 1 82.9
#> 8 CO 3 80.1 2 CO 3 2 80.7
#> 9 CO 3 80.7 3 CO 3 3 80.1
#> 184 pulse 1 71.0 1 pulse 1 1 73.0
#> 185 pulse 1 72.0 2 pulse 1 2 71.0
#> 186 pulse 1 73.0 3 pulse 1 3 72.0
#> 187 pulse 2 68.0 1 pulse 2 1 68.0
#> 188 pulse 2 67.0 2 pulse 2 2 67.0
#> 189 pulse 2 68.0 3 pulse 2 3 68.0
#> 190 pulse 3 82.0 1 pulse 3 1 77.0
#> 191 pulse 3 77.0 2 pulse 3 2 82.0
#> 192 pulse 3 77.0 3 pulse 3 3 77.0
data( ox )
xx <- subset( ox, item<4 )
cbind( xx, perm.repl(xx) )
#> meth item repl y meth item repl y
#> 1 CO 1 1 78.0 CO 1 1 77.2
#> 2 CO 1 2 76.4 CO 1 2 76.4
#> 3 CO 1 3 77.2 CO 1 3 78.0
#> 4 CO 2 1 68.7 CO 2 1 68.7
#> 5 CO 2 2 67.6 CO 2 2 67.6
#> 6 CO 2 3 68.3 CO 2 3 68.3
#> 7 CO 3 1 82.9 CO 3 1 80.1
#> 8 CO 3 2 80.1 CO 3 2 80.7
#> 9 CO 3 3 80.7 CO 3 3 82.9
#> 184 pulse 1 1 71.0 pulse 1 1 73.0
#> 185 pulse 1 2 72.0 pulse 1 2 71.0
#> 186 pulse 1 3 73.0 pulse 1 3 72.0
#> 187 pulse 2 1 68.0 pulse 2 1 68.0
#> 188 pulse 2 2 67.0 pulse 2 2 68.0
#> 189 pulse 2 3 68.0 pulse 2 3 67.0
#> 190 pulse 3 1 82.0 pulse 3 1 77.0
#> 191 pulse 3 2 77.0 pulse 3 2 82.0
#> 192 pulse 3 3 77.0 pulse 3 3 77.0
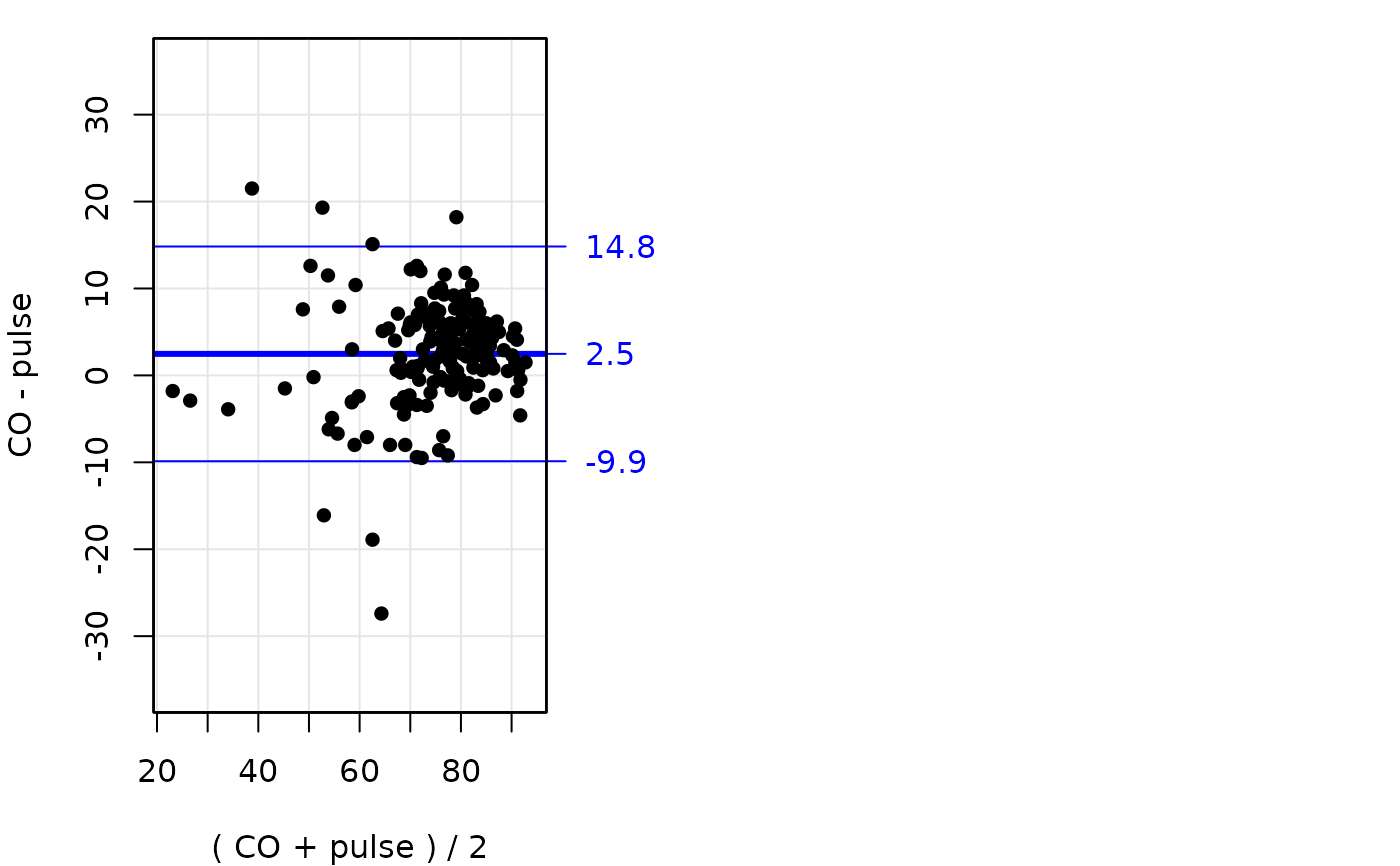
# Replicates are linked in the oximetry dataset, so randomly permuting
# them clearly inflates the limits of agreement:
par( mfrow=c(1,2), mar=c(4,4,1,4) )
BA.plot( ox , ymax=30, digits=1 )
#> Warning: "ymax" is not a graphical parameter
#> Warning: "ymax" is not a graphical parameter
#> Warning: "ymax" is not a graphical parameter
#> Warning: "ymax" is not a graphical parameter
#> Warning: "ymax" is not a graphical parameter
#> Warning: "ymax" is not a graphical parameter
#> Warning: "ymax" is not a graphical parameter
BA.plot( perm.repl(ox), ymax=30, digits=1 )
#> Error in solve.default(do.call("cbind", Xcols), apply(shifted, 2, fun, ...)): Lapack routine dgesv: system is exactly singular: U[9,9] = 0