Implementation of the Passing-Bablok's procedure for assessing of the equality of measurements by two different analytical methods.
PBreg(x, y = NULL, conf.level = 0.05, wh.meth = 1:2)Arguments
- x
a
Methobject, alternatively a numeric vector of measurements by method A, or a data frame of exactly two columns, first column with measurements by method A, second column with measurements by method B.- y
a numeric vector of measurements by method B - must be of the same length as
x. If not provided,xmust be theMethobject or a data frame of exactly 2 columns.- conf.level
confidence level for calculation of confidence boundaries - 0.05 is the default.
- wh.meth
Which of the methods from the
Methobject are used in the regression.
Value
PBreg returns an object of class "PBreg", for which
the print, predict and plot methods are defined.
An object of class "PBreg" is a list composed of the following
elements:
- coefficients
a matrix of 3 columns and 2 rows, containing the estimates of the intercept and slope, along with their confidence boundaries.
- residuals
defined as in the
"lm"class, as the response minus the fitted value.- fitted.values
the fitted values.
- model
the model data frame used.
- n
a vector of two values: the number of observations read, and the number of observations used.
- S
A vector of all slope estimates.
- I
A vector of all intercept estimates.
- adj
A vector of fit parameters, where Ss is the number of estimated slopes (
length(S)), K is the offset for slopes <(-1), M1 and M2 are the locations of confidence boundaries inS, and l and L are the numbers of points above and below the fitted line, used in cusum calculation.- cusum
A vector of cumulative sums of residuals sorted by the D-rank.
- Di
A vector of D-ranks.
Details
This is an implementation of the original Passing-Bablok procedure of fitting unbiased linear regression line to data in the method comparison studies. It calcualtes the unbiased slope and intercept, along with their confidence intervals. However, the tests for linearity is not yet fully implemented.
It doesn't matter which results are assigned to "Method A" and "Method B",
however the "Method A" results will be plotted on the x-axis by the
plot method.
Note
Please note that this method can become very computationally intensive for larger numbers of observations. One can expect a reasonable computation times for datasets with fewer than 100 observations.
References
Passing, H. and Bablok, W. (1983), A New Biometrical Procedure for Testing the Equality of Measurements from Two Different Analytical Methods. Journal of Clinical Chemistry and Clinical Biochemistry, Vol 21, 709–720
See also
Examples
## Model data frame generation
a <- data.frame(x=seq(1, 30)+rnorm(mean=0, sd=1, n=30),
y=seq(1, 30)*rnorm(mean=1, sd=0.4, n=30))
## Call to PBreg
x <- PBreg(a)
print(x)
#>
#> Passing-Bablok linear regression of y on x
#>
#> Observations read: 30, used: 30
#> Slopes calculated: 435, offset: 38
#>
#> Estimate 2.5%CI 97.5%CI
#> Intercept -1.104044 -5.3923293 0.772813
#> Slope 1.018875 0.8304804 1.336758
#>
#> Unadjusted summary of slopes:
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -66.2310 0.3324 0.8629 1.0447 1.4904 41.8334
#>
#> Summary of residuals:
#> Min. 1st Qu. Median Mean 3rd Qu. Max.
#> -12.024 -3.983 0.000 -1.136 2.437 11.998
#>
#> Test for linearity: (passed)
#> Linearity test not fully implemented in this version.
#>
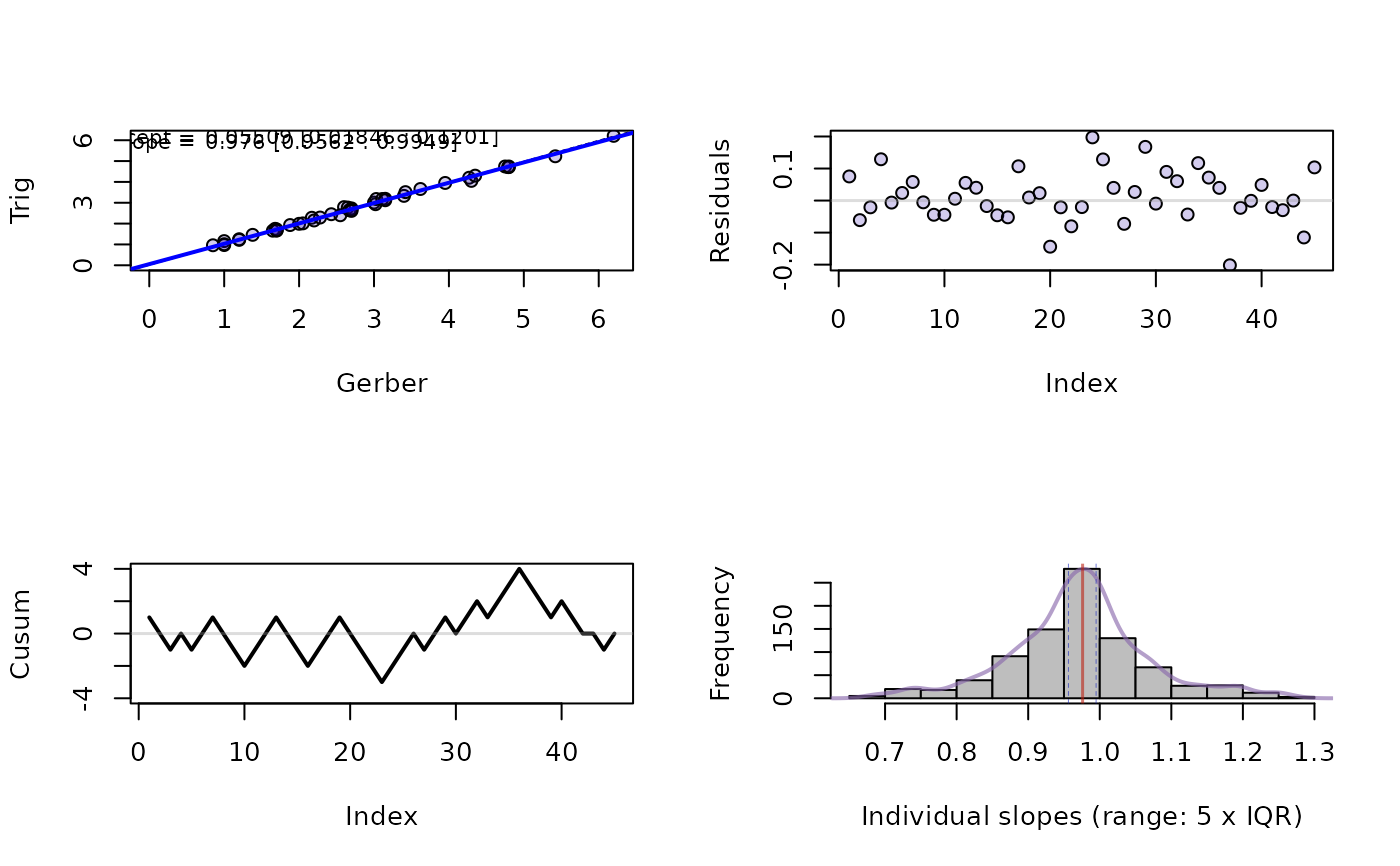
par(mfrow=c(2,2))
plot(x, s=1:4)
 ## A real data example
data(milk)
milk <- Meth(milk)
#> The following variables from the dataframe
#> "milk" are used as the Meth variables:
#> meth: meth
#> item: item
#> y: y
#> #Replicates
#> Method 1 #Items #Obs: 90 Values: min med max
#> Gerber 45 45 45 0.85 2.67 6.20
#> Trig 45 45 45 0.96 2.67 6.21
summary(milk)
#> #Replicates
#> Method 1 #Items #Obs: 90 Values: min med max
#> Gerber 45 45 45 0.85 2.67 6.20
#> Trig 45 45 45 0.96 2.67 6.21
PBmilk <- PBreg(milk)
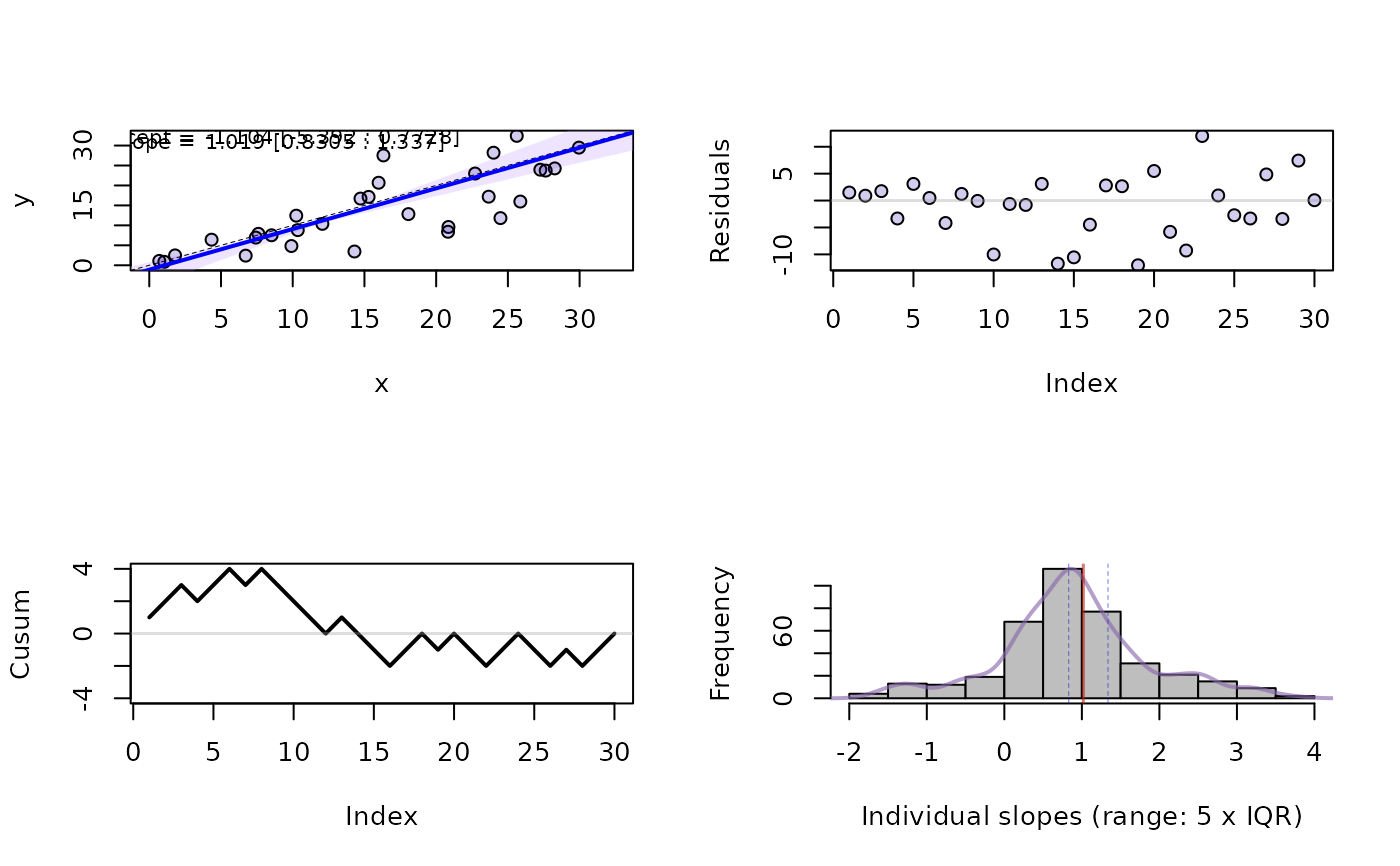
par(mfrow=c(2,2))
plot(PBmilk, s=1:4)
## A real data example
data(milk)
milk <- Meth(milk)
#> The following variables from the dataframe
#> "milk" are used as the Meth variables:
#> meth: meth
#> item: item
#> y: y
#> #Replicates
#> Method 1 #Items #Obs: 90 Values: min med max
#> Gerber 45 45 45 0.85 2.67 6.20
#> Trig 45 45 45 0.96 2.67 6.21
summary(milk)
#> #Replicates
#> Method 1 #Items #Obs: 90 Values: min med max
#> Gerber 45 45 45 0.85 2.67 6.20
#> Trig 45 45 45 0.96 2.67 6.21
PBmilk <- PBreg(milk)
par(mfrow=c(2,2))
plot(PBmilk, s=1:4)