Rank Correlation for Censored Data
rcorr.cens.RdComputes the c index and the corresponding generalization of Somers' Dxy rank correlation for a censored response variable. Also works for uncensored and binary responses, although its use of all possible pairings makes it slow for this purpose. Dxy and c are related by \(Dxy=2(c-0.5)\).
rcorr.cens handles one predictor variable. rcorrcens
computes rank correlation measures separately by a series of
predictors. In addition, rcorrcens has a rough way of handling
categorical predictors. If a categorical (factor) predictor has two
levels, it is coverted to a numeric having values 1 and 2. If it has
more than 2 levels, an indicator variable is formed for the most
frequently level vs. all others, and another indicator for the second
most frequent level and all others. The correlation is taken as the
maximum of the two (in absolute value).
Usage
rcorr.cens(x, S, outx=FALSE)
# S3 method for class 'formula'
rcorrcens(formula, data=NULL, subset=NULL,
na.action=na.retain, exclude.imputed=TRUE, outx=FALSE,
...)Arguments
- x
a numeric predictor variable
- S
an
Survobject or a vector. If a vector, assumes that every observation is uncensored.- outx
set to
TRUEto not count pairs of observations tied onxas a relevant pair. This results in a Goodman–Kruskal gamma type rank correlation.- formula
a formula with a
Survobject or a numeric vector on the left-hand side- data, subset, na.action
the usual options for models. Default for
na.actionis to retain all values, NA or not, so that NAs can be deleted in only a pairwise fashion.- exclude.imputed
set to
FALSEto include imputed values (created byimpute) in the calculations.- ...
extra arguments passed to
biVar.
Value
rcorr.cens returns a vector with the following named elements:
C Index, Dxy, S.D., n, missing,
uncensored, Relevant Pairs, Concordant, and
Uncertain
- n
number of observations not missing on any input variables
- missing
number of observations missing on
xorS- relevant
number of pairs of non-missing observations for which
Scould be ordered- concordant
number of relevant pairs for which
xandSare concordant.- uncertain
number of pairs of non-missing observations for which censoring prevents classification of concordance of
xandS.
rcorrcens.formula returns an object of class biVar
which is documented with the biVar function.
Author
Frank Harrell
Department of Biostatistics
Vanderbilt University
fh@fharrell.com
References
Newson R: Confidence intervals for rank statistics: Somers' D and extensions. Stata Journal 6:309-334; 2006.
Examples
set.seed(1)
x <- round(rnorm(200))
y <- rnorm(200)
rcorr.cens(x, y, outx=TRUE) # can correlate non-censored variables
#> C Index Dxy S.D. n missing
#> 4.76e-01 -4.74e-02 6.49e-02 2.00e+02 0.00e+00
#> uncensored Relevant Pairs Concordant Uncertain
#> 2.00e+02 2.83e+04 1.35e+04 0.00e+00
library(survival)
age <- rnorm(400, 50, 10)
bp <- rnorm(400,120, 15)
bp[1] <- NA
d.time <- rexp(400)
cens <- runif(400,.5,2)
death <- d.time <= cens
d.time <- pmin(d.time, cens)
rcorr.cens(age, Surv(d.time, death))
#> C Index Dxy S.D. n missing
#> 4.99e-01 -1.49e-03 3.67e-02 4.00e+02 0.00e+00
#> uncensored Relevant Pairs Concordant Uncertain
#> 2.79e+02 1.33e+05 6.66e+04 2.63e+04
r <- rcorrcens(Surv(d.time, death) ~ age + bp)
r
#>
#> Somers' Rank Correlation for Censored Data Response variable:Surv(d.time, death)
#>
#> C Dxy aDxy SD Z P n
#> age 0.499 -0.001 0.001 0.037 0.04 0.968 400
#> bp 0.497 -0.006 0.006 0.038 0.15 0.877 399
plot(r)
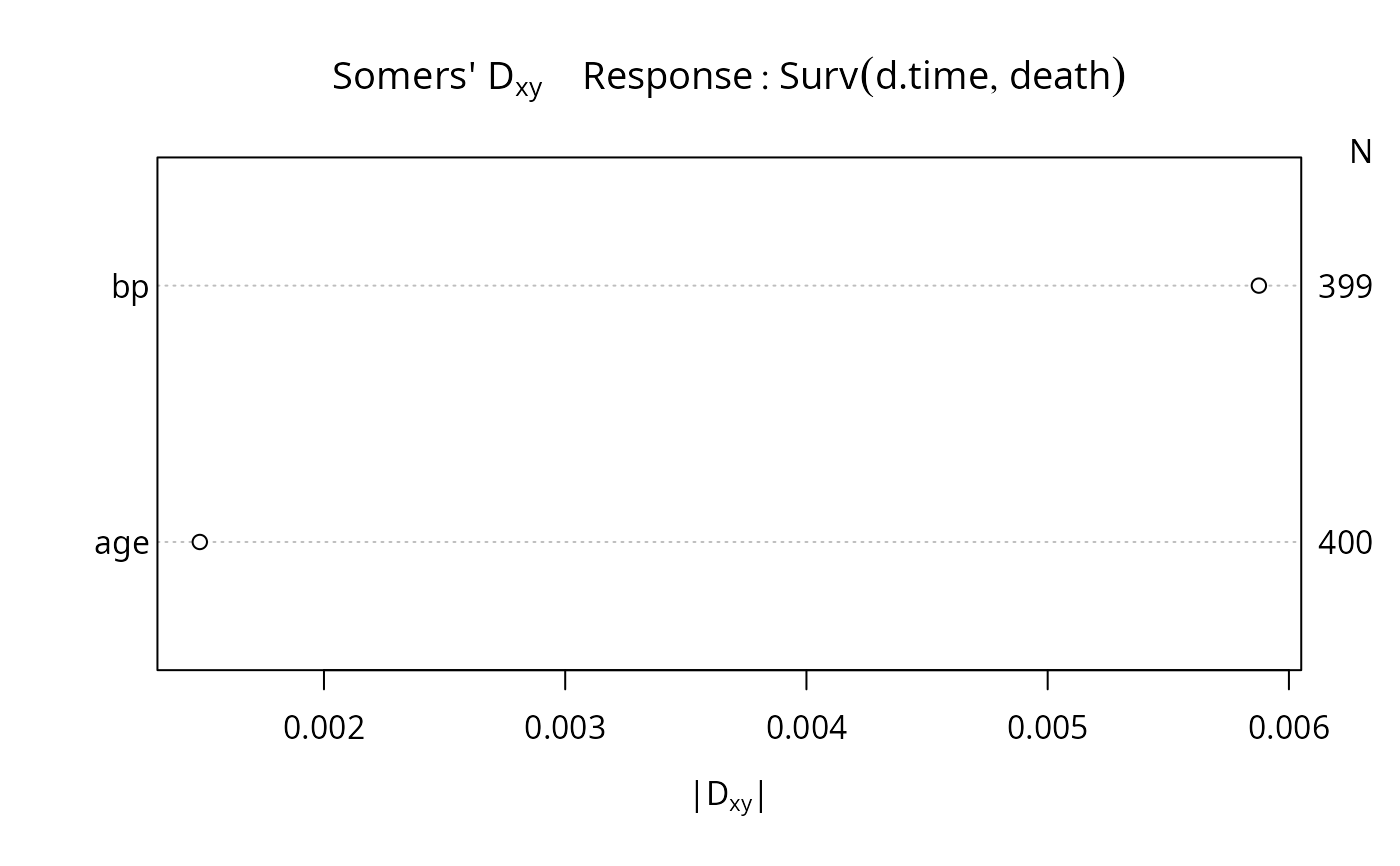 # Show typical 0.95 confidence limits for ROC areas for a sample size
# with 24 events and 62 non-events, for varying population ROC areas
# Repeat for 138 events and 102 non-events
set.seed(8)
par(mfrow=c(2,1))
for(i in 1:2) {
n1 <- c(24,138)[i]
n0 <- c(62,102)[i]
y <- c(rep(0,n0), rep(1,n1))
deltas <- seq(-3, 3, by=.25)
C <- se <- deltas
j <- 0
for(d in deltas) {
j <- j + 1
x <- c(rnorm(n0, 0), rnorm(n1, d))
w <- rcorr.cens(x, y)
C[j] <- w['C Index']
se[j] <- w['S.D.']/2
}
low <- C-1.96*se; hi <- C+1.96*se
print(cbind(C, low, hi))
errbar(deltas, C, C+1.96*se, C-1.96*se,
xlab='True Difference in Mean X',
ylab='ROC Area and Approx. 0.95 CI')
title(paste('n1=',n1,' n0=',n0,sep=''))
abline(h=.5, v=0, col='gray')
true <- 1 - pnorm(0, deltas, sqrt(2))
lines(deltas, true, col='blue')
}
#> C low hi
#> [1,] 0.0276 -0.001510 0.0566
#> [2,] 0.0336 -0.000378 0.0676
#> [3,] 0.0108 -0.004450 0.0260
#> [4,] 0.1042 0.028482 0.1799
#> [5,] 0.0679 0.014300 0.1215
#> [6,] 0.0981 0.026285 0.1700
#> [7,] 0.1472 0.068762 0.2256
#> [8,] 0.0927 0.023150 0.1623
#> [9,] 0.1532 0.060669 0.2458
#> [10,] 0.2386 0.132475 0.3447
#> [11,] 0.3085 0.194064 0.4229
#> [12,] 0.5141 0.385697 0.6425
#> [13,] 0.4335 0.297805 0.5691
#> [14,] 0.5410 0.409413 0.6726
#> [15,] 0.6136 0.486172 0.7410
#> [16,] 0.6418 0.508400 0.7752
#> [17,] 0.7695 0.666004 0.8730
#> [18,] 0.8233 0.718352 0.9282
#> [19,] 0.7977 0.689130 0.9063
#> [20,] 0.9079 0.849626 0.9662
#> [21,] 0.9362 0.882644 0.9897
#> [22,] 0.9577 0.921863 0.9935
#> [23,] 0.9630 0.928010 0.9981
#> [24,] 0.9906 0.978049 1.0031
#> [25,] 0.9718 0.935365 1.0082
#> C low hi
#> [1,] 0.0220 0.00644 0.0375
#> [2,] 0.0239 0.00933 0.0385
#> [3,] 0.0279 0.01162 0.0442
#> [4,] 0.0635 0.03341 0.0936
#> [5,] 0.0634 0.03448 0.0924
#> [6,] 0.1398 0.09424 0.1854
#> [7,] 0.1701 0.11980 0.2205
#> [8,] 0.1961 0.14033 0.2518
#> [9,] 0.2230 0.16465 0.2814
#> [10,] 0.2956 0.22882 0.3624
#> [11,] 0.3382 0.26882 0.4077
#> [12,] 0.3661 0.29566 0.4365
#> [13,] 0.5075 0.43368 0.5812
#> [14,] 0.6153 0.54412 0.6865
#> [15,] 0.6569 0.58852 0.7252
#> [16,] 0.6523 0.58239 0.7222
#> [17,] 0.7237 0.65994 0.7875
#> [18,] 0.8255 0.77347 0.8776
#> [19,] 0.8291 0.77867 0.8796
#> [20,] 0.8939 0.85444 0.9333
#> [21,] 0.8813 0.84016 0.9224
#> [22,] 0.9587 0.93522 0.9821
#> [23,] 0.9716 0.95470 0.9885
#> [24,] 0.9743 0.95601 0.9926
#> [25,] 0.9899 0.98147 0.9983
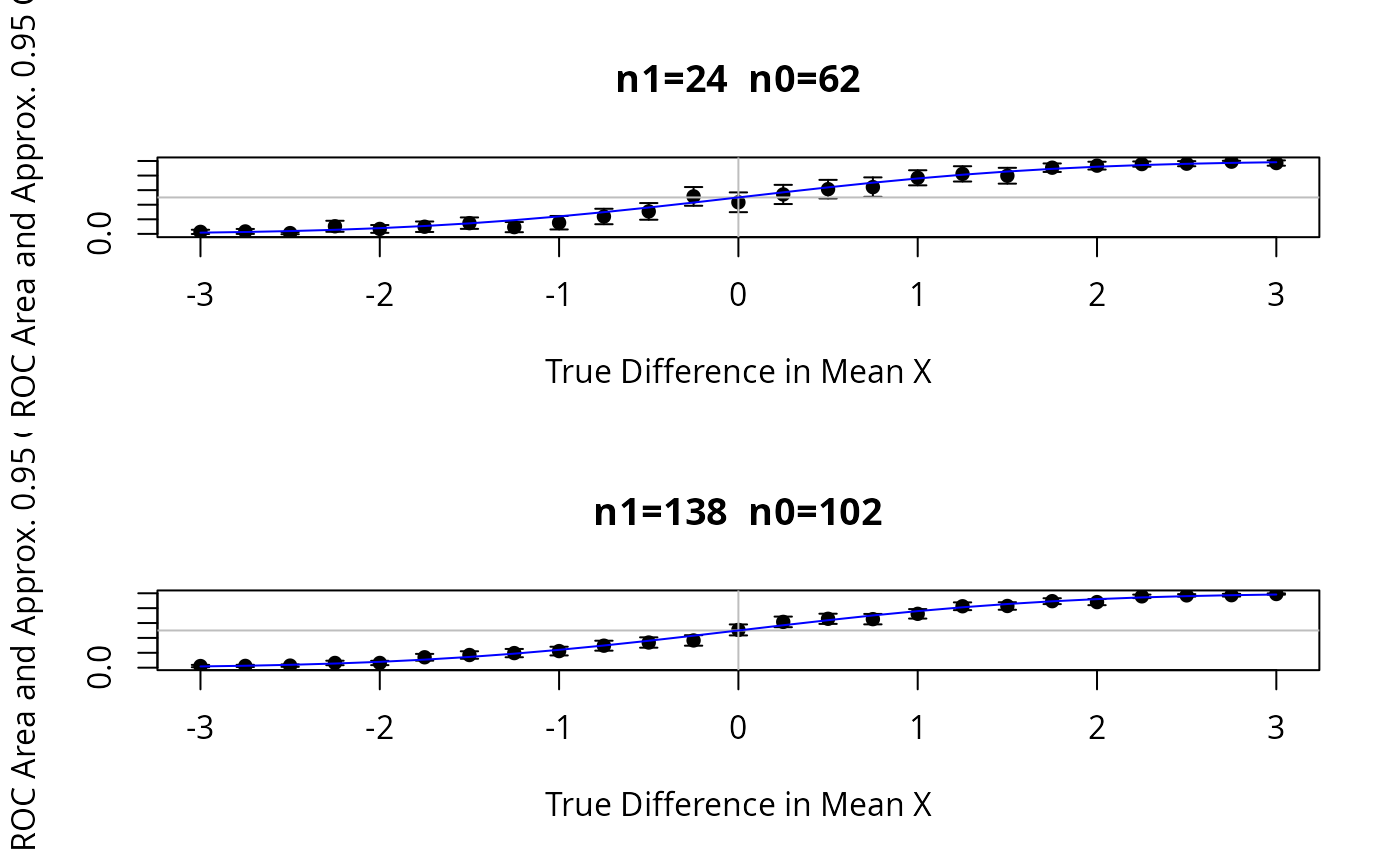
# Show typical 0.95 confidence limits for ROC areas for a sample size
# with 24 events and 62 non-events, for varying population ROC areas
# Repeat for 138 events and 102 non-events
set.seed(8)
par(mfrow=c(2,1))
for(i in 1:2) {
n1 <- c(24,138)[i]
n0 <- c(62,102)[i]
y <- c(rep(0,n0), rep(1,n1))
deltas <- seq(-3, 3, by=.25)
C <- se <- deltas
j <- 0
for(d in deltas) {
j <- j + 1
x <- c(rnorm(n0, 0), rnorm(n1, d))
w <- rcorr.cens(x, y)
C[j] <- w['C Index']
se[j] <- w['S.D.']/2
}
low <- C-1.96*se; hi <- C+1.96*se
print(cbind(C, low, hi))
errbar(deltas, C, C+1.96*se, C-1.96*se,
xlab='True Difference in Mean X',
ylab='ROC Area and Approx. 0.95 CI')
title(paste('n1=',n1,' n0=',n0,sep=''))
abline(h=.5, v=0, col='gray')
true <- 1 - pnorm(0, deltas, sqrt(2))
lines(deltas, true, col='blue')
}
#> C low hi
#> [1,] 0.0276 -0.001510 0.0566
#> [2,] 0.0336 -0.000378 0.0676
#> [3,] 0.0108 -0.004450 0.0260
#> [4,] 0.1042 0.028482 0.1799
#> [5,] 0.0679 0.014300 0.1215
#> [6,] 0.0981 0.026285 0.1700
#> [7,] 0.1472 0.068762 0.2256
#> [8,] 0.0927 0.023150 0.1623
#> [9,] 0.1532 0.060669 0.2458
#> [10,] 0.2386 0.132475 0.3447
#> [11,] 0.3085 0.194064 0.4229
#> [12,] 0.5141 0.385697 0.6425
#> [13,] 0.4335 0.297805 0.5691
#> [14,] 0.5410 0.409413 0.6726
#> [15,] 0.6136 0.486172 0.7410
#> [16,] 0.6418 0.508400 0.7752
#> [17,] 0.7695 0.666004 0.8730
#> [18,] 0.8233 0.718352 0.9282
#> [19,] 0.7977 0.689130 0.9063
#> [20,] 0.9079 0.849626 0.9662
#> [21,] 0.9362 0.882644 0.9897
#> [22,] 0.9577 0.921863 0.9935
#> [23,] 0.9630 0.928010 0.9981
#> [24,] 0.9906 0.978049 1.0031
#> [25,] 0.9718 0.935365 1.0082
#> C low hi
#> [1,] 0.0220 0.00644 0.0375
#> [2,] 0.0239 0.00933 0.0385
#> [3,] 0.0279 0.01162 0.0442
#> [4,] 0.0635 0.03341 0.0936
#> [5,] 0.0634 0.03448 0.0924
#> [6,] 0.1398 0.09424 0.1854
#> [7,] 0.1701 0.11980 0.2205
#> [8,] 0.1961 0.14033 0.2518
#> [9,] 0.2230 0.16465 0.2814
#> [10,] 0.2956 0.22882 0.3624
#> [11,] 0.3382 0.26882 0.4077
#> [12,] 0.3661 0.29566 0.4365
#> [13,] 0.5075 0.43368 0.5812
#> [14,] 0.6153 0.54412 0.6865
#> [15,] 0.6569 0.58852 0.7252
#> [16,] 0.6523 0.58239 0.7222
#> [17,] 0.7237 0.65994 0.7875
#> [18,] 0.8255 0.77347 0.8776
#> [19,] 0.8291 0.77867 0.8796
#> [20,] 0.8939 0.85444 0.9333
#> [21,] 0.8813 0.84016 0.9224
#> [22,] 0.9587 0.93522 0.9821
#> [23,] 0.9716 0.95470 0.9885
#> [24,] 0.9743 0.95601 0.9926
#> [25,] 0.9899 0.98147 0.9983
 par(mfrow=c(1,1))
par(mfrow=c(1,1))