The Shrinkage t Statistic
shrinkt.stat.Rdshrinkt.stat and shrinkt.fun compute the “shrinkage t” statistic
of Opgen-Rhein and Strimmer (2007).
Usage
shrinkt.stat(X, L, lambda.var, lambda.freqs, var.equal=TRUE,
paired=FALSE, verbose=TRUE)
shrinkt.fun(L, lambda.var, lambda.freqs, var.equal=TRUE, verbose=TRUE)Arguments
- X
data matrix. Note that the columns correspond to variables (“genes”) and the rows to samples.
- L
factor with class labels for the two groups. If only a single label is given then a one-sample t-score against 0 is computed.
- lambda.var
Shrinkage intensity for the variances. If not specified it is estimated from the data.
lambda.var=0implies no shrinkage andlambda.var=1complete shrinkage.- lambda.freqs
Shrinkage intensity for the frequencies. If not specified it is estimated from the data.
lambda.freqs=0implies no shrinkage (i.e. empirical frequencies).- var.equal
assume equal (default) or unequal variances in each group.
- paired
compute paired t-score (default is to use unpaired t-score).
- verbose
print out some (more or less useful) information during computation.
Details
The “shrinkage t” statistic is similar to the usual t statistic, with the replacement of the sample variances by corresponding shrinkage estimates. These are derived in a distribution-free fashion and with little a priori assumptions. Using the “shrinkage t” statistic procduces highly accurate rankings - see Opgen-Rhein and Strimmer (2007).
The“shrinkage t” statistic can be generalized to include gene-wise correlation,
see shrinkcat.stat.
The scale factor in the ”shrinkage t” statistic is computed from the estimated frequencies
(to use the standard empirical scale factor set lambda.freqs=0).
Value
shrinkt.stat returns a vector containing the “shrinkage t”
statistic for each variable/gene.
The corresponding shrinkt.fun functions return a function that
produces the “shrinkage t” statistics when applied to a data matrix
(this is very useful for simulations).
References
Opgen-Rhein, R., and K. Strimmer. 2007. Accurate ranking of differentially expressed genes by a distribution-free shrinkage approach. Statist. Appl. Genet. Mol. Biol. 6:9. <DOI:10.2202/1544-6115.1252>
Author
Rainer Opgen-Rhein, Verena Zuber, and Korbinian Strimmer (https://strimmerlab.github.io).
Examples
# load st library
library("st")
# load Choe et al. (2005) data
data(choedata)
X <- choe2.mat
dim(X) # 6 11475
#> [1] 6 11475
L <- choe2.L
L
#> [1] 1 1 1 2 2 2
# L may also contain some real labels
L = c("group 1", "group 1", "group 1", "group 2", "group 2", "group 2")
# shrinkage t statistic (equal variances)
score = shrinkt.stat(X, L)
#> Number of variables: 11475
#> Number of observations: 6
#> Number of classes: 2
#>
#> Estimating optimal shrinkage intensity lambda.freq (frequencies): 1
#> Estimating variances (pooled across classes)
#> Estimating optimal shrinkage intensity lambda.var (variance vector): 0.3882
#>
order(score^2, decreasing=TRUE)[1:10]
#> [1] 10979 11068 50 1022 724 5762 43 4790 10936 9939
# [1] 10979 11068 50 1022 724 5762 43 4790 10936 9939
# lambda.var (variances): 0.3882
# lambda.freqs (frequencies): 1
# shrinkage t statistic (unequal variances)
score = shrinkt.stat(X, L, var.equal=FALSE)
#> Number of variables: 11475
#> Number of observations: 6
#> Number of classes: 2
#>
#> Estimating optimal shrinkage intensity lambda.freq (frequencies): 1
#> Estimating variances (class #1)
#> Estimating optimal shrinkage intensity lambda.var (variance vector): 0.3673
#>
#> Estimating variances (class #2)
#> Estimating optimal shrinkage intensity lambda.var (variance vector): 0.3362
#>
#> Estimating variances (pooled across classes)
#> Estimating optimal shrinkage intensity lambda.var (variance vector): 0.3882
#>
order(score^2, decreasing=TRUE)[1:10]
#> [1] 11068 50 10979 724 43 1022 5762 10936 9939 9769
# [1] 11068 50 10979 724 43 1022 5762 10936 9939 9769
# lambda.var class #1 and class #2 (variances): 0.3673 0.3362
# lambda.freqs (frequencies): 1
# compute q-values and local false discovery rates
library("fdrtool")
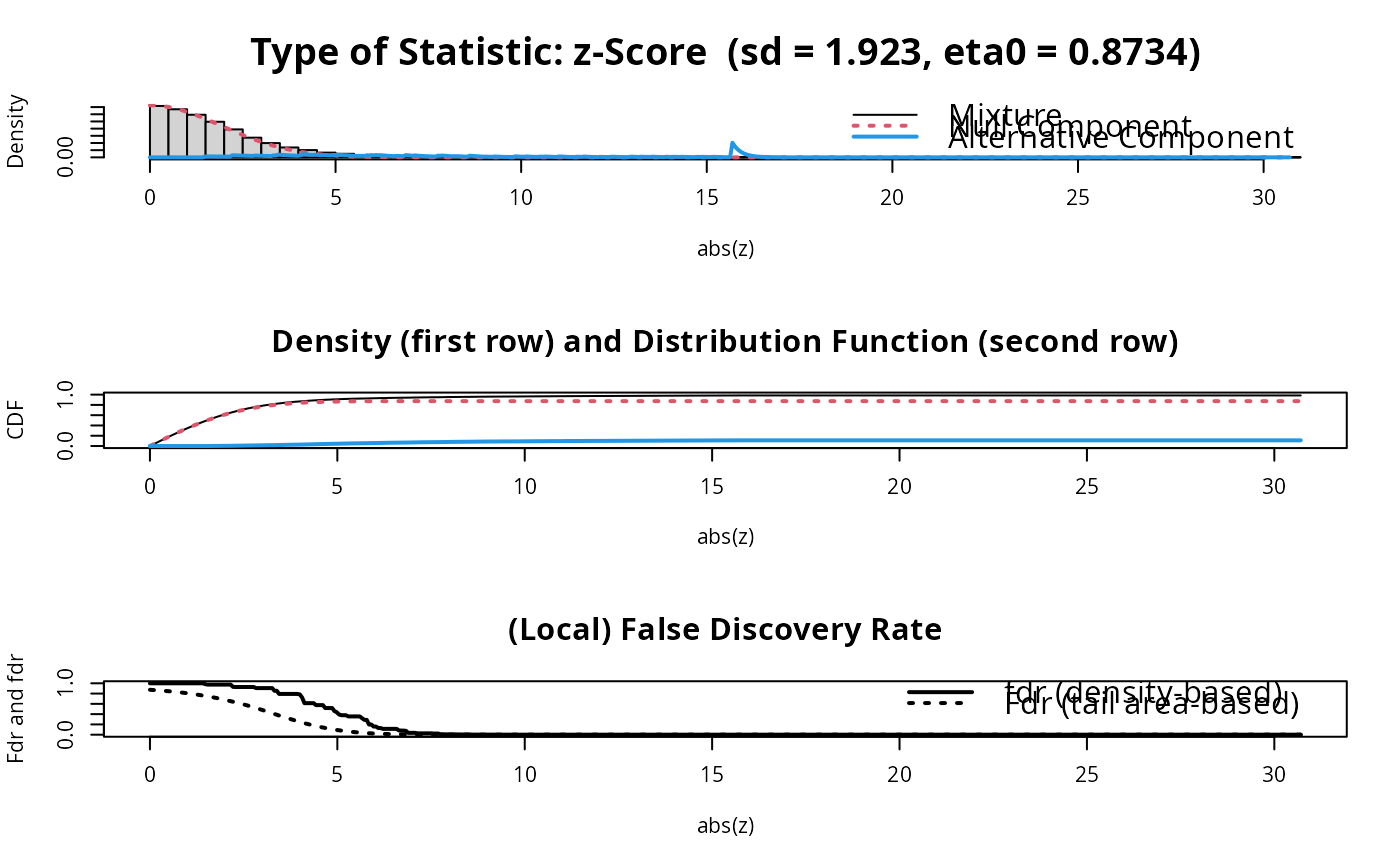
fdr.out = fdrtool(score)
#> Step 1... determine cutoff point
#> Step 2... estimate parameters of null distribution and eta0
#> Step 3... compute p-values and estimate empirical PDF/CDF
#> Step 4... compute q-values and local fdr
#> Step 5... prepare for plotting
 #>
sum( fdr.out$qval < 0.05 )
#> [1] 894
sum( fdr.out$lfdr < 0.2 )
#> [1] 846
fdr.out$param
#> cutoff N.cens eta0 eta0.SE sd sd.SE
#> [1,] 1.158258 4541 0.873391 0.01007508 1.92272 0.2685543
# computation of paired t-score
# paired shrinkage t statistic
score = shrinkt.stat(X, L, paired=TRUE)
#> Number of variables: 11475
#> Number of observations: 3
#> Number of classes: 1
#>
#> Estimating optimal shrinkage intensity lambda.freq (frequencies): 1
#> Estimating variances (pooled across classes)
#> Estimating optimal shrinkage intensity lambda.var (variance vector): 0.3498
#>
order(score^2, decreasing=TRUE)[1:10]
#> [1] 50 4790 5393 11068 5762 10238 9939 708 728 68
# [1] 50 4790 5393 11068 5762 10238 9939 708 728 68
# if there is no shrinkage the paired shrinkage t score reduces
# to the conventional paired student t statistic
score = studentt.stat(X, L, paired=TRUE)
score2 = shrinkt.stat(X, L, lambda.var=0, lambda.freqs=0, paired=TRUE, verbose=FALSE)
sum((score-score2)^2)
#> [1] 0
#>
sum( fdr.out$qval < 0.05 )
#> [1] 894
sum( fdr.out$lfdr < 0.2 )
#> [1] 846
fdr.out$param
#> cutoff N.cens eta0 eta0.SE sd sd.SE
#> [1,] 1.158258 4541 0.873391 0.01007508 1.92272 0.2685543
# computation of paired t-score
# paired shrinkage t statistic
score = shrinkt.stat(X, L, paired=TRUE)
#> Number of variables: 11475
#> Number of observations: 3
#> Number of classes: 1
#>
#> Estimating optimal shrinkage intensity lambda.freq (frequencies): 1
#> Estimating variances (pooled across classes)
#> Estimating optimal shrinkage intensity lambda.var (variance vector): 0.3498
#>
order(score^2, decreasing=TRUE)[1:10]
#> [1] 50 4790 5393 11068 5762 10238 9939 708 728 68
# [1] 50 4790 5393 11068 5762 10238 9939 708 728 68
# if there is no shrinkage the paired shrinkage t score reduces
# to the conventional paired student t statistic
score = studentt.stat(X, L, paired=TRUE)
score2 = shrinkt.stat(X, L, lambda.var=0, lambda.freqs=0, paired=TRUE, verbose=FALSE)
sum((score-score2)^2)
#> [1] 0