For each subset of a data frame, apply function then combine results into a
data frame.
To apply a function for each row, use adply with
.margins set to 1.
ddply(
.data,
.variables,
.fun = NULL,
...,
.progress = "none",
.inform = FALSE,
.drop = TRUE,
.parallel = FALSE,
.paropts = NULL
)Arguments
- .data
data frame to be processed
- .variables
variables to split data frame by, as
as.quotedvariables, a formula or character vector- .fun
function to apply to each piece
- ...
other arguments passed on to
.fun- .progress
name of the progress bar to use, see
create_progress_bar- .inform
produce informative error messages? This is turned off by default because it substantially slows processing speed, but is very useful for debugging
- .drop
should combinations of variables that do not appear in the input data be preserved (FALSE) or dropped (TRUE, default)
- .parallel
if
TRUE, apply function in parallel, using parallel backend provided by foreach- .paropts
a list of additional options passed into the
foreachfunction when parallel computation is enabled. This is important if (for example) your code relies on external data or packages: use the.exportand.packagesarguments to supply them so that all cluster nodes have the correct environment set up for computing.
Value
A data frame, as described in the output section.
Input
This function splits data frames by variables.
Output
The most unambiguous behaviour is achieved when .fun returns a
data frame - in that case pieces will be combined with
rbind.fill. If .fun returns an atomic vector of
fixed length, it will be rbinded together and converted to a data
frame. Any other values will result in an error.
If there are no results, then this function will return a data
frame with zero rows and columns (data.frame()).
References
Hadley Wickham (2011). The Split-Apply-Combine Strategy for Data Analysis. Journal of Statistical Software, 40(1), 1-29. https://www.jstatsoft.org/v40/i01/.
See also
Examples
# Summarize a dataset by two variables
dfx <- data.frame(
group = c(rep('A', 8), rep('B', 15), rep('C', 6)),
sex = sample(c("M", "F"), size = 29, replace = TRUE),
age = runif(n = 29, min = 18, max = 54)
)
# Note the use of the '.' function to allow
# group and sex to be used without quoting
ddply(dfx, .(group, sex), summarize,
mean = round(mean(age), 2),
sd = round(sd(age), 2))
#> group sex mean sd
#> 1 A F 37.70 11.81
#> 2 A M 31.53 5.47
#> 3 B F 34.00 10.17
#> 4 B M 36.01 7.17
#> 5 C F 28.36 8.50
#> 6 C M 29.74 9.78
# An example using a formula for .variables
ddply(baseball[1:100,], ~ year, nrow)
#> year V1
#> 1 1871 7
#> 2 1872 13
#> 3 1873 13
#> 4 1874 15
#> 5 1875 17
#> 6 1876 15
#> 7 1877 17
#> 8 1878 3
# Applying two functions; nrow and ncol
ddply(baseball, .(lg), c("nrow", "ncol"))
#> lg nrow ncol
#> 1 65 22
#> 2 AA 171 22
#> 3 AL 10007 22
#> 4 FL 37 22
#> 5 NL 11378 22
#> 6 PL 32 22
#> 7 UA 9 22
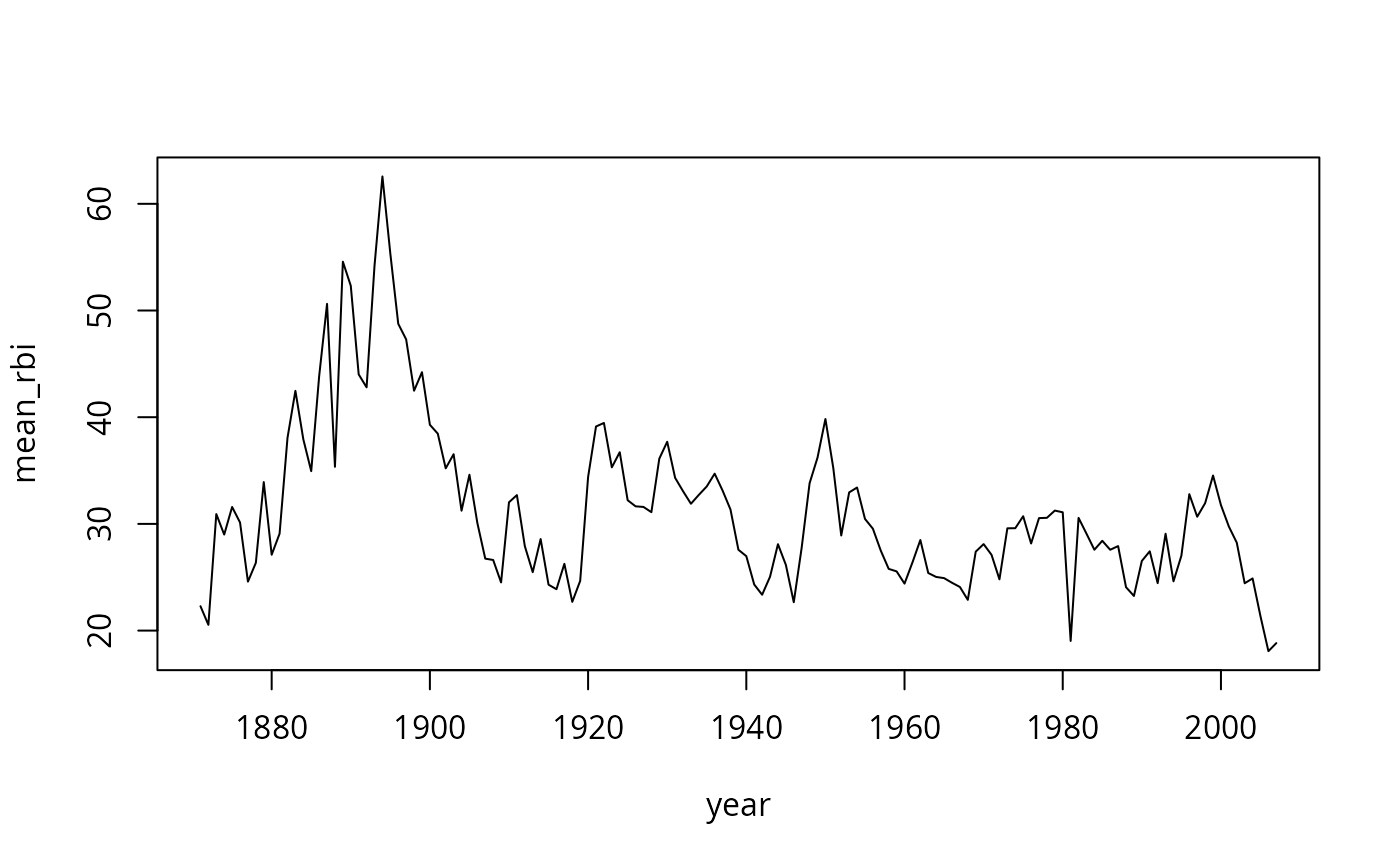
# Calculate mean runs batted in for each year
rbi <- ddply(baseball, .(year), summarise,
mean_rbi = mean(rbi, na.rm = TRUE))
# Plot a line chart of the result
plot(mean_rbi ~ year, type = "l", data = rbi)
 # make new variable career_year based on the
# start year for each player (id)
base2 <- ddply(baseball, .(id), mutate,
career_year = year - min(year) + 1
)
# make new variable career_year based on the
# start year for each player (id)
base2 <- ddply(baseball, .(id), mutate,
career_year = year - min(year) + 1
)