Slice the likelihood of a clm
slice.clm.RdSlice likelihood and plot the slice. This is usefull for illustrating the likelihood surface around the MLE (maximum likelihood estimate) and provides graphics to substantiate (non-)convergence of a model fit. Also, the closeness of a quadratic approximation to the log-likelihood function can be inspected for relevant parameters. A slice is considerably less computationally demanding than a profile.
Usage
slice(object, ...)
# S3 method for class 'clm'
slice(object, parm = seq_along(par), lambda = 3,
grid = 100, quad.approx = TRUE, ...)
# S3 method for class 'slice.clm'
plot(x, parm = seq_along(x),
type = c("quadratic", "linear"), plot.mle = TRUE,
ask = prod(par("mfcol")) < length(parm) && dev.interactive(), ...)Arguments
- object
for the
clmmethod an object of class"clm", i.e., the result of a call toclm.- x
a
slice.clmobject, i.e., the result ofslice(clm.object).- parm
for
slice.clma numeric or character vector indexing parameters, forplot.slice.clmonly a numeric vector is accepted. By default all parameters are selected.- lambda
the number of curvature units on each side of the MLE the slice should cover.
- grid
the number of values at which to compute the log-likelihood for each parameter.
- quad.approx
compute and include the quadratic approximation to the log-likelihood function?
- type
"quadratic"plots the log-likelihood function which is approximately quadratic, and"linear"plots the signed square root of the log-likelihood function which is approximately linear.- plot.mle
include a vertical line at the MLE (maximum likelihood estimate) when
type = "quadratic"? Ignored fortype = "linear".- ask
logical; if
TRUE, the user is asked before each plot, seepar(ask=.).- ...
further arguments to
plot.defaultfor the plot method. Not used in the slice method.
Value
The slice method returns a list of data.frames with one
data.frame for each parameter slice. Each data.frame
contains in the first column the values of the parameter and in the
second column the values of the (positive) log-likelihood
"logLik". A third column is present if quad.approx = TRUE
and contains the corresponding quadratic approximation to the
log-likelihood. The original model fit is included as the attribute
"original.fit".
The plot method produces a plot of the likelihood slice for
each parameter.
Examples
## fit model:
fm1 <- clm(rating ~ contact + temp, data = wine)
## slice the likelihood:
sl1 <- slice(fm1)
## three different ways to plot the slices:
par(mfrow = c(2,3))
plot(sl1)
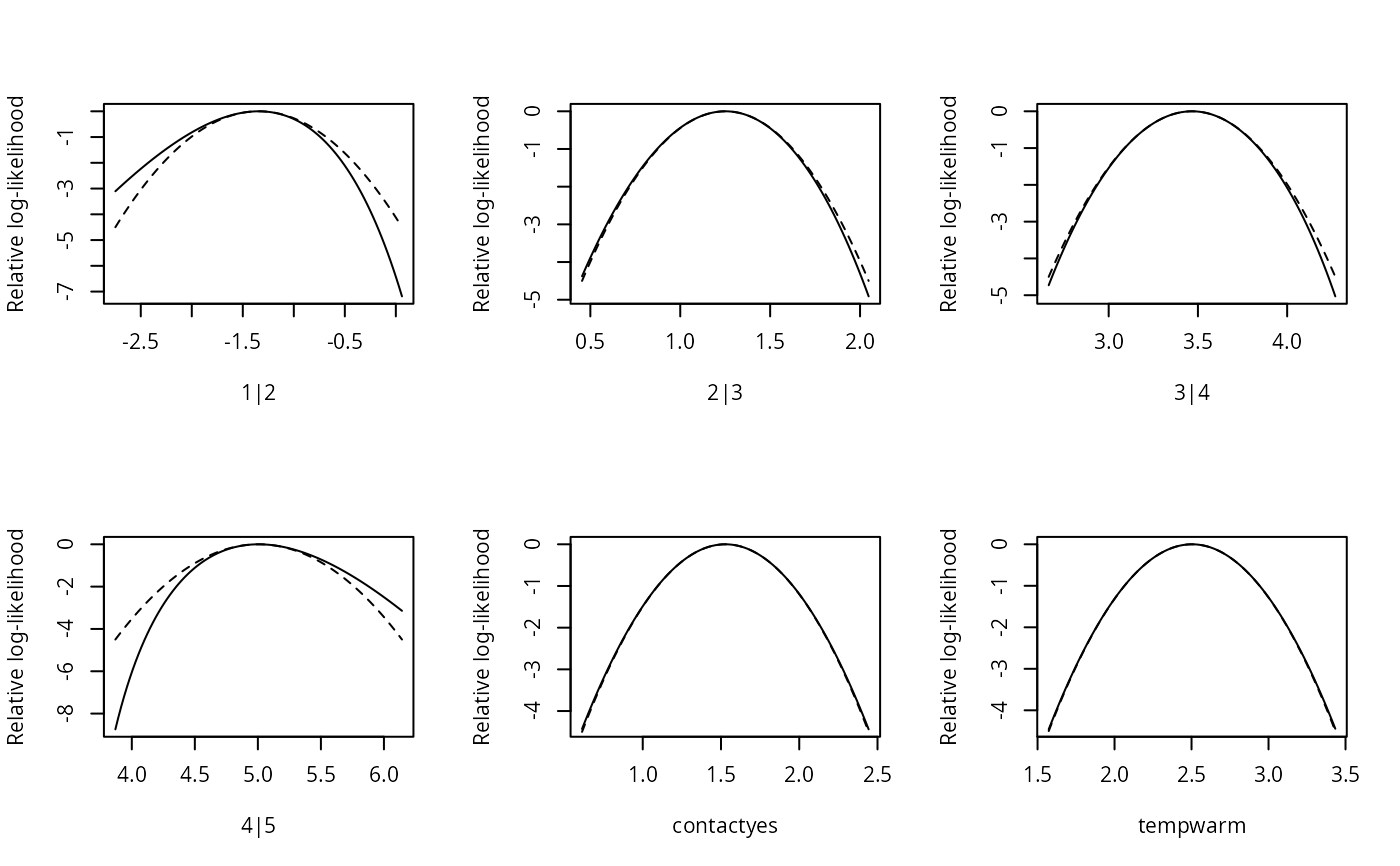
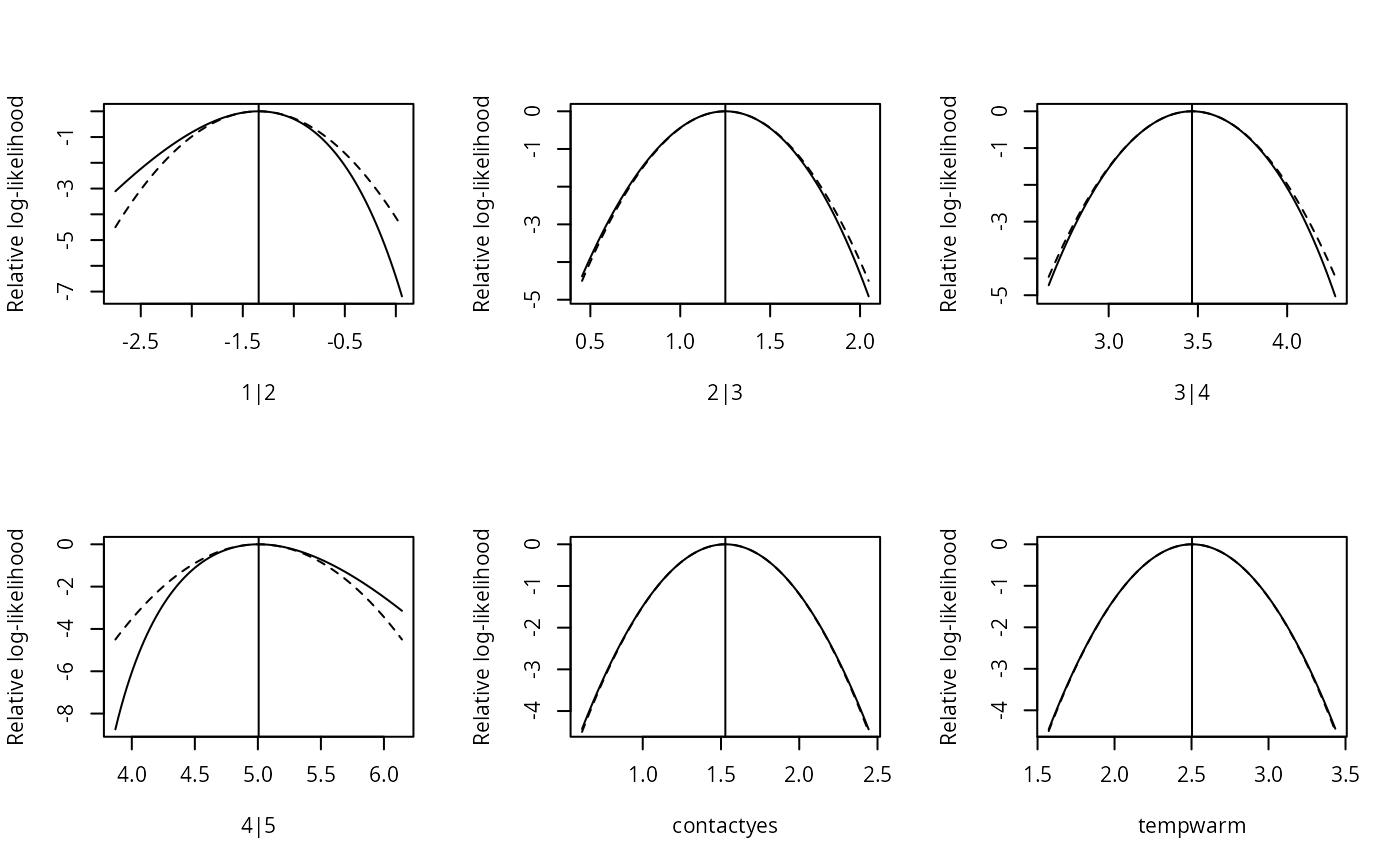 plot(sl1, type = "quadratic", plot.mle = FALSE)
plot(sl1, type = "quadratic", plot.mle = FALSE)
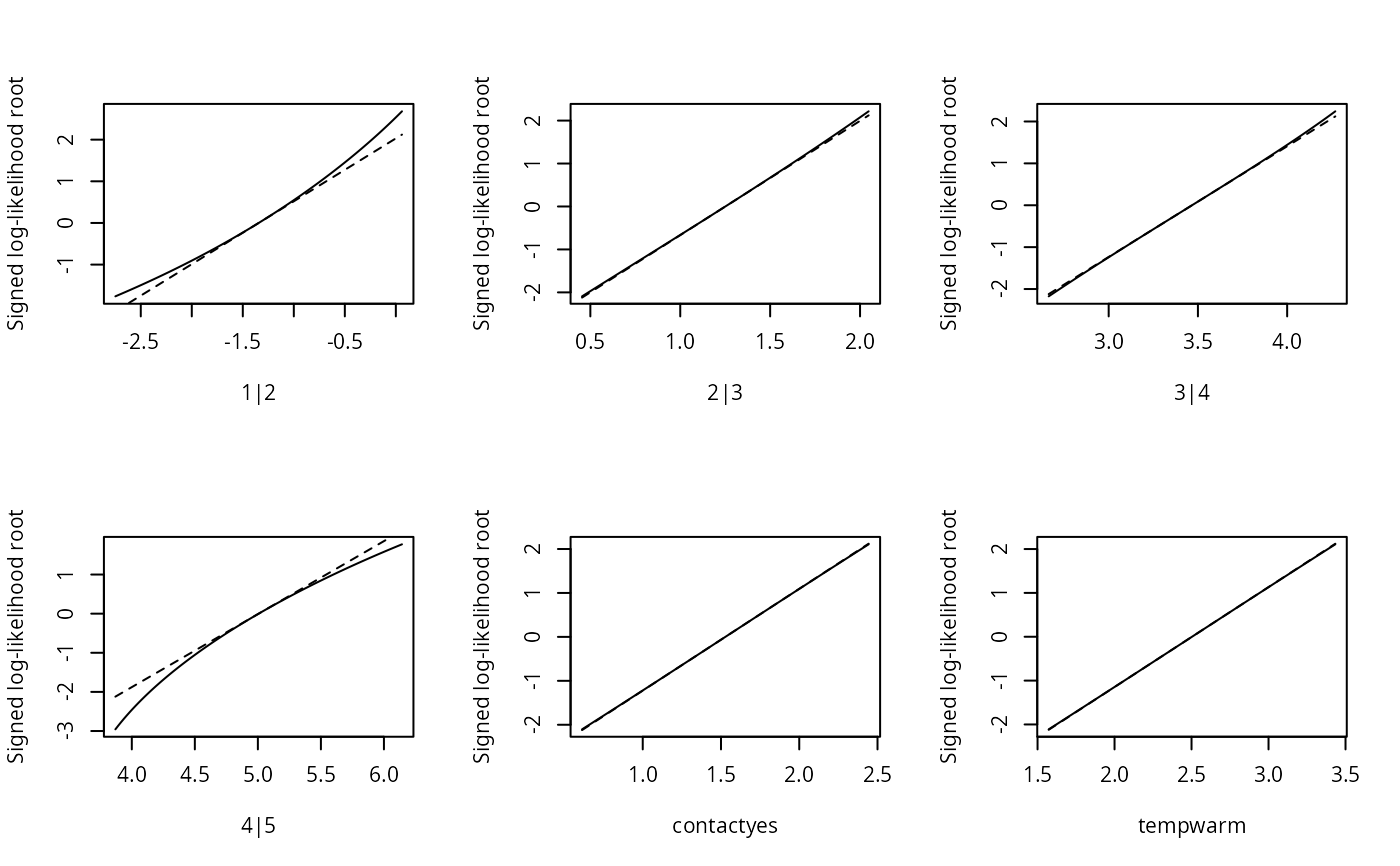
 plot(sl1, type = "linear")
plot(sl1, type = "linear")
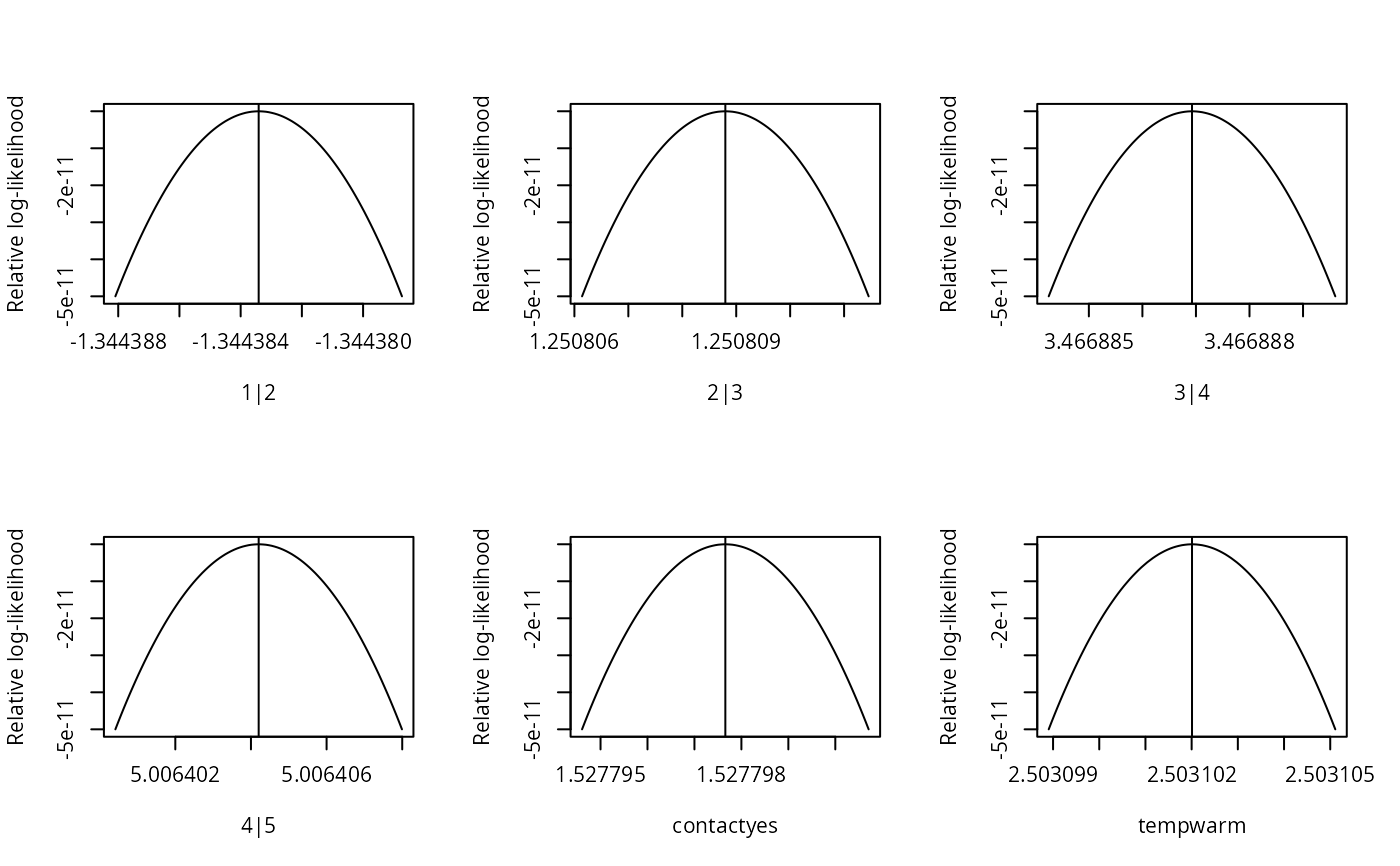
 ## Verify convergence to the optimum:
sl2 <- slice(fm1, lambda = 1e-5, quad.approx = FALSE)
plot(sl2)
## Verify convergence to the optimum:
sl2 <- slice(fm1, lambda = 1e-5, quad.approx = FALSE)
plot(sl2)