Set up a compact letter display of all pair-wise comparisons
cld.RdExtract information from glht, summary.glht or
confint.glht objects which is required to create
and plot compact letter displays of all pair-wise comparisons.
Usage
# S3 method for class 'summary.glht'
cld(object, level = 0.05, decreasing = FALSE, ...)
# S3 method for class 'glht'
cld(object, level = 0.05, decreasing = FALSE, ...)
# S3 method for class 'confint.glht'
cld(object, decreasing = FALSE, ...)Details
This function extracts all the information from glht,
summary.glht or confint.glht objects that is required
to create a compact letter display of all pair-wise comparisons
multcomp::piepho2004.
In case the contrast matrix is not of type "Tukey", an error
is issued. In case of confint.glht objects, a pair-wise comparison
is termed significant whenever a particular confidence interval contains 0.
Otherwise, p-values are compared to the value of "level".
Once, this information is extracted, plotting of all pair-wise
comparisons can be carried out.
Value
An object of class cld, a list with items:
- y
Values of the response variable of the original model.
- yname
Name of the response variable.
- x
Values of the variable used to compute Tukey contrasts.
- weights
Weights used in the fitting process.
- lp
Predictions from the fitted model.
- covar
A logical indicating whether the fitted model contained covariates.
- signif
Vector of logicals indicating significant differences with hyphenated names that identify pair-wise comparisons.
Examples
### multiple comparison procedures
### set up a one-way ANOVA
data(warpbreaks)
amod <- aov(breaks ~ tension, data = warpbreaks)
### specify all pair-wise comparisons among levels of variable "tension"
tuk <- glht(amod, linfct = mcp(tension = "Tukey"))
### extract information
tuk.cld <- cld(tuk)
### use sufficiently large upper margin
old.par <- par(mai=c(1,1,1.25,1), no.readonly = TRUE)
### plot
plot(tuk.cld)
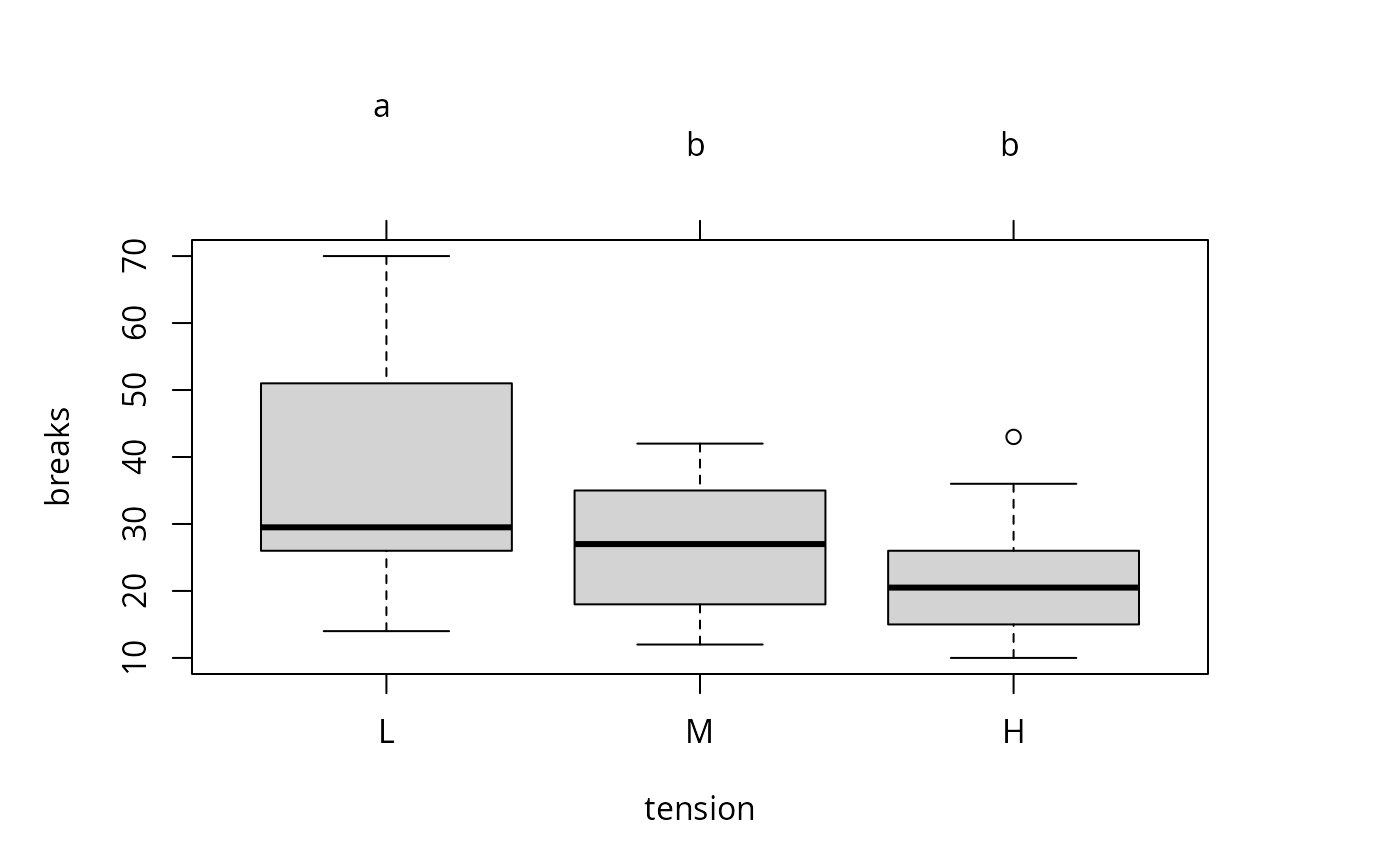 par(old.par)
### now using covariates
data(warpbreaks)
amod2 <- aov(breaks ~ tension + wool, data = warpbreaks)
### specify all pair-wise comparisons among levels of variable "tension"
tuk2 <- glht(amod2, linfct = mcp(tension = "Tukey"))
### extract information
tuk.cld2 <- cld(tuk2)
### use sufficiently large upper margin
old.par <- par(mai=c(1,1,1.25,1), no.readonly = TRUE)
### plot using different colors
plot(tuk.cld2, col=c("black", "red", "blue"))
par(old.par)
### now using covariates
data(warpbreaks)
amod2 <- aov(breaks ~ tension + wool, data = warpbreaks)
### specify all pair-wise comparisons among levels of variable "tension"
tuk2 <- glht(amod2, linfct = mcp(tension = "Tukey"))
### extract information
tuk.cld2 <- cld(tuk2)
### use sufficiently large upper margin
old.par <- par(mai=c(1,1,1.25,1), no.readonly = TRUE)
### plot using different colors
plot(tuk.cld2, col=c("black", "red", "blue"))
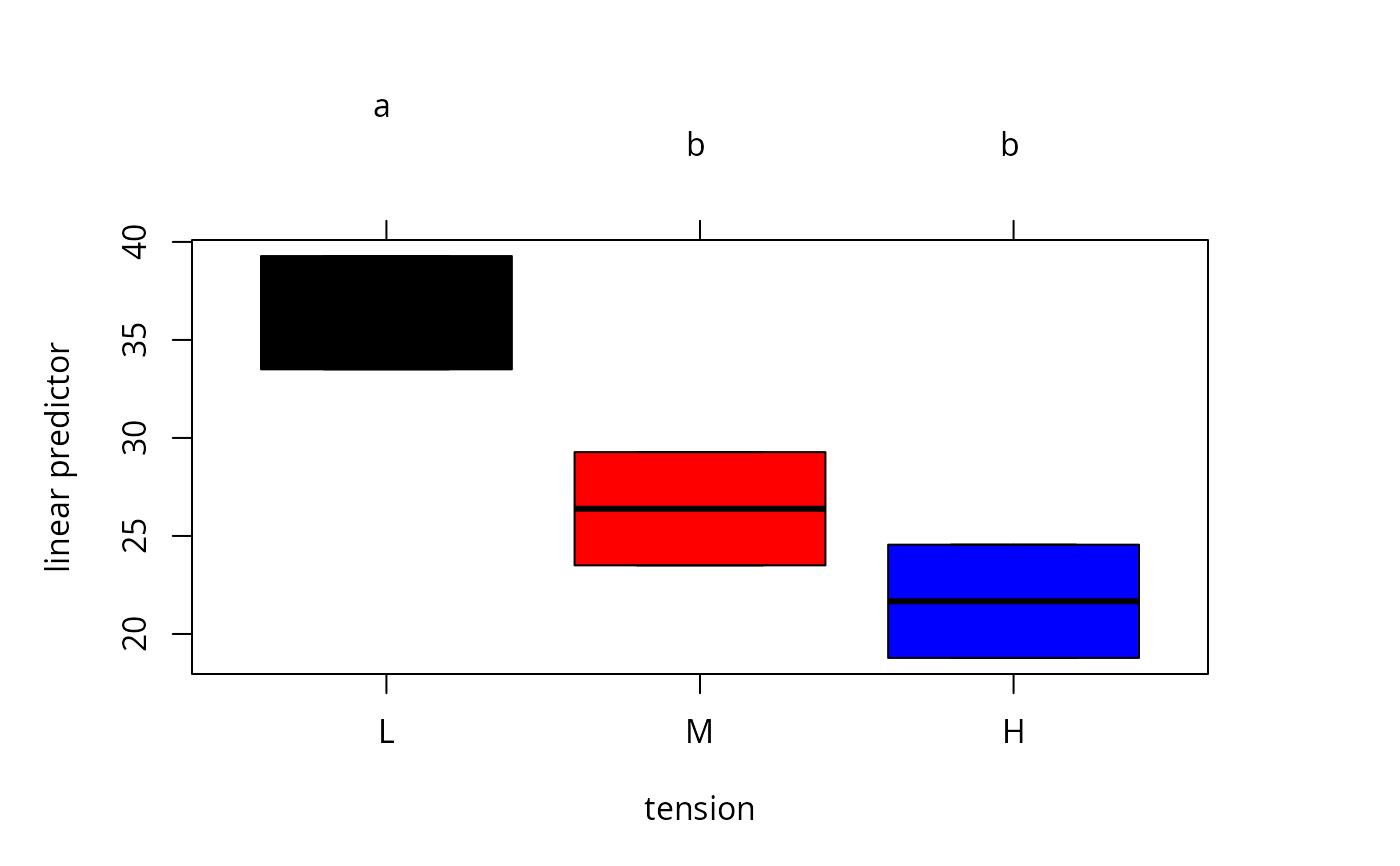 par(old.par)
### set up all pair-wise comparisons for count data
data(Titanic)
mod <- glm(Survived ~ Class, data = as.data.frame(Titanic), weights = Freq, family = binomial())
### specify all pair-wise comparisons among levels of variable "Class"
glht.mod <- glht(mod, mcp(Class = "Tukey"))
### extract information
mod.cld <- cld(glht.mod)
### use sufficiently large upper margin
old.par <- par(mai=c(1,1,1.5,1), no.readonly = TRUE)
### plot
plot(mod.cld)
par(old.par)
### set up all pair-wise comparisons for count data
data(Titanic)
mod <- glm(Survived ~ Class, data = as.data.frame(Titanic), weights = Freq, family = binomial())
### specify all pair-wise comparisons among levels of variable "Class"
glht.mod <- glht(mod, mcp(Class = "Tukey"))
### extract information
mod.cld <- cld(glht.mod)
### use sufficiently large upper margin
old.par <- par(mai=c(1,1,1.5,1), no.readonly = TRUE)
### plot
plot(mod.cld)
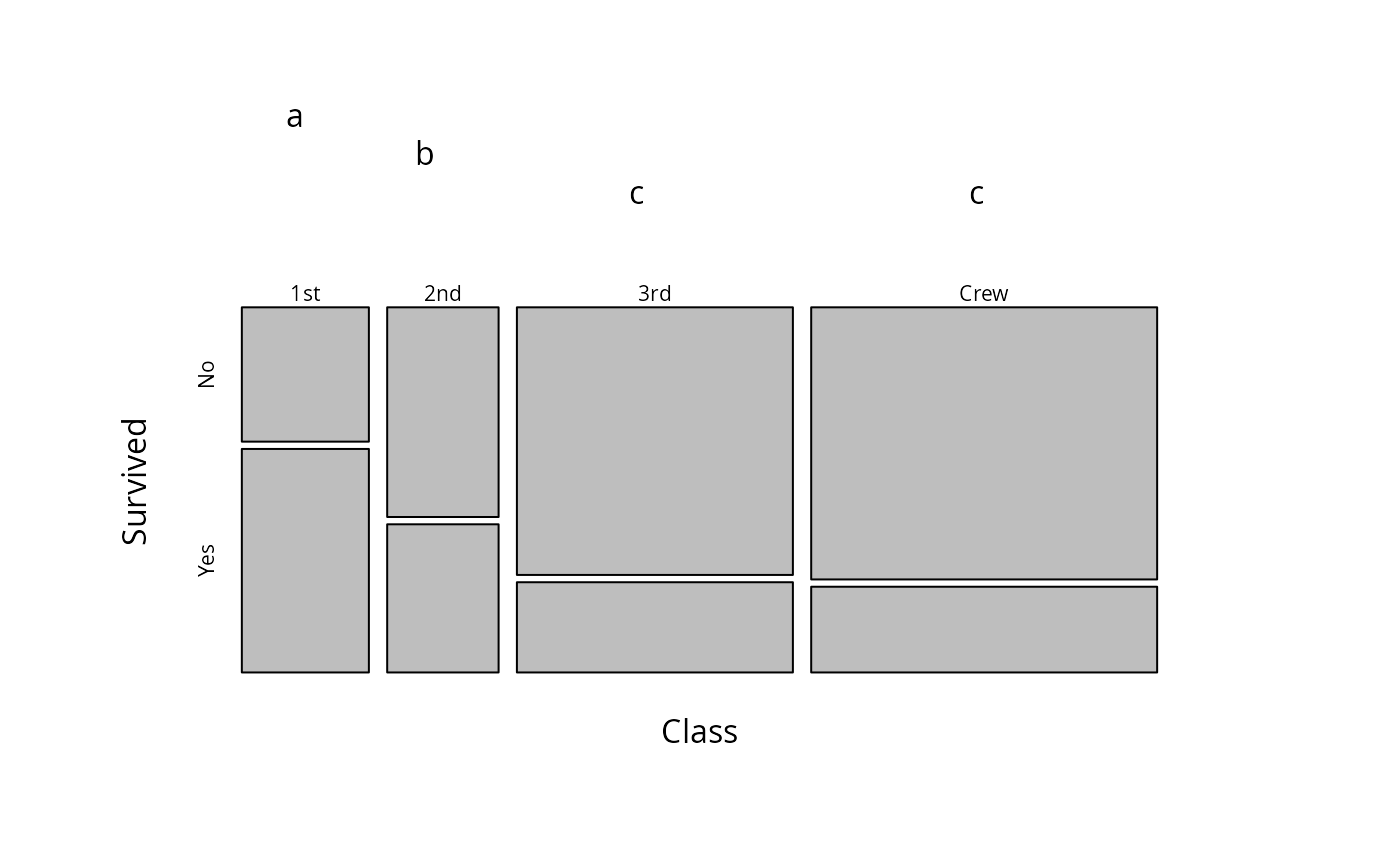 par(old.par)
par(old.par)