Sub-millisecond accurate timing of expression evaluation.
Source:R/microbenchmark.R
microbenchmark.Rdmicrobenchmark serves as a more accurate replacement of the
often seen system.time(replicate(1000, expr))
expression. It tries hard to accurately measure only the time it
takes to evaluate expr. To achieved this, the
sub-millisecond (supposedly nanosecond) accurate timing functions
most modern operating systems provide are used. Additionally all
evaluations of the expressions are done in C code to minimize any
overhead.
microbenchmark(
...,
list = NULL,
times = 100L,
unit = NULL,
check = NULL,
control = list(),
setup = NULL
)Arguments
- ...
Expressions to benchmark.
- list
List of unevaluated expressions to benchmark.
- times
Number of times to evaluate each expression.
- unit
Default unit used in
summaryandprint.- check
A function to check if the expressions are equal. By default
NULLwhich omits the check. In addition to a function, a string can be supplied. The string ‘equal’ will compare all values usingall.equal, ‘equivalent’ will compare all values usingall.equaland check.attributes = FALSE, and ‘identical’ will compare all values usingidentical.- control
List of control arguments. See Details.
- setup
An unevaluated expression to be run (untimed) before each benchmark expression.
Value
Object of class ‘microbenchmark’, a data frame with
columns expr and time. expr contains the
deparsed expression as passed to microbenchmark or the name
of the argument if the expression was passed as a named
argument. time is the measured execution time of the
expression in nanoseconds. The order of the observations in the
data frame is the order in which they were executed.
Details
This function is only meant for micro-benchmarking small pieces of source code and to compare their relative performance characteristics. You should generally avoid benchmarking larger chunks of your code using this function. Instead, try using the R profiler to detect hot spots and consider rewriting them in C/C++ or FORTRAN.
The control list can contain the following entries:
- order
the order in which the expressions are evaluated. “random” (the default) randomizes the execution order, “inorder” executes each expression in order and “block” executes all repetitions of each expression as one block.
- warmup
the number of iterations to run the timing code before evaluating the expressions in .... These warm-up iterations are used to estimate the timing overhead as well as spinning up the processor from any sleep or idle states it might be in. The default value is 2.
Note
Depending on the underlying operating system, different
methods are used for timing. On Windows the
QueryPerformanceCounter interface is used to measure the
time passed. For Linux the clock_gettime API is used and on
Solaris the gethrtime function. Finally on MacOS X the,
undocumented, mach_absolute_time function is used to avoid
a dependency on the CoreServices Framework.
Before evaluating each expression times times, the overhead
of calling the timing functions and the C function call overhead
are estimated. This estimated overhead is subtracted from each
measured evaluation time. Should the resulting timing be negative,
a warning is thrown and the respective value is replaced by
0. If the timing is zero, a warning is raised.
Should all evaluations result in one of the two error conditions described above, an error is raised.
One platform on which the clock resolution is known to be too low to measure short runtimes with the required precision is Oracle® Solaris on some SPARC® hardware. Reports of other platforms with similar problems are welcome. Please contact the package maintainer.
See also
print.microbenchmark to display and
boxplot.microbenchmark or
autoplot.microbenchmark to plot the results.
Examples
## Measure the time it takes to dispatch a simple function call
## compared to simply evaluating the constant \code{NULL}
f <- function() NULL
res <- microbenchmark(NULL, f(), times=1000L)
## Print results:
print(res)
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> NULL 5 17 23.534 22.0 29 317 1000 a
#> f() 107 173 200.668 183.5 194 4827 1000 b
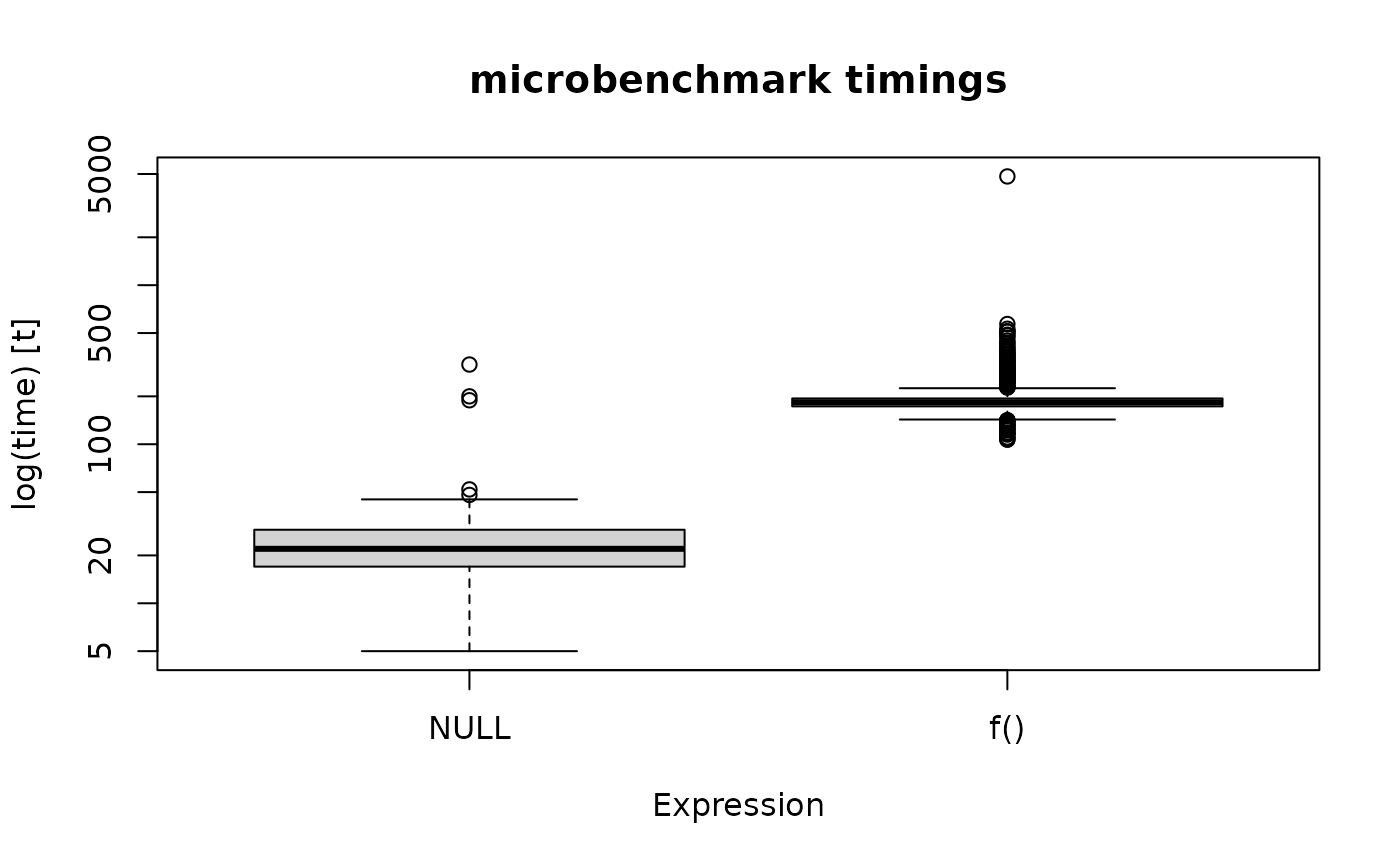
## Plot results:
boxplot(res)
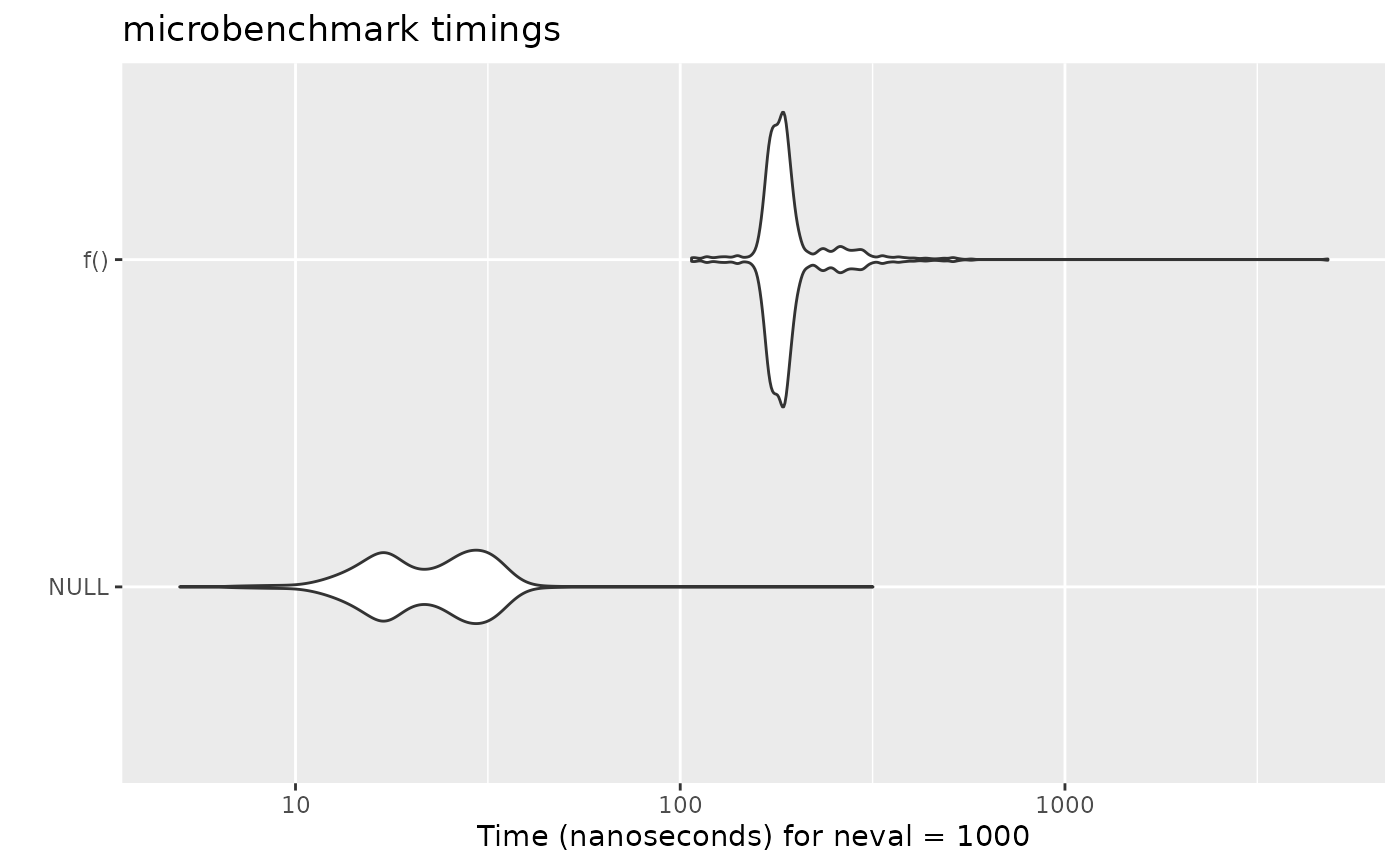
 ## Pretty plot:
if (requireNamespace("ggplot2")) {
ggplot2::autoplot(res)
}
## Pretty plot:
if (requireNamespace("ggplot2")) {
ggplot2::autoplot(res)
}
 ## Example check usage
my_check <- function(values) {
all(sapply(values[-1], function(x) identical(values[[1]], x)))
}
f <- function(a, b)
2 + 2
a <- 2
## Check passes
microbenchmark(2 + 2, 2 + a, f(2, a), f(2, 2), check=my_check)
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 56 69.0 90.79 76.5 91.0 355 100 a
#> 2 + a 67 79.0 110.09 84.0 95.0 846 100 a
#> f(2, a) 228 249.0 402.86 284.5 488.5 1063 100 b
#> f(2, 2) 226 247.5 425.18 262.0 393.5 7644 100 b
if (FALSE) { # \dontrun{
a <- 3
## Check fails
microbenchmark(2 + 2, 2 + a, f(2, a), f(2, 2), check=my_check)
} # }
## Example setup usage
set.seed(21)
x <- rnorm(10)
microbenchmark(x, rnorm(10), check=my_check, setup=set.seed(21))
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> x 51 68 87.24 78.0 85 1091 100 a
#> rnorm(10) 1830 2292 2492.73 2395.5 2477 13074 100 b
## Will fail without setup
if (FALSE) { # \dontrun{
microbenchmark(x, rnorm(10), check=my_check)
} # }
## using check
a <- 2
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='identical')
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 75 134.5 176.65 158.0 177.5 1317 100 a
#> 2 + a 106 155.5 184.94 179.0 195.5 594 100 a
#> sum(2, a) 254 387.5 517.06 426.5 456.5 7199 100 b
#> sum(2, 2) 256 374.0 427.70 409.5 436.0 915 100 b
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='equal')
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 106 172.0 195.66 185.0 198.0 651 100 a
#> 2 + a 133 176.5 218.40 199.5 223.0 1868 100 a
#> sum(2, a) 339 458.0 517.02 493.5 522.0 1061 100 b
#> sum(2, 2) 291 427.0 545.54 477.5 499.5 6531 100 b
attr(a, 'abc') <- 123
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='equivalent')
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 53 67.0 95.58 72.5 87.5 732 100 a
#> 2 + a 126 140.5 234.57 151.0 187.0 4554 100 ab
#> sum(2, a) 179 191.5 363.03 200.0 295.5 11155 100 b
#> sum(2, 2) 171 187.0 271.80 197.5 302.0 811 100 ab
## check='equal' will fail due to difference in attribute
if (FALSE) { # \dontrun{
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='equal')
} # }
## Example check usage
my_check <- function(values) {
all(sapply(values[-1], function(x) identical(values[[1]], x)))
}
f <- function(a, b)
2 + 2
a <- 2
## Check passes
microbenchmark(2 + 2, 2 + a, f(2, a), f(2, 2), check=my_check)
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 56 69.0 90.79 76.5 91.0 355 100 a
#> 2 + a 67 79.0 110.09 84.0 95.0 846 100 a
#> f(2, a) 228 249.0 402.86 284.5 488.5 1063 100 b
#> f(2, 2) 226 247.5 425.18 262.0 393.5 7644 100 b
if (FALSE) { # \dontrun{
a <- 3
## Check fails
microbenchmark(2 + 2, 2 + a, f(2, a), f(2, 2), check=my_check)
} # }
## Example setup usage
set.seed(21)
x <- rnorm(10)
microbenchmark(x, rnorm(10), check=my_check, setup=set.seed(21))
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> x 51 68 87.24 78.0 85 1091 100 a
#> rnorm(10) 1830 2292 2492.73 2395.5 2477 13074 100 b
## Will fail without setup
if (FALSE) { # \dontrun{
microbenchmark(x, rnorm(10), check=my_check)
} # }
## using check
a <- 2
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='identical')
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 75 134.5 176.65 158.0 177.5 1317 100 a
#> 2 + a 106 155.5 184.94 179.0 195.5 594 100 a
#> sum(2, a) 254 387.5 517.06 426.5 456.5 7199 100 b
#> sum(2, 2) 256 374.0 427.70 409.5 436.0 915 100 b
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='equal')
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 106 172.0 195.66 185.0 198.0 651 100 a
#> 2 + a 133 176.5 218.40 199.5 223.0 1868 100 a
#> sum(2, a) 339 458.0 517.02 493.5 522.0 1061 100 b
#> sum(2, 2) 291 427.0 545.54 477.5 499.5 6531 100 b
attr(a, 'abc') <- 123
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='equivalent')
#> Unit: nanoseconds
#> expr min lq mean median uq max neval cld
#> 2 + 2 53 67.0 95.58 72.5 87.5 732 100 a
#> 2 + a 126 140.5 234.57 151.0 187.0 4554 100 ab
#> sum(2, a) 179 191.5 363.03 200.0 295.5 11155 100 b
#> sum(2, 2) 171 187.0 271.80 197.5 302.0 811 100 ab
## check='equal' will fail due to difference in attribute
if (FALSE) { # \dontrun{
microbenchmark(2 + 2, 2 + a, sum(2, a), sum(2, 2), check='equal')
} # }