library(ggplot2)
library(dplyr)
#>
#> Attaching package: 'dplyr'
#> The following objects are masked from 'package:stats':
#>
#> filter, lag
#> The following objects are masked from 'package:base':
#>
#> intersect, setdiff, setequal, union
library(tidyr)
library(hyperion)
#>
#>
#> ── pharos configuration ────────────────────────────────────────────────────────
#> ✔ pharos.toml found: /data/user-homes/tariq/projects/prism-pkgdocs-build/installed-pkgs/2026-03-02/hyperion_0.3.2/vignettes/pharos.toml
#> ── hyperion options ────────────────────────────────────────────────────────────
#> ✔ hyperion.significant_number_display : 4
#> ── hyperion nonmem object options ──────────────────────────────────────────────
#> ✔ hyperion.nonmem_model.show_included_columns : FALSE
#> ✔ hyperion.nonmem_summary.rse_threshold : 50
#> ✔ hyperion.nonmem_summary.shrinkage_threshold : 30
test_data_dir <- system.file("extdata", package = "hyperion")
get parameter estimates
get_parameters(file.path(test_data_dir, "models", "onecmt", "run002"))
#> kind name random_effect estimate sd corr stderr
#> 1 THETA TVCL <NA> 1.24679000 NA NA 0.1288330
#> 2 THETA TVV <NA> 40.84820000 NA NA 3.0272100
#> 3 THETA TVKA <NA> 1.24394000 NA NA 0.1134100
#> 4 OMEGA OM1 (TVCL) ETA1 0.13041700 0.3611320 NA 0.0601864
#> 5 OMEGA OM2 (TVV) ETA2 0.13633200 0.3692310 NA 0.0397114
#> 6 OMEGA OM3 (TVKA) ETA3 0.11440800 0.3382430 NA 0.0614429
#> 7 SIGMA SIGMA(1,1) EPS1 0.03723180 0.1929550 NA 0.0115996
#> 8 SIGMA SIGMA(2,2) EPS2 0.00660722 0.0812848 NA 0.0279207
#> rse shrinkage fixed diagonal
#> 1 10.333176 NA FALSE NA
#> 2 7.410877 NA FALSE NA
#> 3 9.116999 NA FALSE NA
#> 4 46.149198 18.0601 FALSE TRUE
#> 5 29.128451 4.9859 FALSE TRUE
#> 6 53.705073 27.1894 FALSE TRUE
#> 7 31.155088 15.4384 FALSE TRUE
#> 8 422.578634 15.4384 FALSE TRUE
get_parameters(file.path(test_data_dir, "models", "onecmt", "run002.mod"))
#> kind name random_effect estimate sd corr stderr
#> 1 THETA TVCL <NA> 1.24679000 NA NA 0.1288330
#> 2 THETA TVV <NA> 40.84820000 NA NA 3.0272100
#> 3 THETA TVKA <NA> 1.24394000 NA NA 0.1134100
#> 4 OMEGA OM1 (TVCL) ETA1 0.13041700 0.3611320 NA 0.0601864
#> 5 OMEGA OM2 (TVV) ETA2 0.13633200 0.3692310 NA 0.0397114
#> 6 OMEGA OM3 (TVKA) ETA3 0.11440800 0.3382430 NA 0.0614429
#> 7 SIGMA SIGMA(1,1) EPS1 0.03723180 0.1929550 NA 0.0115996
#> 8 SIGMA SIGMA(2,2) EPS2 0.00660722 0.0812848 NA 0.0279207
#> rse shrinkage fixed diagonal
#> 1 10.333176 NA FALSE NA
#> 2 7.410877 NA FALSE NA
#> 3 9.116999 NA FALSE NA
#> 4 46.149198 18.0601 FALSE TRUE
#> 5 29.128451 4.9859 FALSE TRUE
#> 6 53.705073 27.1894 FALSE TRUE
#> 7 31.155088 15.4384 FALSE TRUE
#> 8 422.578634 15.4384 FALSE TRUE
get_parameters(file.path(test_data_dir, "models", "onecmt", "run002", "run002.ext"))
#> kind name random_effect estimate sd corr stderr
#> 1 THETA TVCL <NA> 1.24679000 NA NA 0.1288330
#> 2 THETA TVV <NA> 40.84820000 NA NA 3.0272100
#> 3 THETA TVKA <NA> 1.24394000 NA NA 0.1134100
#> 4 OMEGA OM1 (TVCL) ETA1 0.13041700 0.3611320 NA 0.0601864
#> 5 OMEGA OM2 (TVV) ETA2 0.13633200 0.3692310 NA 0.0397114
#> 6 OMEGA OM3 (TVKA) ETA3 0.11440800 0.3382430 NA 0.0614429
#> 7 SIGMA SIGMA(1,1) EPS1 0.03723180 0.1929550 NA 0.0115996
#> 8 SIGMA SIGMA(2,2) EPS2 0.00660722 0.0812848 NA 0.0279207
#> rse shrinkage fixed diagonal
#> 1 10.333176 NA FALSE NA
#> 2 7.410877 NA FALSE NA
#> 3 9.116999 NA FALSE NA
#> 4 46.149198 18.0601 FALSE TRUE
#> 5 29.128451 4.9859 FALSE TRUE
#> 6 53.705073 27.1894 FALSE TRUE
#> 7 31.155088 15.4384 FALSE TRUE
#> 8 422.578634 15.4384 FALSE TRUE
get_parameters(file.path(test_data_dir, "models", "onecmt", "run002_metadata.json"))
#> kind name random_effect estimate sd corr stderr
#> 1 THETA TVCL <NA> 1.24679000 NA NA 0.1288330
#> 2 THETA TVV <NA> 40.84820000 NA NA 3.0272100
#> 3 THETA TVKA <NA> 1.24394000 NA NA 0.1134100
#> 4 OMEGA OM1 (TVCL) ETA1 0.13041700 0.3611320 NA 0.0601864
#> 5 OMEGA OM2 (TVV) ETA2 0.13633200 0.3692310 NA 0.0397114
#> 6 OMEGA OM3 (TVKA) ETA3 0.11440800 0.3382430 NA 0.0614429
#> 7 SIGMA SIGMA(1,1) EPS1 0.03723180 0.1929550 NA 0.0115996
#> 8 SIGMA SIGMA(2,2) EPS2 0.00660722 0.0812848 NA 0.0279207
#> rse shrinkage fixed diagonal
#> 1 10.333176 NA FALSE NA
#> 2 7.410877 NA FALSE NA
#> 3 9.116999 NA FALSE NA
#> 4 46.149198 18.0601 FALSE TRUE
#> 5 29.128451 4.9859 FALSE TRUE
#> 6 53.705073 27.1894 FALSE TRUE
#> 7 31.155088 15.4384 FALSE TRUE
#> 8 422.578634 15.4384 FALSE TRUE
get_parameters(
file.path(test_data_dir, "models", "onecmt", "run002")
) |>
mutate(
`95% CI` = paste0(
"(",
round(estimate - 1.96 * stderr, 3)," ,",
round(estimate + 1.96 * stderr, 3),
")"
)
)
#> kind name random_effect estimate sd corr stderr
#> 1 THETA TVCL <NA> 1.24679000 NA NA 0.1288330
#> 2 THETA TVV <NA> 40.84820000 NA NA 3.0272100
#> 3 THETA TVKA <NA> 1.24394000 NA NA 0.1134100
#> 4 OMEGA OM1 (TVCL) ETA1 0.13041700 0.3611320 NA 0.0601864
#> 5 OMEGA OM2 (TVV) ETA2 0.13633200 0.3692310 NA 0.0397114
#> 6 OMEGA OM3 (TVKA) ETA3 0.11440800 0.3382430 NA 0.0614429
#> 7 SIGMA SIGMA(1,1) EPS1 0.03723180 0.1929550 NA 0.0115996
#> 8 SIGMA SIGMA(2,2) EPS2 0.00660722 0.0812848 NA 0.0279207
#> rse shrinkage fixed diagonal 95% CI
#> 1 10.333176 NA FALSE NA (0.994 ,1.499)
#> 2 7.410877 NA FALSE NA (34.915 ,46.782)
#> 3 9.116999 NA FALSE NA (1.022 ,1.466)
#> 4 46.149198 18.0601 FALSE TRUE (0.012 ,0.248)
#> 5 29.128451 4.9859 FALSE TRUE (0.058 ,0.214)
#> 6 53.705073 27.1894 FALSE TRUE (-0.006 ,0.235)
#> 7 31.155088 15.4384 FALSE TRUE (0.014 ,0.06)
#> 8 422.578634 15.4384 FALSE TRUE (-0.048 ,0.061)
untransformed_df <- get_parameters(
file.path(test_data_dir, "models", "onecmt", "run002")
)
transformed_df <- untransformed_df |>
mutate(
transform = case_when(
kind == "THETA" ~ "Identity",
kind == "OMEGA" & diagonal ~ "LogNormal",
kind == "OMEGA" & !diagonal ~ "Identity",
name == "SIGMA(1,1)" ~ "Proportional",
name == "SIGMA(2,2)" ~ "AddErr",
.default = "Identity"
),
cv = compute_cv(estimate, kind, transform),
rse = compute_rse(estimate, stderr, kind, transform),
ci_low = compute_ci(estimate, stderr, 0.95, transform)$lower,
ci_high = compute_ci(estimate, stderr, 0.95, transform)$upper,
)
transformed_df
#> kind name random_effect estimate sd corr stderr
#> 1 THETA TVCL <NA> 1.24679000 NA NA 0.1288330
#> 2 THETA TVV <NA> 40.84820000 NA NA 3.0272100
#> 3 THETA TVKA <NA> 1.24394000 NA NA 0.1134100
#> 4 OMEGA OM1 (TVCL) ETA1 0.13041700 0.3611320 NA 0.0601864
#> 5 OMEGA OM2 (TVV) ETA2 0.13633200 0.3692310 NA 0.0397114
#> 6 OMEGA OM3 (TVKA) ETA3 0.11440800 0.3382430 NA 0.0614429
#> 7 SIGMA SIGMA(1,1) EPS1 0.03723180 0.1929550 NA 0.0115996
#> 8 SIGMA SIGMA(2,2) EPS2 0.00660722 0.0812848 NA 0.0279207
#> rse shrinkage fixed diagonal transform cv ci_low
#> 1 10.333176 NA FALSE NA Identity NA 0.99428196
#> 2 7.410877 NA FALSE NA Identity NA 34.91497743
#> 3 9.116999 NA FALSE NA Identity NA 1.02166048
#> 4 46.149198 18.0601 FALSE TRUE LogNormal 37.32337 1.01253170
#> 5 29.128451 4.9859 FALSE TRUE LogNormal 38.21810 1.06024402
#> 6 53.705073 27.1894 FALSE TRUE LogNormal 34.81515 0.99400020
#> 7 31.155088 15.4384 FALSE TRUE Proportional 19.29554 0.01449700
#> 8 422.578634 15.4384 FALSE TRUE AddErr NA -0.04811635
#> ci_high
#> 1 1.49929804
#> 2 46.78142257
#> 3 1.46621952
#> 4 1.28194721
#> 5 1.23882694
#> 6 1.26469865
#> 7 0.05996660
#> 8 0.06133079
Read ext file
read_ext_file(
file.path(test_data_dir, "ext", "bql.ext"),
line_prefixes = "-1000000000",
parameters_only = TRUE
)
#> iteration method THETA1 THETA2 THETA3 THETA4 THETA5 THETA6 THETA7
#> 1 -1000000000 FOCE 26.4911 282.616 297.044 58.7497 1.50949 0.75 1
#> THETA8 THETA9 SIGMA.1.1. OMEGA.1.1. OMEGA.2.1. OMEGA.2.2. OMEGA.3.1.
#> 1 1 0.75 0.00245104 0.100615 0 0.0360015 0
#> OMEGA.3.2. OMEGA.3.3.
#> 1 0 0.0111728
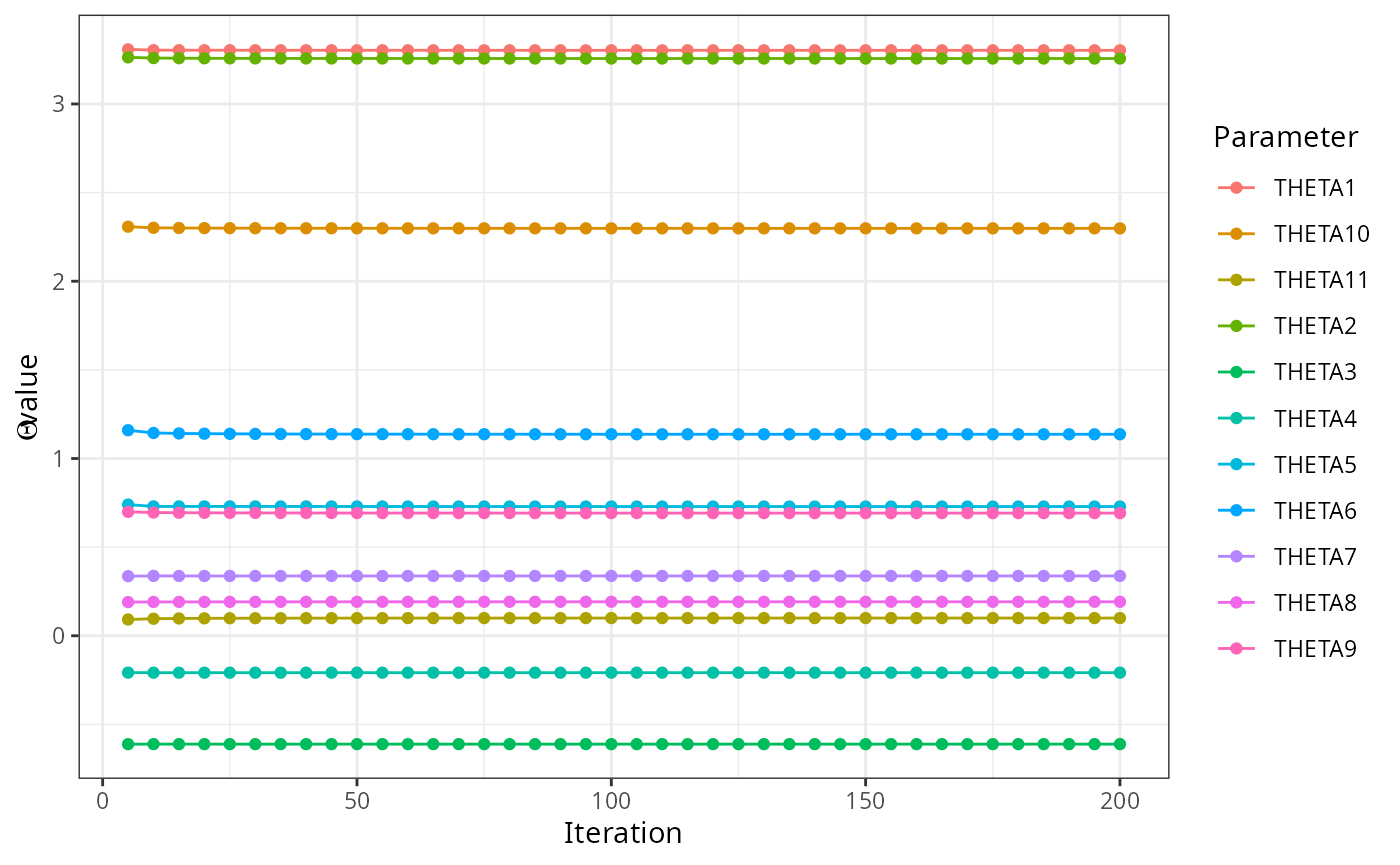
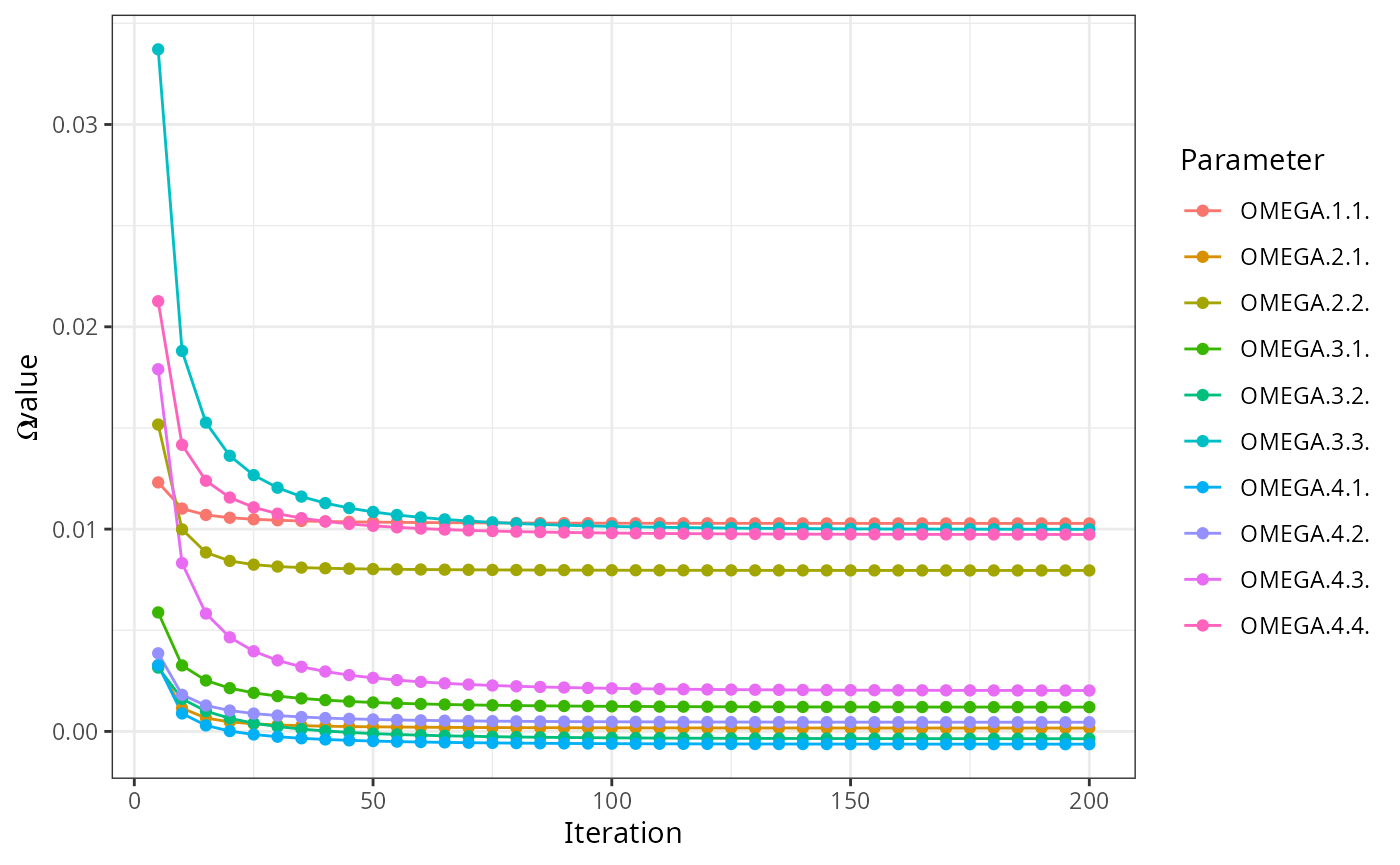
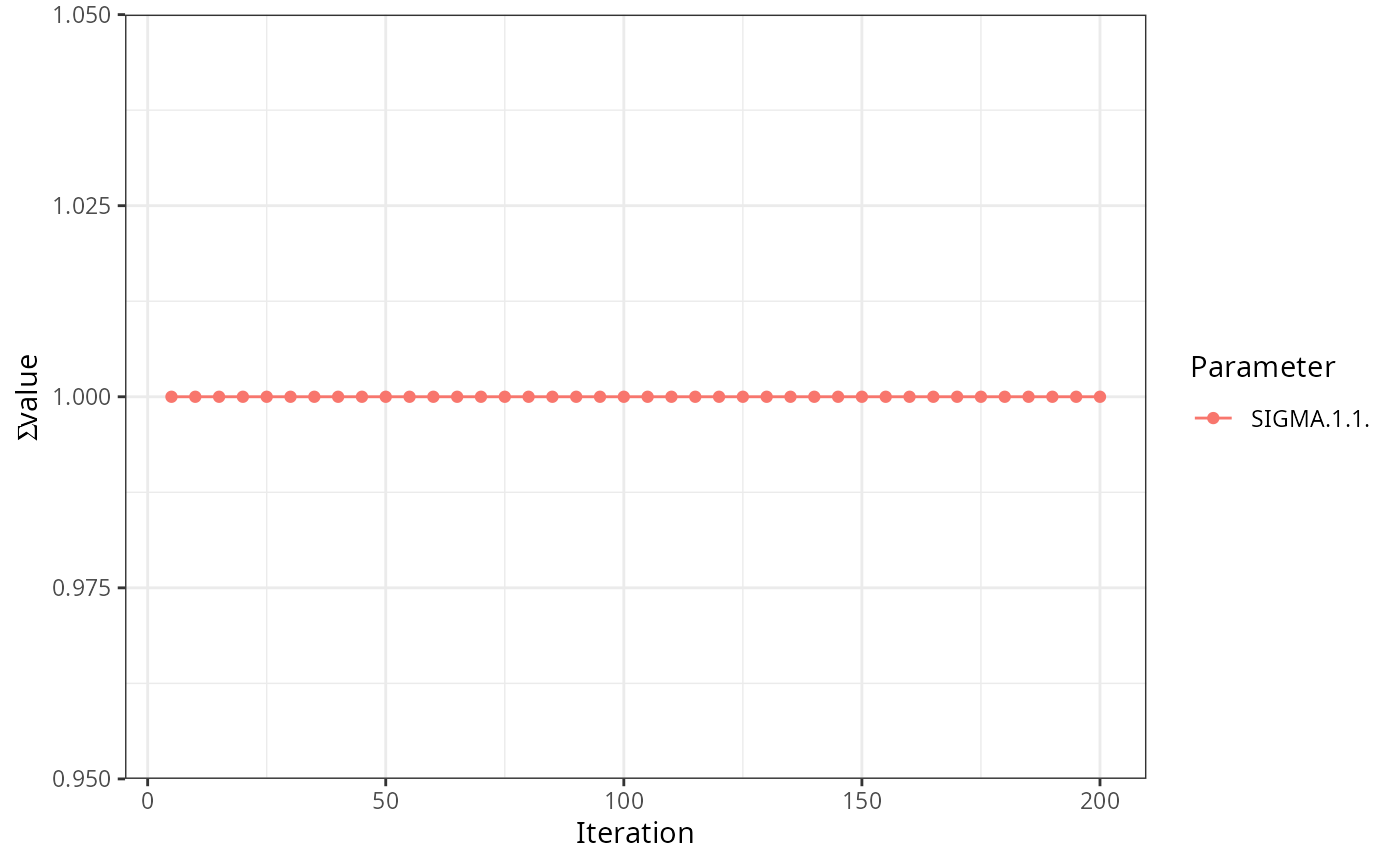
read_ext_file(file.path(test_data_dir, "ext", "example6.txt.ext")) |>
filter(iteration == -1000000000)
#> iteration method ITERATION THETA1 THETA2 THETA3 THETA4 THETA5
#> 1 -1000000000 Bayes -1e+09 3.90686 -2.21701 0.552837 -0.183461 2.26824
#> THETA6 THETA7 THETA8 SIGMA.1.1. SIGMA.2.1. SIGMA.2.2. OMEGA.1.1.
#> 1 0.238698 3.71188 -0.703886 0.00931609 0 0.0223709 0.281875
#> OMEGA.2.1. OMEGA.2.2. OMEGA.3.1. OMEGA.3.2. OMEGA.3.3. OMEGA.4.1. OMEGA.4.2.
#> 1 -0.0347915 0.213765 0.0451677 -0.00980444 0.14051 0.030207 0.0548359
#> OMEGA.4.3. OMEGA.4.4. OMEGA.5.1. OMEGA.5.2. OMEGA.5.3. OMEGA.5.4.
#> 1 -0.0143056 0.26165 0.0272446 0.0164947 -0.000745322 -0.0322041
#> OMEGA.5.5. OMEGA.6.1. OMEGA.6.2. OMEGA.6.3. OMEGA.6.4. OMEGA.6.5. OMEGA.6.6.
#> 1 0.206086 -0.0268258 0.00474677 0.0158401 0.0146191 -0.0737751 0.234941
#> OMEGA.7.1. OMEGA.7.2. OMEGA.7.3. OMEGA.7.4. OMEGA.7.5. OMEGA.7.6.
#> 1 0.0297134 -0.0475722 0.0318595 -0.0738601 0.0236572 -0.000315573
#> OMEGA.7.7. OMEGA.8.1. OMEGA.8.2. OMEGA.8.3. OMEGA.8.4. OMEGA.8.5. OMEGA.8.6.
#> 1 0.247069 0.0958203 0.0735577 0.0413009 0.0461165 0.00398651 -0.051391
#> OMEGA.8.7. OMEGA.8.8. MCMCOBJ
#> 1 0.0573565 0.237051 -6490.95


