Visualizing error.
Usage
ggerrorplot(
data,
x,
y,
desc_stat = "mean_se",
numeric.x.axis = FALSE,
combine = FALSE,
merge = FALSE,
color = "black",
fill = "white",
palette = NULL,
size = NULL,
width = NULL,
title = NULL,
xlab = NULL,
ylab = NULL,
facet.by = NULL,
panel.labs = NULL,
short.panel.labs = TRUE,
select = NULL,
remove = NULL,
order = NULL,
add = "none",
add.params = list(),
error.plot = "pointrange",
ci = 0.95,
position = position_dodge(),
ggtheme = theme_pubr(),
...
)Arguments
- data
a data frame
- x, y
x and y variables for drawing.
- desc_stat
descriptive statistics to be used for visualizing errors. Default value is "mean_se". Allowed values are one of , "mean", "mean_se", "mean_sd", "mean_ci", "mean_range", "median", "median_iqr", "median_hilow", "median_q1q3", "median_mad", "median_range"; see
desc_statbyfor more details.- numeric.x.axis
logical. If TRUE, x axis will be treated as numeric. Default is FALSE.
- combine
logical value. Default is FALSE. Used only when y is a vector containing multiple variables to plot. If TRUE, create a multi-panel plot by combining the plot of y variables.
- merge
logical or character value. Default is FALSE. Used only when y is a vector containing multiple variables to plot. If TRUE, merge multiple y variables in the same plotting area. Allowed values include also "asis" (TRUE) and "flip". If merge = "flip", then y variables are used as x tick labels and the x variable is used as grouping variable.
- color, fill
outline and fill colors.
- palette
the color palette to be used for coloring or filling by groups. Allowed values include "grey" for grey color palettes; brewer palettes e.g. "RdBu", "Blues", ...; or custom color palette e.g. c("blue", "red"); and scientific journal palettes from ggsci R package, e.g.: "npg", "aaas", "lancet", "jco", "ucscgb", "uchicago", "simpsons" and "rickandmorty".
- size
Numeric value (e.g.: size = 1). change the size of points and outlines.
- width
numeric value between 0 and 1 specifying box width.
- title
plot main title.
- xlab
character vector specifying x axis labels. Use xlab = FALSE to hide xlab.
- ylab
character vector specifying y axis labels. Use ylab = FALSE to hide ylab.
- facet.by
character vector, of length 1 or 2, specifying grouping variables for faceting the plot into multiple panels. Should be in the data.
- panel.labs
a list of one or two character vectors to modify facet panel labels. For example, panel.labs = list(sex = c("Male", "Female")) specifies the labels for the "sex" variable. For two grouping variables, you can use for example panel.labs = list(sex = c("Male", "Female"), rx = c("Obs", "Lev", "Lev2") ).
- short.panel.labs
logical value. Default is TRUE. If TRUE, create short labels for panels by omitting variable names; in other words panels will be labelled only by variable grouping levels.
- select
character vector specifying which items to display.
- remove
character vector specifying which items to remove from the plot.
- order
character vector specifying the order of items. Considered only when x axis is a factor variable.
- add
character vector for adding another plot element (e.g.: dot plot or error bars). Allowed values are one or the combination of: "none", "dotplot", "jitter", "boxplot", "point", "mean", "mean_se", "mean_sd", "mean_ci", "mean_range", "median", "median_iqr", "median_hilow", "median_q1q3", "median_mad", "median_range"; see ?desc_statby for more details.
- add.params
parameters (color, shape, size, fill, linetype) for the argument 'add'; e.g.: add.params = list(color = "red").
- error.plot
plot type used to visualize error. Allowed values are one of c("pointrange", "linerange", "crossbar", "errorbar", "upper_errorbar", "lower_errorbar", "upper_pointrange", "lower_pointrange", "upper_linerange", "lower_linerange"). Default value is "pointrange" or "errorbar". Used only when add != "none" and add contains one "mean_*" or "med_*" where "*" = sd, se, ....
- ci
the percent range of the confidence interval (default is 0.95).
- position
A position adjustment to use on the data for this layer. This can be used in various ways, including to prevent overplotting and improving the display. The
positionargument accepts the following:The result of calling a position function, such as
position_jitter(). This method allows for passing extra arguments to the position.A string naming the position adjustment. To give the position as a string, strip the function name of the
position_prefix. For example, to useposition_jitter(), give the position as"jitter".For more information and other ways to specify the position, see the layer position documentation.
- ggtheme
function, ggplot2 theme name. Default value is theme_pubr(). Allowed values include ggplot2 official themes: theme_gray(), theme_bw(), theme_minimal(), theme_classic(), theme_void(), ....
- ...
other arguments to be passed to be passed to ggpar().
Details
The plot can be easily customized using the function ggpar(). Read ?ggpar for changing:
main title and axis labels: main, xlab, ylab
axis limits: xlim, ylim (e.g.: ylim = c(0, 30))
axis scales: xscale, yscale (e.g.: yscale = "log2")
color palettes: palette = "Dark2" or palette = c("gray", "blue", "red")
legend title, labels and position: legend = "right"
plot orientation : orientation = c("vertical", "horizontal", "reverse")
Examples
# Data: ToothGrowth data set we'll be used.
df<- ToothGrowth
head(df, 10)
#> len supp dose
#> 1 4.2 VC 0.5
#> 2 11.5 VC 0.5
#> 3 7.3 VC 0.5
#> 4 5.8 VC 0.5
#> 5 6.4 VC 0.5
#> 6 10.0 VC 0.5
#> 7 11.2 VC 0.5
#> 8 11.2 VC 0.5
#> 9 5.2 VC 0.5
#> 10 7.0 VC 0.5
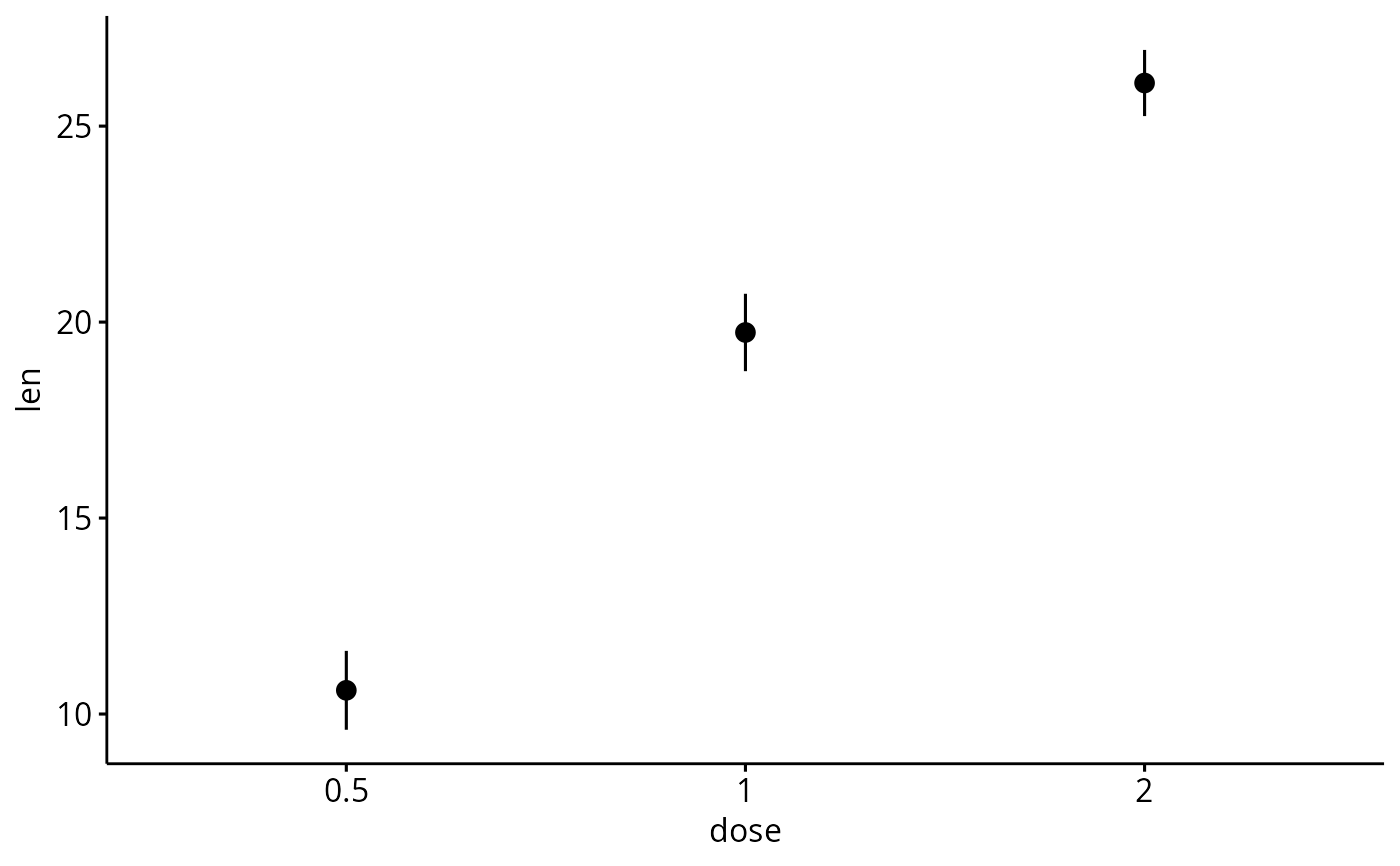
# Plot mean_se
ggerrorplot(df, x = "dose", y = "len")
 # Change desc_stat to mean_sd
# (other values include: mean_sd, mean_ci, median_iqr, ....)
# Add labels
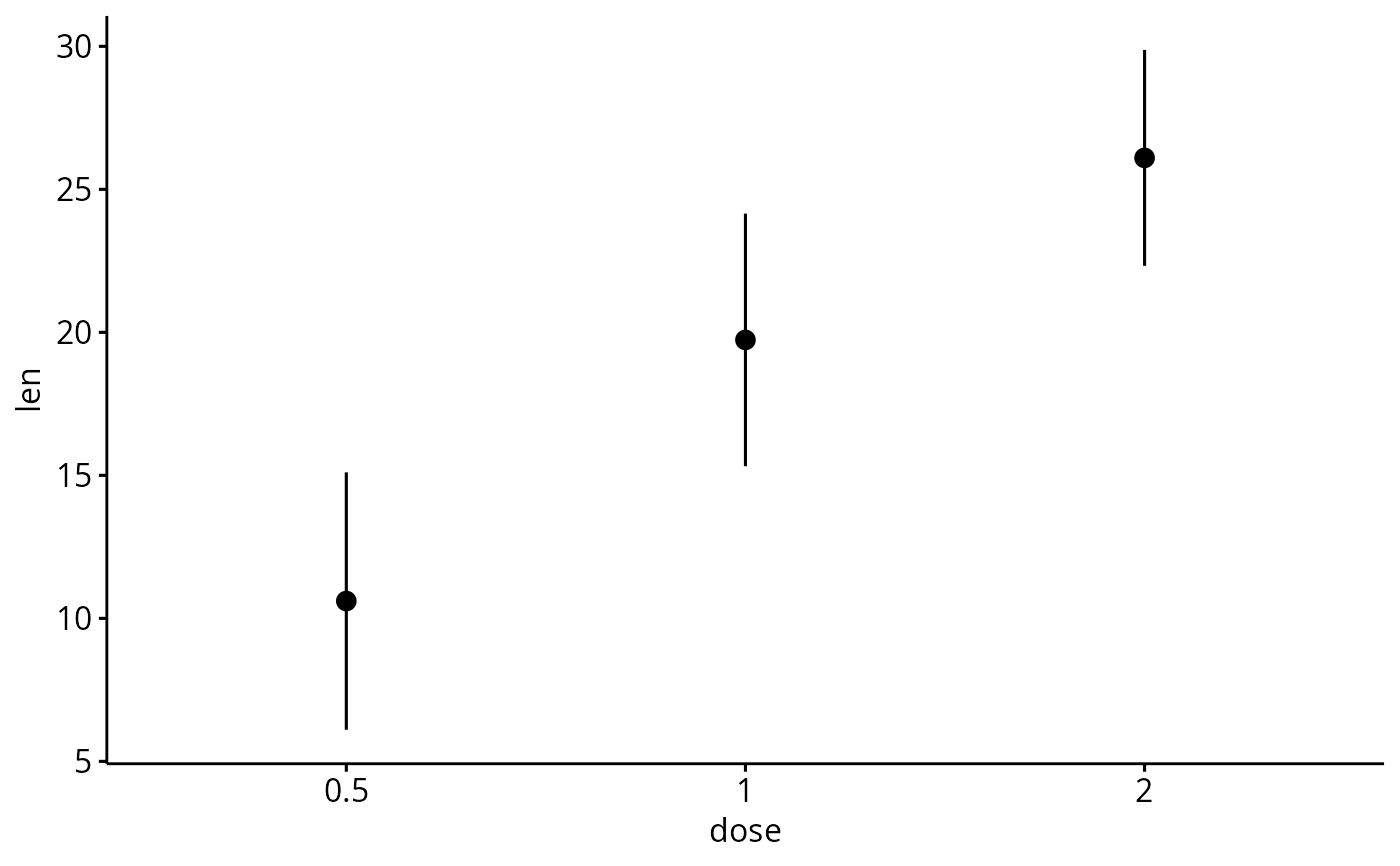
ggerrorplot(df, x = "dose", y = "len",
desc_stat = "mean_sd")
# Change desc_stat to mean_sd
# (other values include: mean_sd, mean_ci, median_iqr, ....)
# Add labels
ggerrorplot(df, x = "dose", y = "len",
desc_stat = "mean_sd")
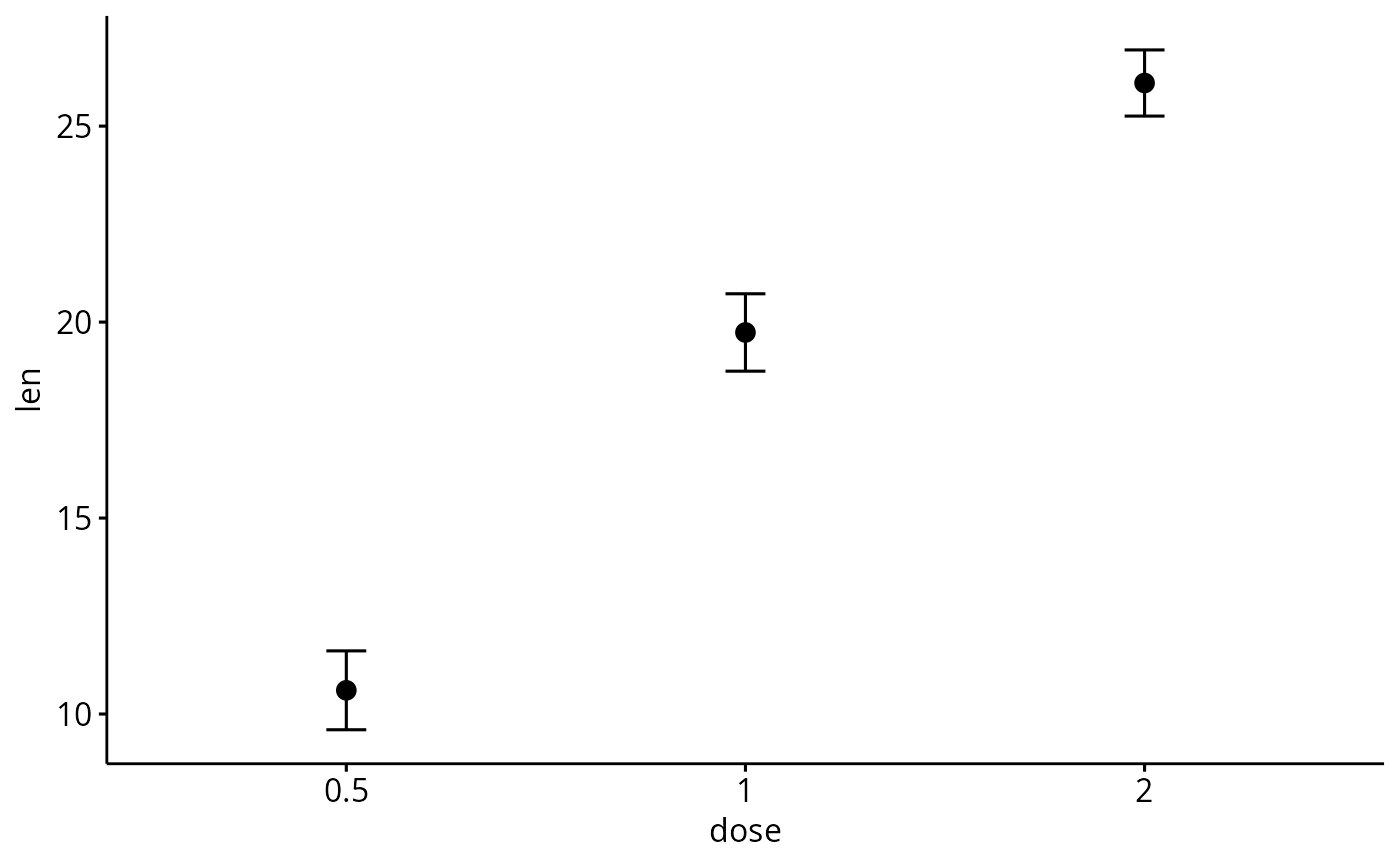
 # Change error.plot to "errorbar" and add mean point
# Visualize the mean of each group
ggerrorplot(df, x = "dose", y = "len",
add = "mean", error.plot = "errorbar")
# Change error.plot to "errorbar" and add mean point
# Visualize the mean of each group
ggerrorplot(df, x = "dose", y = "len",
add = "mean", error.plot = "errorbar")
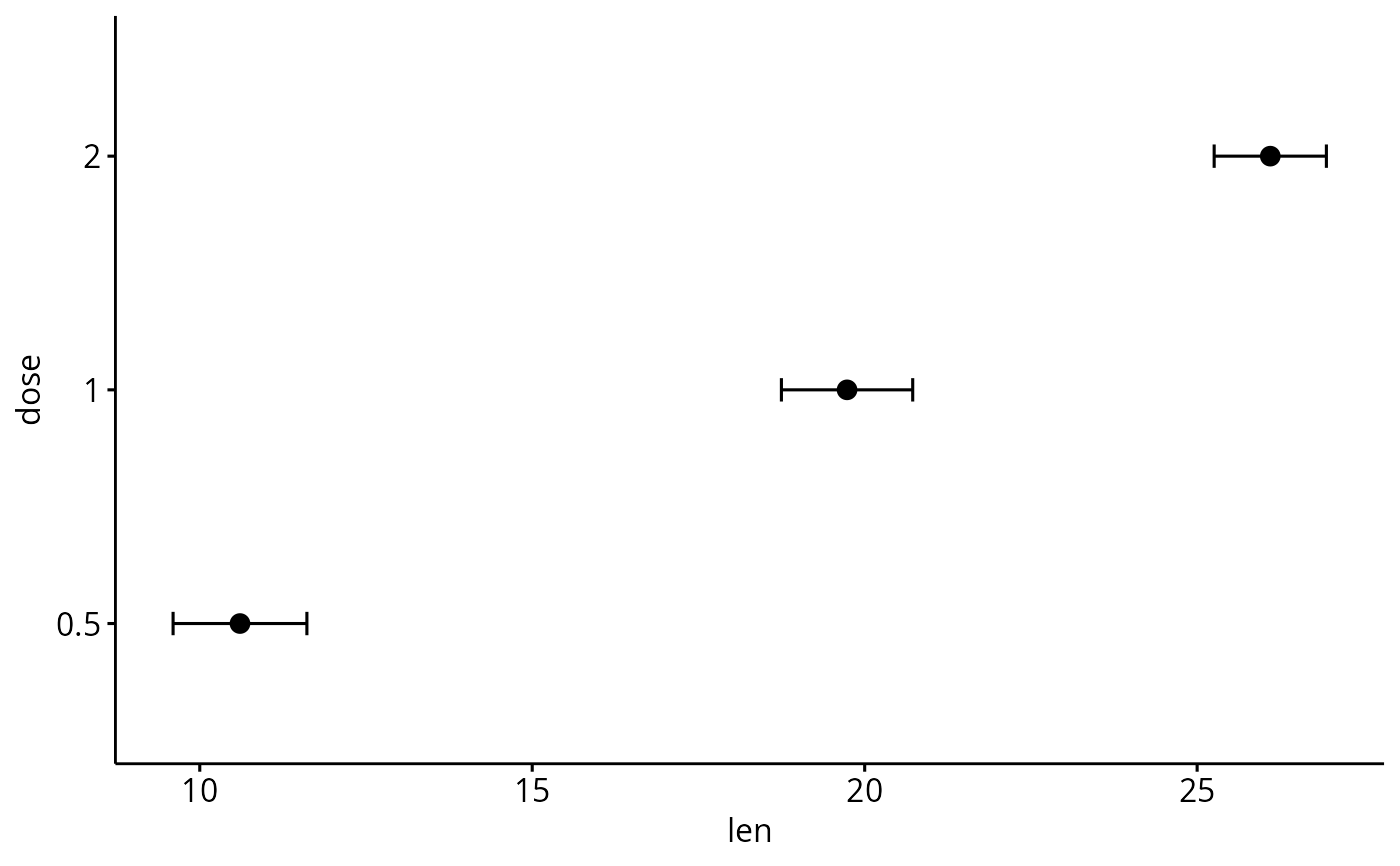
 # Horizontal plot
ggerrorplot(df, x = "dose", y = "len",
add = "mean", error.plot = "errorbar",
orientation = "horizontal")
# Horizontal plot
ggerrorplot(df, x = "dose", y = "len",
add = "mean", error.plot = "errorbar",
orientation = "horizontal")
 # Change error.plot to "crossbar"
ggerrorplot(df, x = "dose", y = "len",
error.plot = "crossbar", width = 0.5)
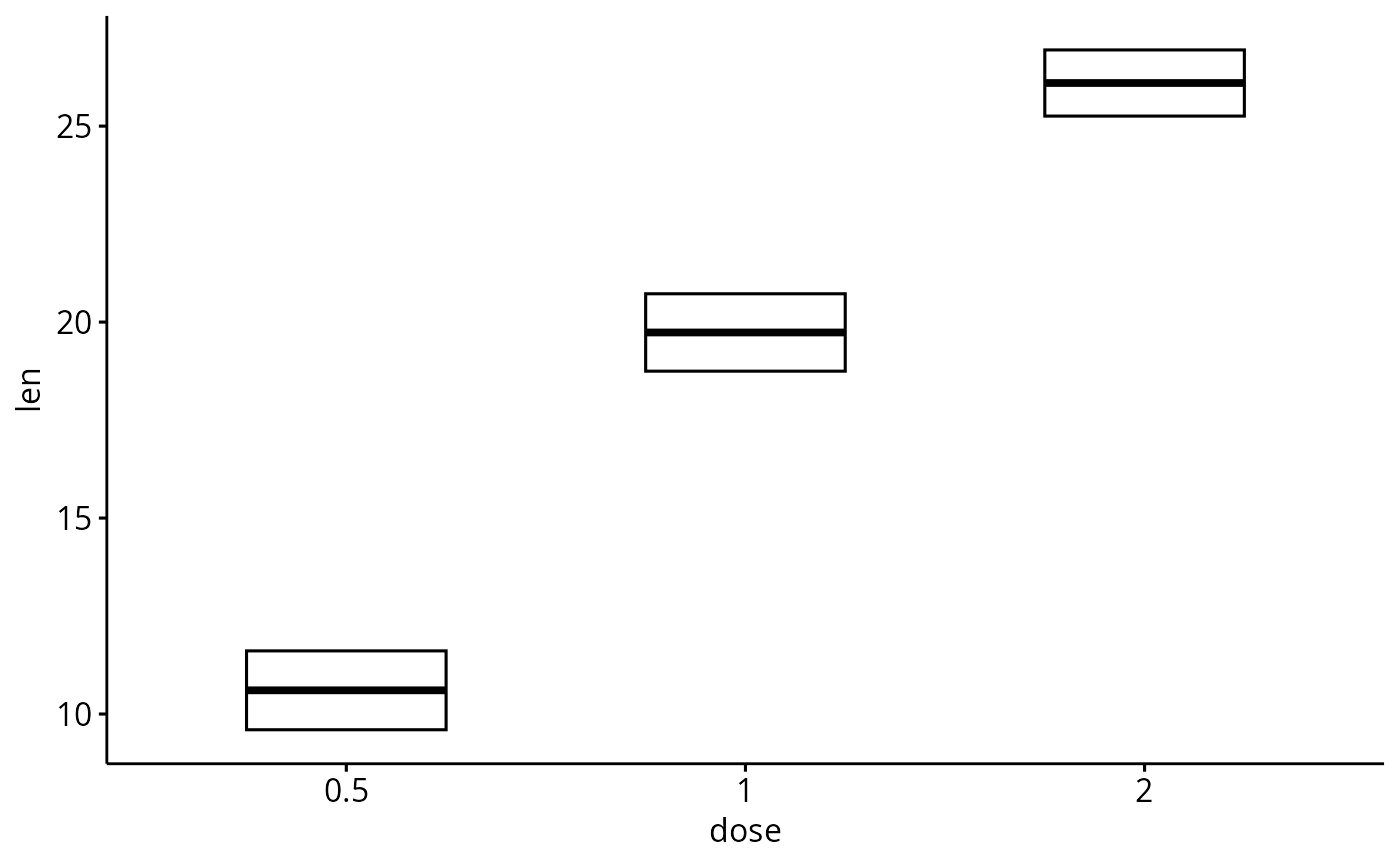
# Change error.plot to "crossbar"
ggerrorplot(df, x = "dose", y = "len",
error.plot = "crossbar", width = 0.5)
 # Add jitter points and errors (mean_se)
ggerrorplot(df, x = "dose", y = "len",
add = "jitter")
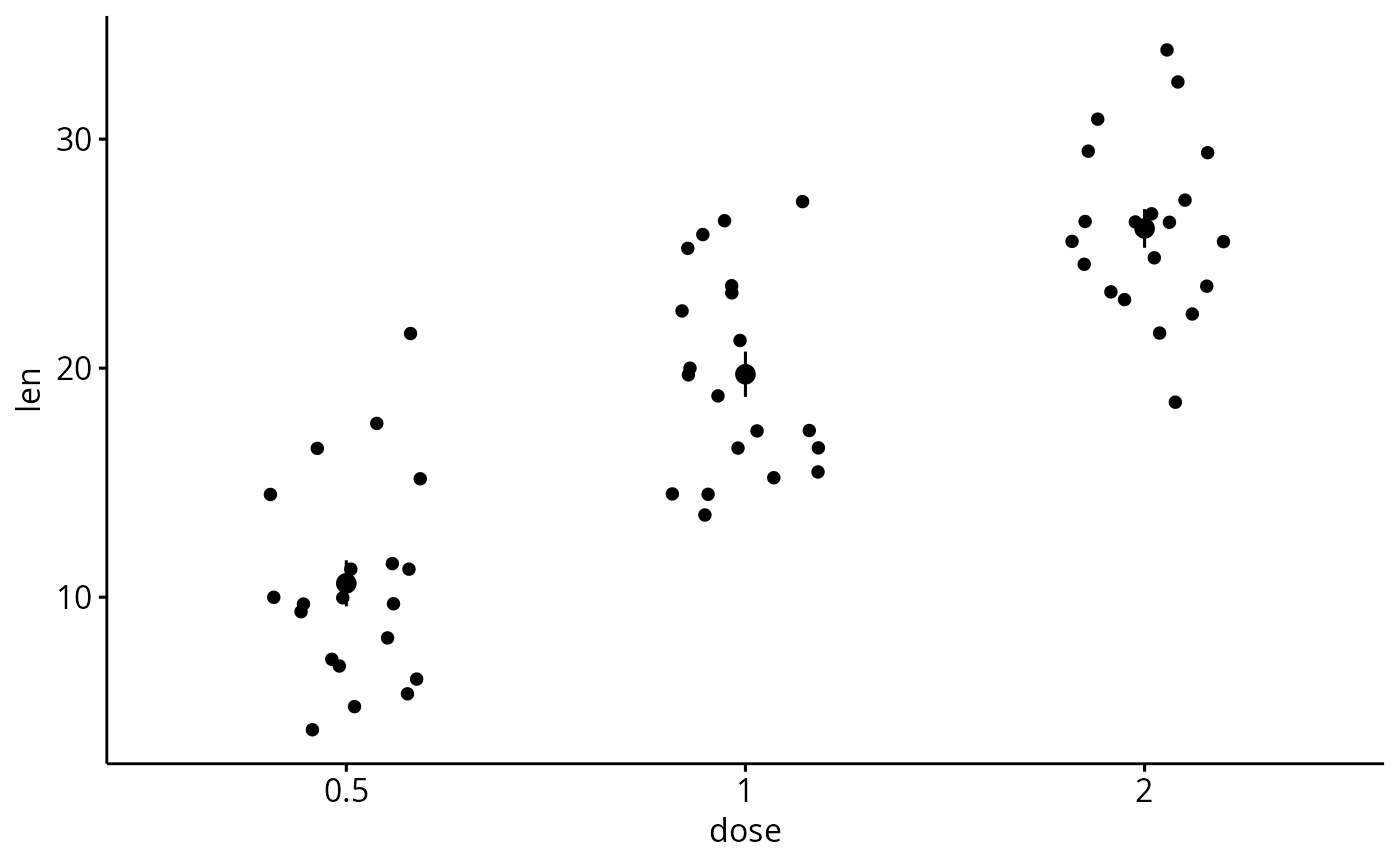
# Add jitter points and errors (mean_se)
ggerrorplot(df, x = "dose", y = "len",
add = "jitter")
 # Add dot and errors (mean_se)
ggerrorplot(df, x = "dose", y = "len",
add = "dotplot")
#> Bin width defaults to 1/30 of the range of the data. Pick better value with
#> `binwidth`.
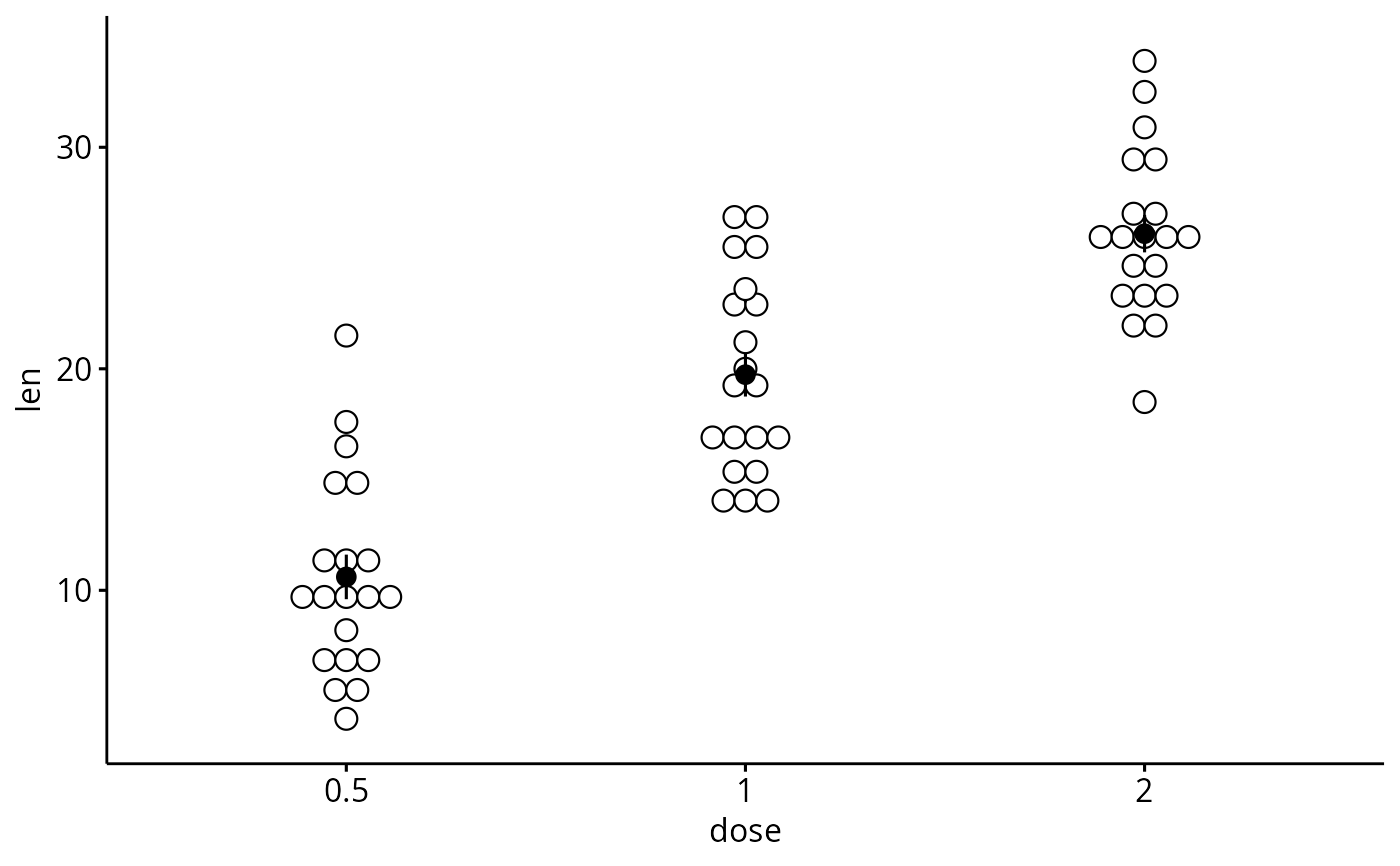
# Add dot and errors (mean_se)
ggerrorplot(df, x = "dose", y = "len",
add = "dotplot")
#> Bin width defaults to 1/30 of the range of the data. Pick better value with
#> `binwidth`.
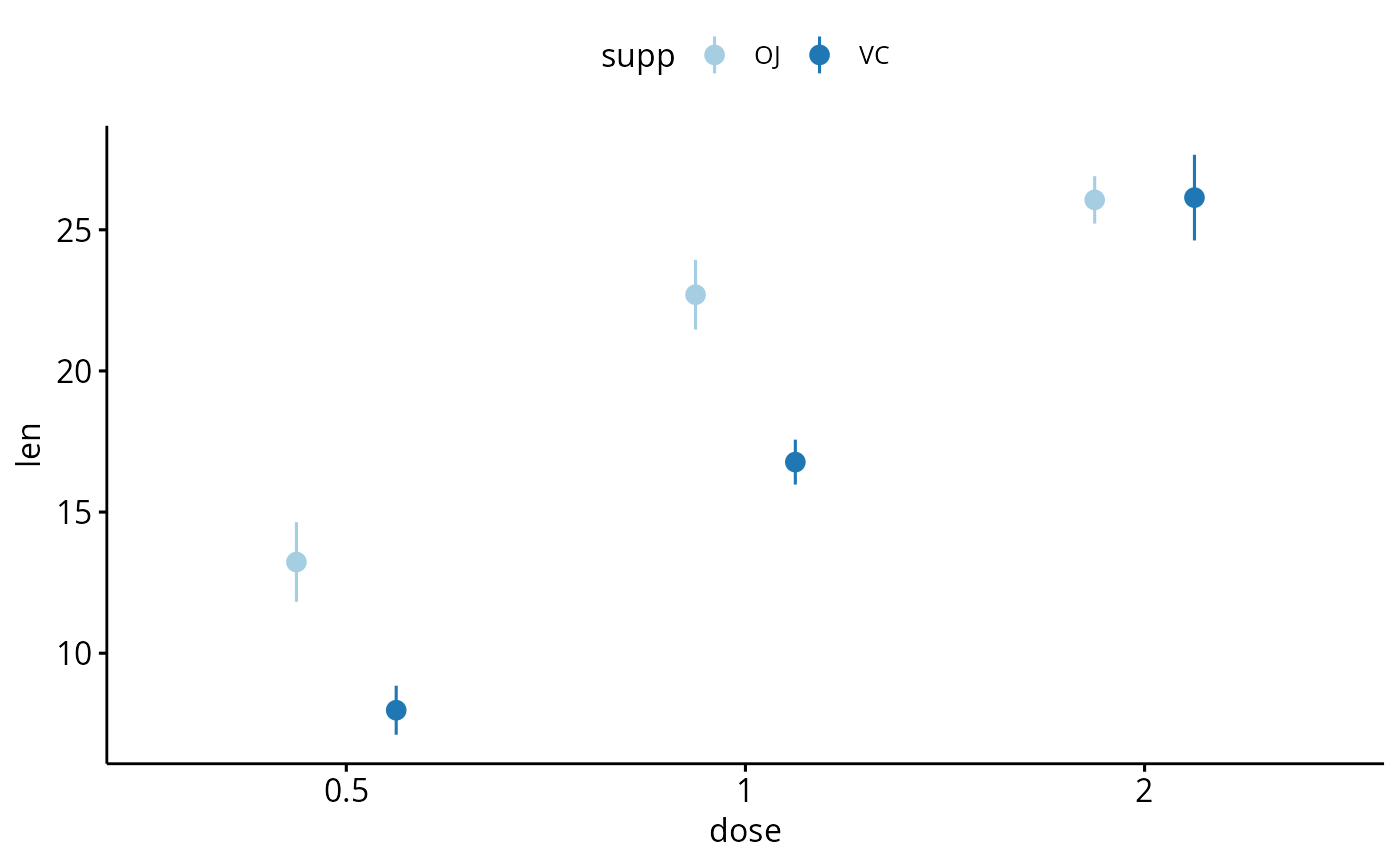
 # Multiple groups with error bars and jitter point
ggerrorplot(df, x = "dose", y = "len",
color = "supp", palette = "Paired",
error.plot = "pointrange",
position = position_dodge(0.5))
# Multiple groups with error bars and jitter point
ggerrorplot(df, x = "dose", y = "len",
color = "supp", palette = "Paired",
error.plot = "pointrange",
position = position_dodge(0.5))