Equation, p-value, \(R^2\), AIC and BIC of fitted polynomial
Source:R/stat-poly-eq.R
stat_poly_eq.Rdstat_poly_eq fits a polynomial, by default with stats::lm(),
but alternatively using robust regression or generalized least squares. Using
the fitted model it generates several labels including the fitted model
equation, p-value, F-value, coefficient of determination (R^2), 'AIC', 'BIC',
number of observations and method name, if available.
stat_poly_eq(
mapping = NULL,
data = NULL,
geom = "text_npc",
position = "identity",
...,
formula = NULL,
method = "lm",
method.args = list(),
n.min = 2L,
eq.with.lhs = TRUE,
eq.x.rhs = NULL,
small.r = getOption("ggpmisc.small.r", default = FALSE),
small.p = getOption("ggpmisc.small.p", default = FALSE),
CI.brackets = c("[", "]"),
rsquared.conf.level = 0.95,
coef.digits = 3,
coef.keep.zeros = TRUE,
decreasing = getOption("ggpmisc.decreasing.poly.eq", FALSE),
rr.digits = 2,
f.digits = 3,
p.digits = 3,
label.x = "left",
label.y = "top",
hstep = 0,
vstep = NULL,
output.type = NULL,
na.rm = FALSE,
orientation = NA,
parse = NULL,
show.legend = FALSE,
inherit.aes = TRUE
)Arguments
- mapping
The aesthetic mapping, usually constructed with
aes. Only needs to be set at the layer level if you are overriding the plot defaults.- data
A layer specific dataset, only needed if you want to override the plot defaults.
- geom
The geometric object to use display the data
- position
The position adjustment to use for overlapping points on this layer
- ...
other arguments passed on to
layer. This can include aesthetics whose values you want to set, not map. Seelayerfor more details.- formula
a formula object. Using aesthetic names
xandyinstead of original variable names.- method
function or character If character, "lm", "rlm", "lqs". "gls" or the name of a model fit function are accepted, possibly followed by the fit function's
methodargument separated by a colon (e.g."rlm:M"). If a function is different tolm(),rlm(),lqs()orgls(), it must have formal parameters namedformula,data,weights, andmethod, and return a model fit object of class"lm", class"lqs"or class"gls".- method.args
named list with additional arguments.
- n.min
integer Minimum number of distinct values in the explanatory variable (on the rhs of formula) for fitting to the attempted.
- eq.with.lhs
If
characterthe string is pasted to the front of the equation label before parsing or alogical(see note).- eq.x.rhs
characterthis string will be used as replacement for"x"in the model equation when generating the label before parsing it.- small.r, small.p
logical Flags to switch use of lower case r and p for coefficient of determination and p-value.
- CI.brackets
character vector of length 2. The opening and closing brackets used for the CI label.
- rsquared.conf.level
numeric Confidence level for the returned confidence interval. Set to NA to skip CI computation.
- coef.digits, f.digits
integer Number of significant digits to use for the fitted coefficients and F-value.
- coef.keep.zeros
logical Keep or drop trailing zeros when formatting the fitted coefficients and F-value.
- decreasing
logical It specifies the order of the terms in the returned character string; in increasing (default) or decreasing powers.
- rr.digits, p.digits
integer Number of digits after the decimal point to use for \(R^2\) and P-value in labels. If
Inf, use exponential notation with three decimal places.- label.x, label.y
numericwith range 0..1 "normalized parent coordinates" (npc units) or character if usinggeom_text_npc()orgeom_label_npc(). If usinggeom_text()orgeom_label()numeric in native data units. If too short they will be recycled.- hstep, vstep
numeric in npc units, the horizontal and vertical step used between labels for different groups.
- output.type
character One of "expression", "LaTeX", "text", "markdown" or "numeric".
- na.rm
a logical indicating whether NA values should be stripped before the computation proceeds.
- orientation
character Either "x" or "y" controlling the default for
formula.- parse
logical Passed to the geom. If
TRUE, the labels will be parsed into expressions and displayed as described in?plotmath. Default isTRUEifoutput.type = "expression"andFALSEotherwise.- show.legend
logical. Should this layer be included in the legends?
NA, the default, includes if any aesthetics are mapped.FALSEnever includes, andTRUEalways includes.- inherit.aes
If
FALSE, overrides the default aesthetics, rather than combining with them. This is most useful for helper functions that define both data and aesthetics and shouldn't inherit behaviour from the default plot specification, e.g.borders.
Value
A data frame, with a single row and columns as described under
Computed variables. In cases when the number of observations is
less than n.min a data frame with no rows or columns is returned,
and rendered as an empty/invisible plot layer.
Details
This statistic can be used to automatically annotate a plot with
\(R^2\), adjusted \(R^2\) or the fitted model equation. It supports
linear regression and polynomial fits, and robust regression fitted
with functions lm, or rlm, respectively.
While strings for \(R^2\), adjusted \(R^2\), \(F\), and \(P\)
annotations are returned for all valid linear models, A character string
for the fitted model is returned only for polynomials (see below), in which
case the equation can still be assembled by the user. In addition, a label
for the confidence interval of \(R^2\), based on values computed with
function ci_rsquared from package 'confintr' is
also returned.
The model formula should be defined based on the names of aesthetics x
and y, not the names of the variables in the data. Before fitting
the model, data are split based on groupings created by any other mappings
present in a plot panel: fitting is done separately for each group
in each plot panel.
Model formulas can use poly() or be defined algebraically including
the intercept indicated by +1, -1, +0 or implicit. If
defined using poly() the argument raw = TRUE must be passed.
The model formula is checked, and if not recognized as a polynomial
with no missing terms and terms ordered by increasing powers, no equation
label is generated. Thus, as the value returned for eq.label can be
NA, the default aesthetic mapping to label is \(R^2\).
By default, the character strings are generated as suitable for parsing into R's
plotmath expressions. However, LaTeX (use TikZ device), markdown (use package
'ggtext') and plain text are also supported, as well as returning numeric values for
user-generated text labels. The argument of parse is set automatically
based on output-type, but if you assemble labels that need parsing
from numeric output, the default needs to be overridden.
This statistic only generates annotation labels, the predicted values/line
need to be added to the plot as a separate layer using
stat_poly_line (or stat_smooth). Using
the same formula in stat_poly_line() and in stat_poly_eq() in
most cases ensures that the plotted curve and equation are consistent.
Thus, unless the default formula is not overriden, it is best to save the
model formula as an object and supply this named object as argument to the
two statistics.
A ggplot statistic receives as data a data frame that is not the one
passed as argument by the user, but instead a data frame with the variables
mapped to aesthetics. stat_poly_eq() mimics how stat_smooth()
works.
With method "lm", singularity results in terms being dropped with a
message if more numerous than can be fitted with a singular (exact) fit.
In this case or if the model results in a perfect fit due to a low
number of observations, estimates for various parameters are NaN or
NA. When this is the case the corresponding labels are set to
character(0L) and thus not visible in the plot.
With methods other than "lm", the model fit functions simply fail
in case of singularity, e.g., singular fits are not implemented in
"rlm".
In both cases the minimum number of observations with distinct values in
the explanatory variable can be set through parameter n.min. The
default n.min = 2L is the smallest suitable for method "lm"
but too small for method "rlm" for which n.min = 3L is
needed. Anyway, model fits with very few observations are of little
interest and using larger values of n.min than the default is
usually wise.
Note
For backward compatibility a logical is accepted as argument for
eq.with.lhs. If TRUE, the default is used, either
"x" or "y", depending on the argument passed to formula.
However, "x" or "y" can be substituted by providing a
suitable replacement character string through eq.x.rhs.
Parameter orientation is redundant as it only affects the default
for formula but is included for consistency with
ggplot2::stat_smooth().
R option OutDec is obeyed based on its value at the time the plot
is rendered, i.e., displayed or printed. Set options(OutDec = ",")
for languages like Spanish or French.
User-defined methods
User-defined functions can be passed as
argument to method. The requirements are 1) that the signature is
similar to that of function lm() (with parameters formula,
data, weights and any other arguments passed by name through
method.args) and 2) that the value returned by the function is an
object of class "lm" or an atomic NA value.
The formula used to build the equation label is extracted from the
returned "lm" object and can safely differ from the argument passed to
parameter formula in the call to stat_poly_eq(). Thus,
user-defined methods can implement both model selection or conditional
skipping of labelling.
Aesthetics
stat_poly_eq() understands x and y,
to be referenced in the formula and weight passed as argument
to parameter weights. All three must be mapped to numeric
variables. In addition, the aesthetics understood by the geom
("text" is the default) are understood and grouping respected.
If the model formula includes a transformation of x, a
matching argument should be passed to parameter eq.x.rhs
as its default value "x" will not reflect the applied
transformation. In plots, transformation should never be applied to the
left hand side of the model formula, but instead in the mapping of the
variable within aes, as otherwise plotted observations and fitted
curve will not match. In this case it may be necessary to also pass
a matching argument to parameter eq.with.lhs.
Computed variables
If output.type different from "numeric" the returned tibble contains
columns listed below. If the model fit function used does not return a value,
the label is set to character(0L).
- x,npcx
x position
- y,npcy
y position
- eq.label
equation for the fitted polynomial as a character string to be parsed or
NA- rr.label
\(R^2\) of the fitted model as a character string to be parsed
- adj.rr.label
Adjusted \(R^2\) of the fitted model as a character string to be parsed
- rr.confint.label
Confidence interval for \(R^2\) of the fitted model as a character string to be parsed
- f.value.label
F value and degrees of freedom for the fitted model as a whole.
- p.value.label
P-value for the F-value above.
- AIC.label
AIC for the fitted model.
- BIC.label
BIC for the fitted model.
- n.label
Number of observations used in the fit.
- grp.label
Set according to mapping in
aes.- method.label
Set according
methodused.- r.squared, adj.r.squared, p.value, n
numeric values, from the model fit object
If output.type is "numeric" the returned tibble contains columns
listed below. If the model fit function used does not return a value,
the variable is set to NA_real_.
- x,npcx
x position
- y,npcy
y position
- coef.ls
list containing the "coefficients" matrix from the summary of the fit object
- r.squared, rr.confint.level, rr.confint.low, rr.confint.high, adj.r.squared, f.value, f.df1, f.df2, p.value, AIC, BIC, n
numeric values, from the model fit object
- grp.label
Set according to mapping in
aes.- b_0.constant
TRUE is polynomial is forced through the origin
- b_i
One or columns with the coefficient estimates
To explore the computed values returned for a given input we suggest the use
of geom_debug as shown in the last examples below.
Alternatives
stat_regline_equation() in package 'ggpubr' is
a renamed but almost unchanged copy of stat_poly_eq() taken from an
old version of this package (without acknowledgement of source and
authorship). stat_regline_equation() lacks important functionality
and contains bugs that have been fixed in stat_poly_eq().
References
Originally written as an answer to question 7549694 at Stackoverflow but enhanced based on suggestions from users and my own needs.
See also
This statistics fits a model with function lm,
function rlm or a user supplied function returning an
object of class "lm". Consult the documentation of these functions
for the details and additional arguments that can be passed to them by name
through parameter method.args.
For quantile regression stat_quant_eq should be used instead
of stat_poly_eq while for model II or major axis regression
stat_ma_eq should be used. For other types of models such as
non-linear models, statistics stat_fit_glance and
stat_fit_tidy should be used and the code for construction of
character strings from numeric values and their mapping to aesthetic
label needs to be explicitly supplied by the user.
Other ggplot statistics for linear and polynomial regression:
stat_poly_line()
Examples
# generate artificial data
set.seed(4321)
x <- 1:100
y <- (x + x^2 + x^3) + rnorm(length(x), mean = 0, sd = mean(x^3) / 4)
y <- y / max(y)
my.data <- data.frame(x = x, y = y,
group = c("A", "B"),
y2 = y * c(1, 2) + c(0, 0.1),
w = sqrt(x))
# give a name to a formula
formula <- y ~ poly(x, 3, raw = TRUE)
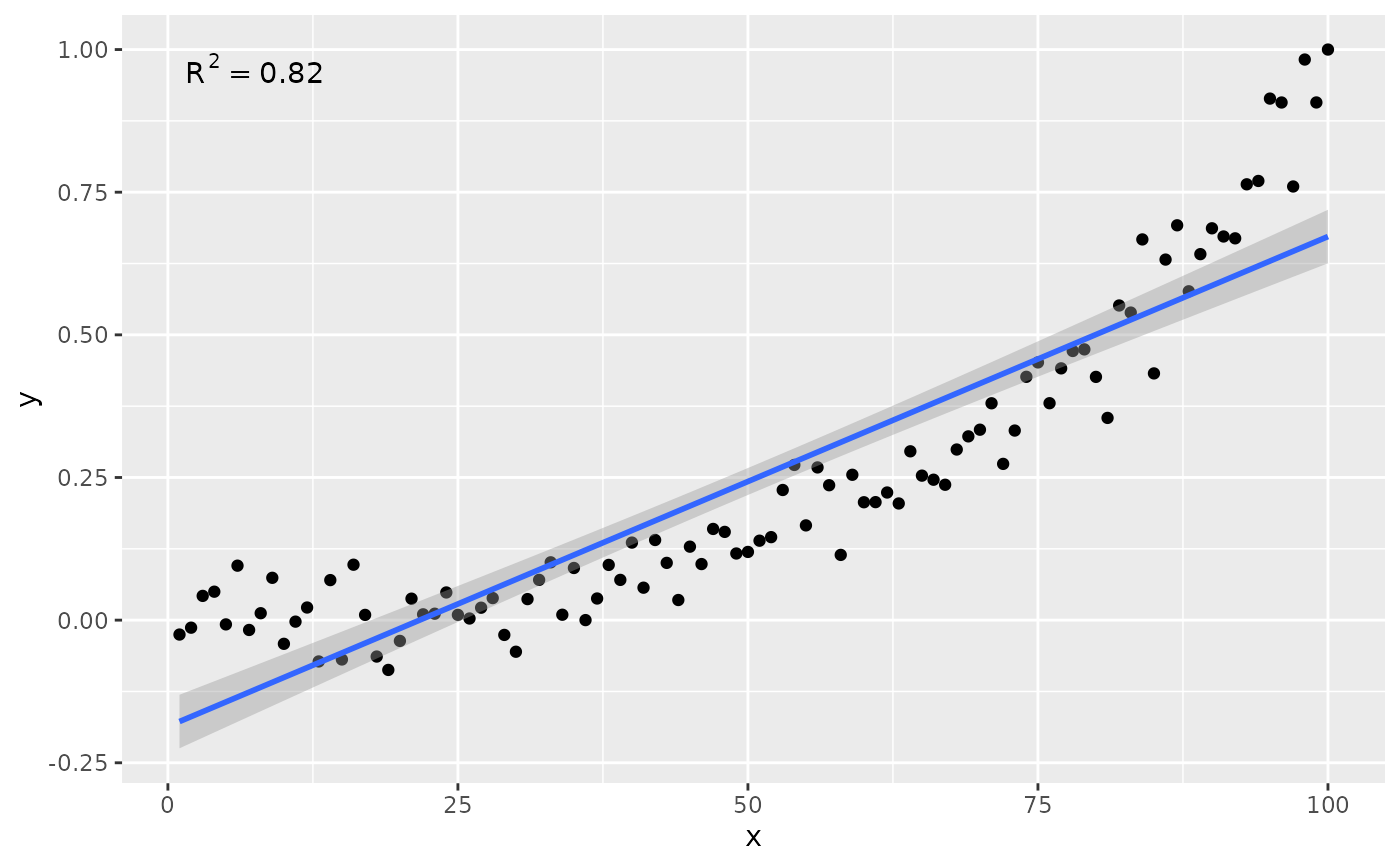
# using defaults
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line() +
stat_poly_eq()
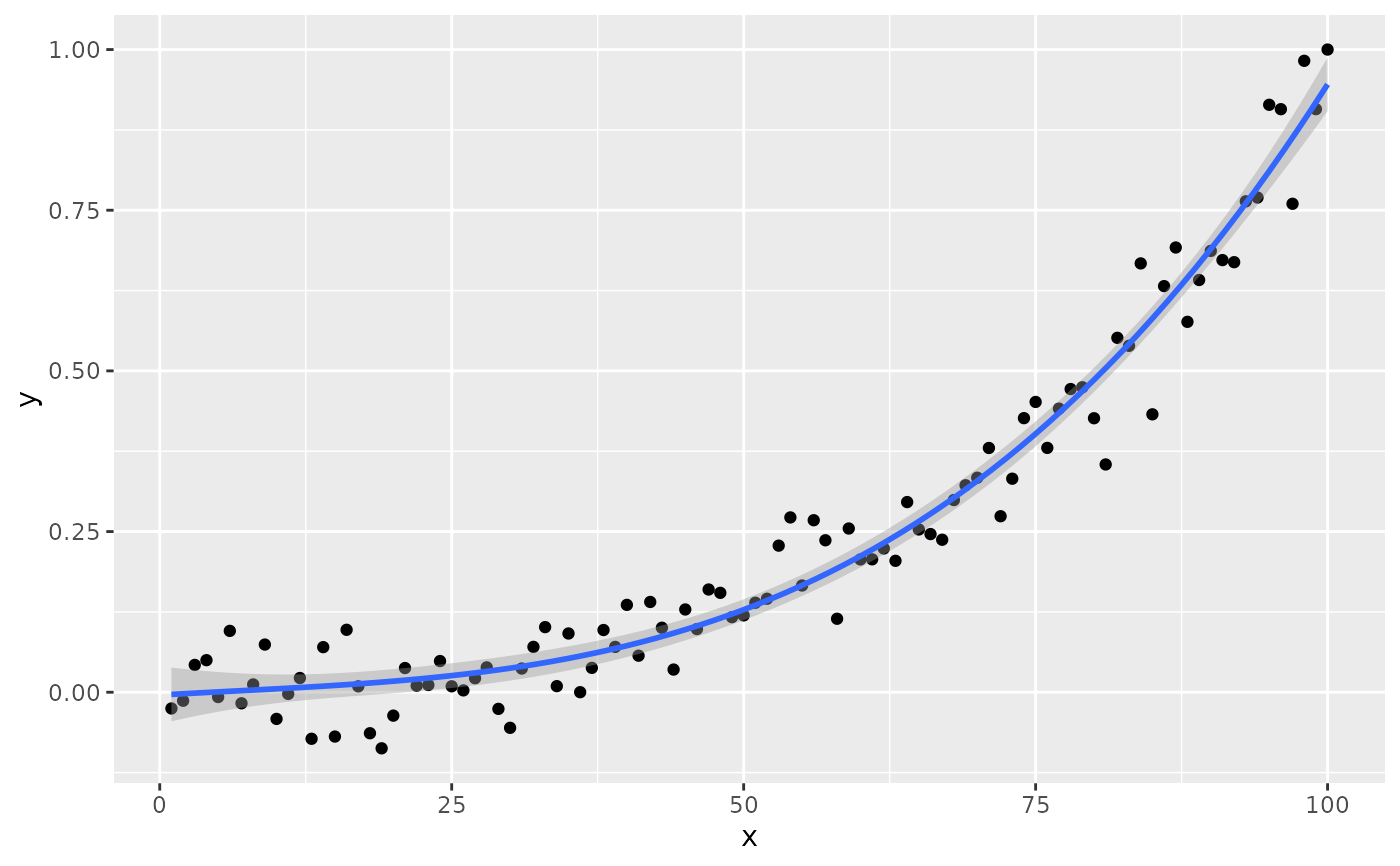
 # no weights
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
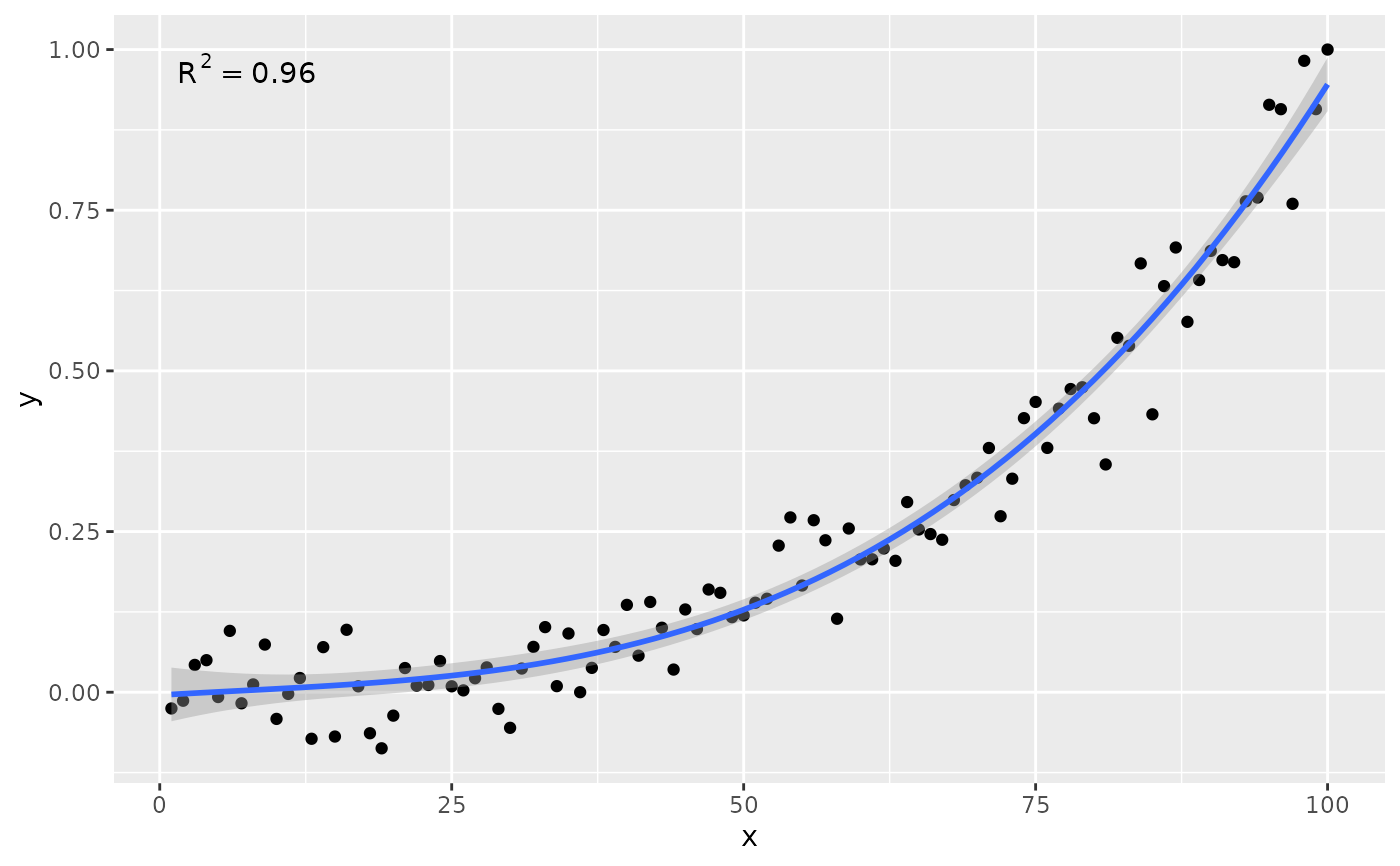
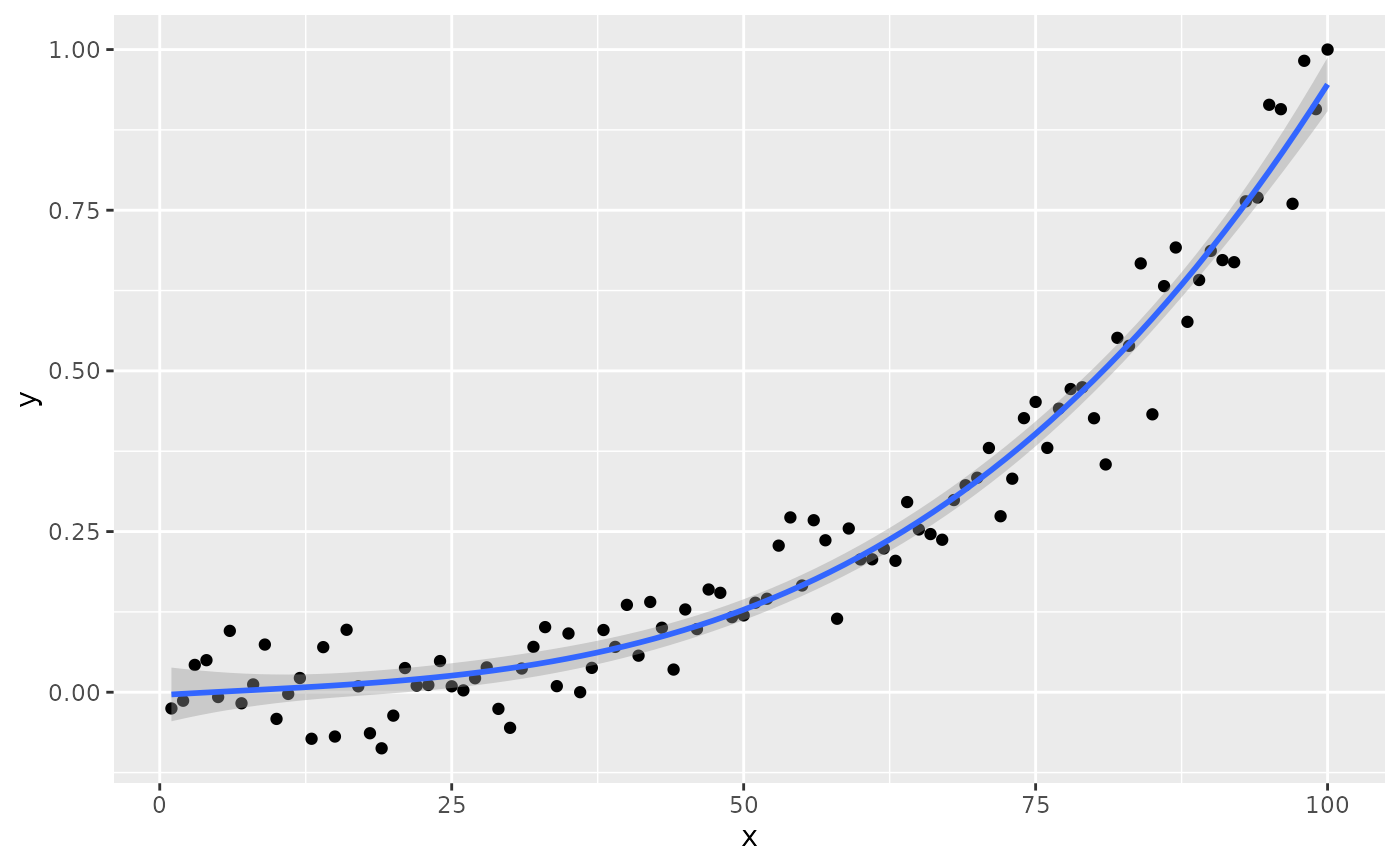
# no weights
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
 # other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula)
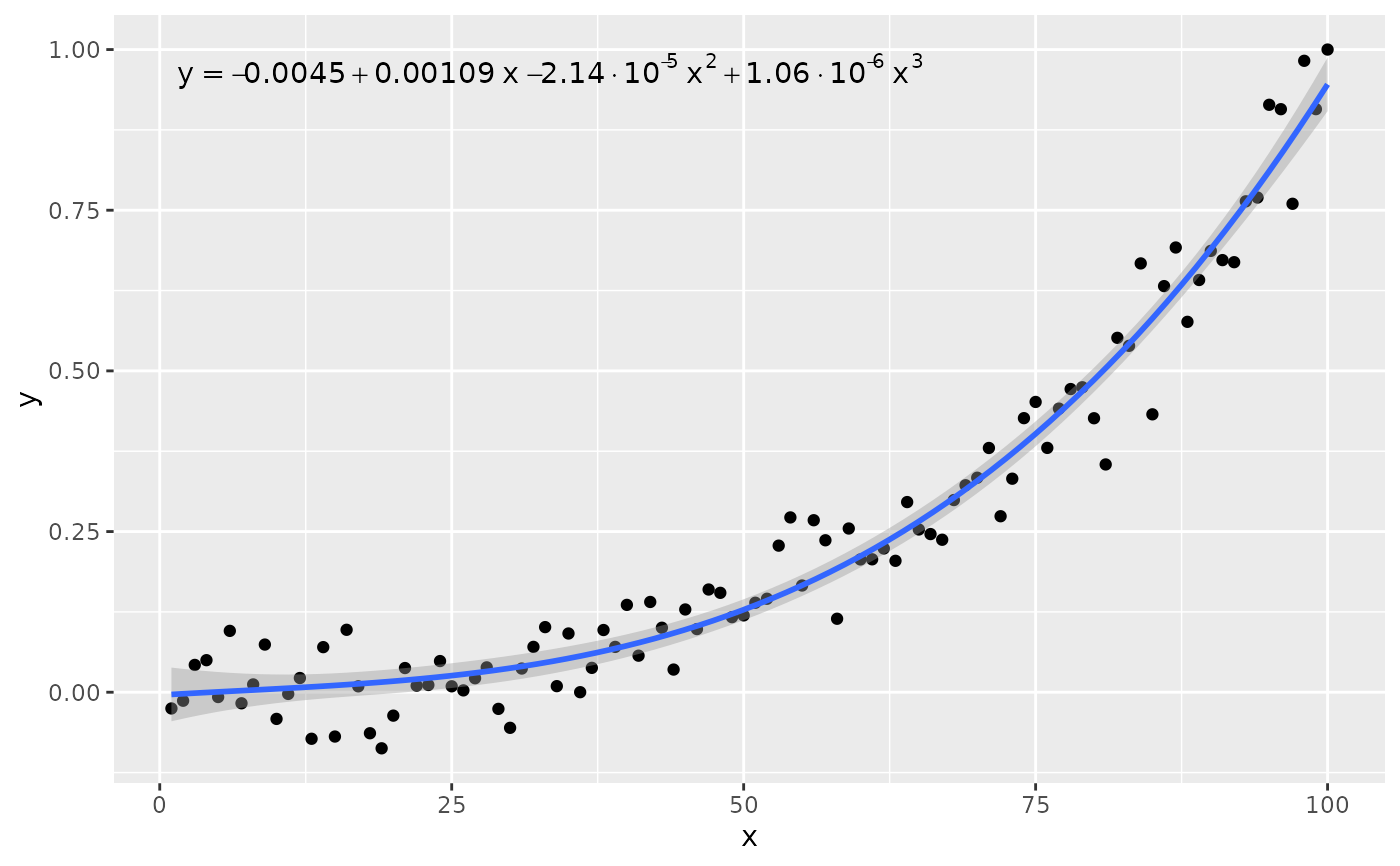
# other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula)
 # other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula, decreasing = TRUE)
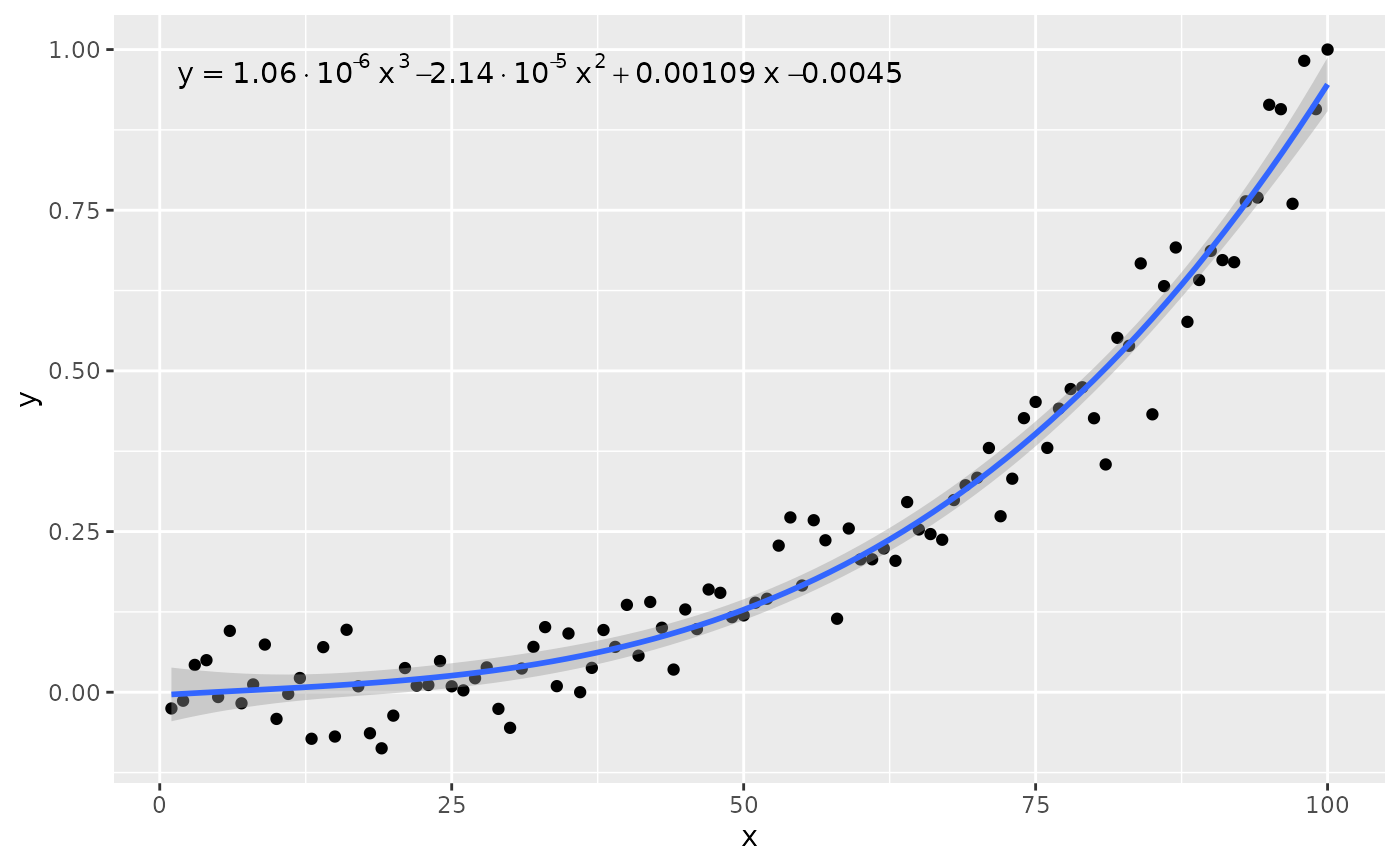
# other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula, decreasing = TRUE)
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq", "R2"), formula = formula)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq", "R2"), formula = formula)
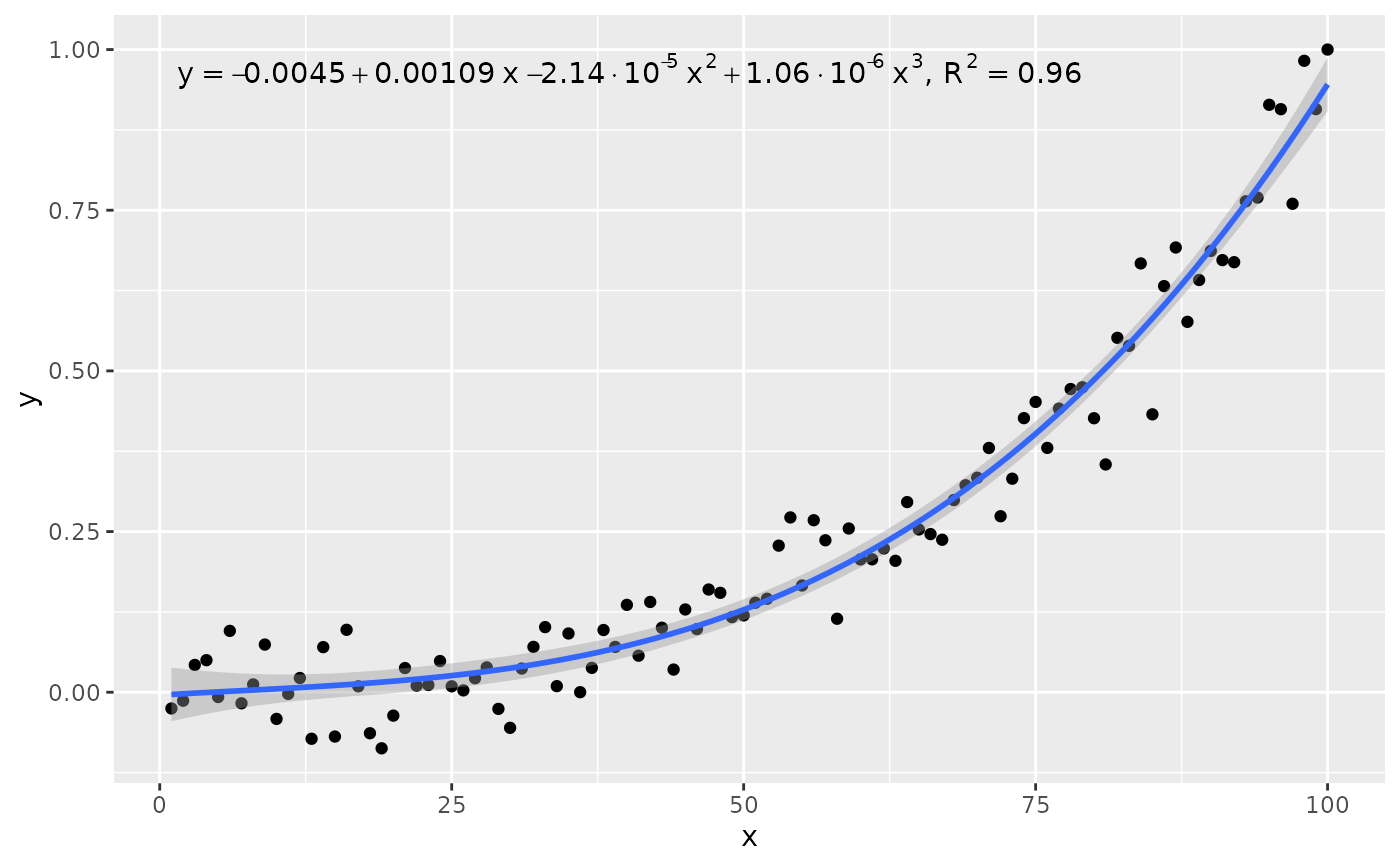 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "R2.CI", "P", "method"), formula = formula)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "R2.CI", "P", "method"), formula = formula)
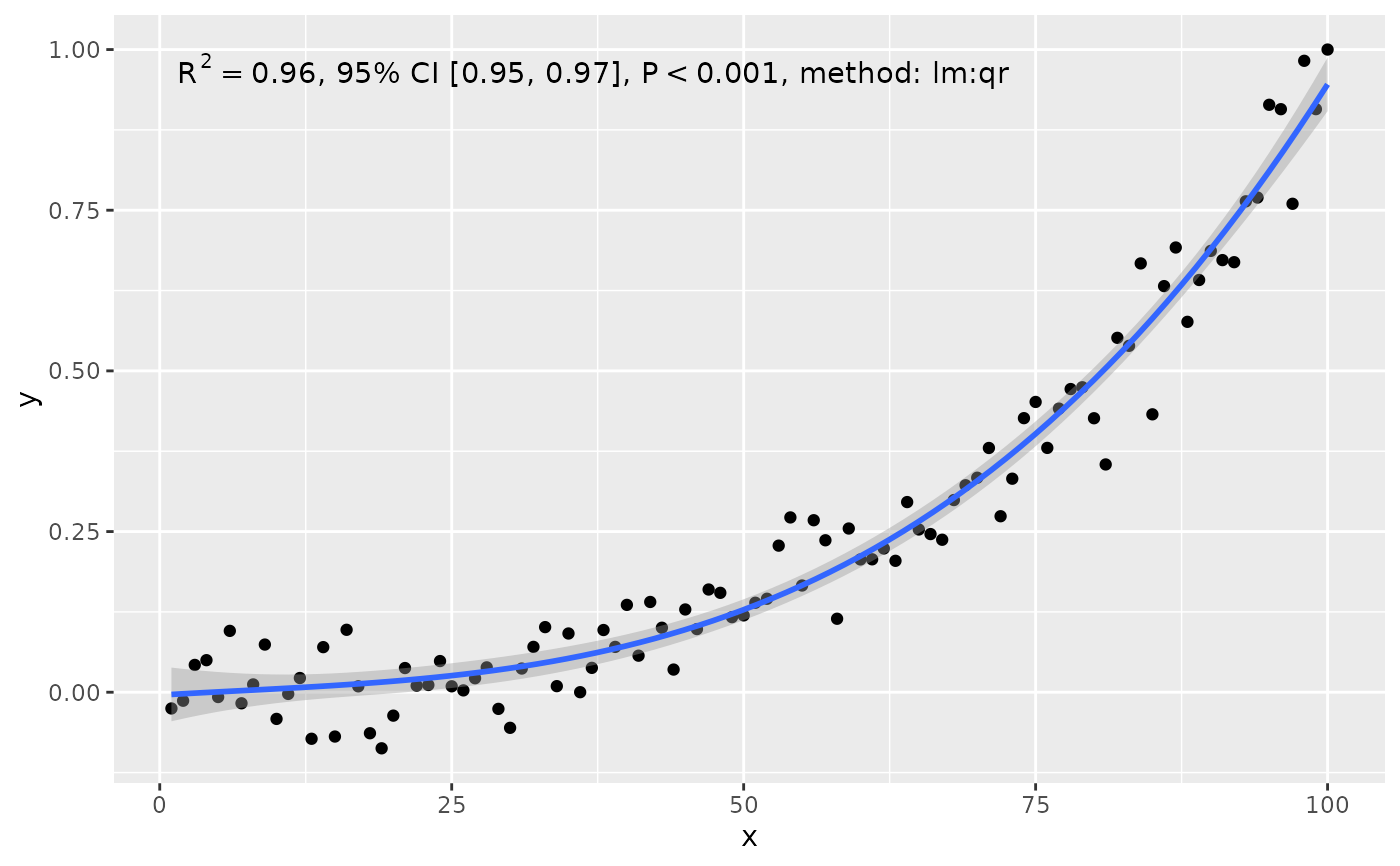 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "F", "P", "n", sep = "*\"; \"*"),
formula = formula)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "F", "P", "n", sep = "*\"; \"*"),
formula = formula)
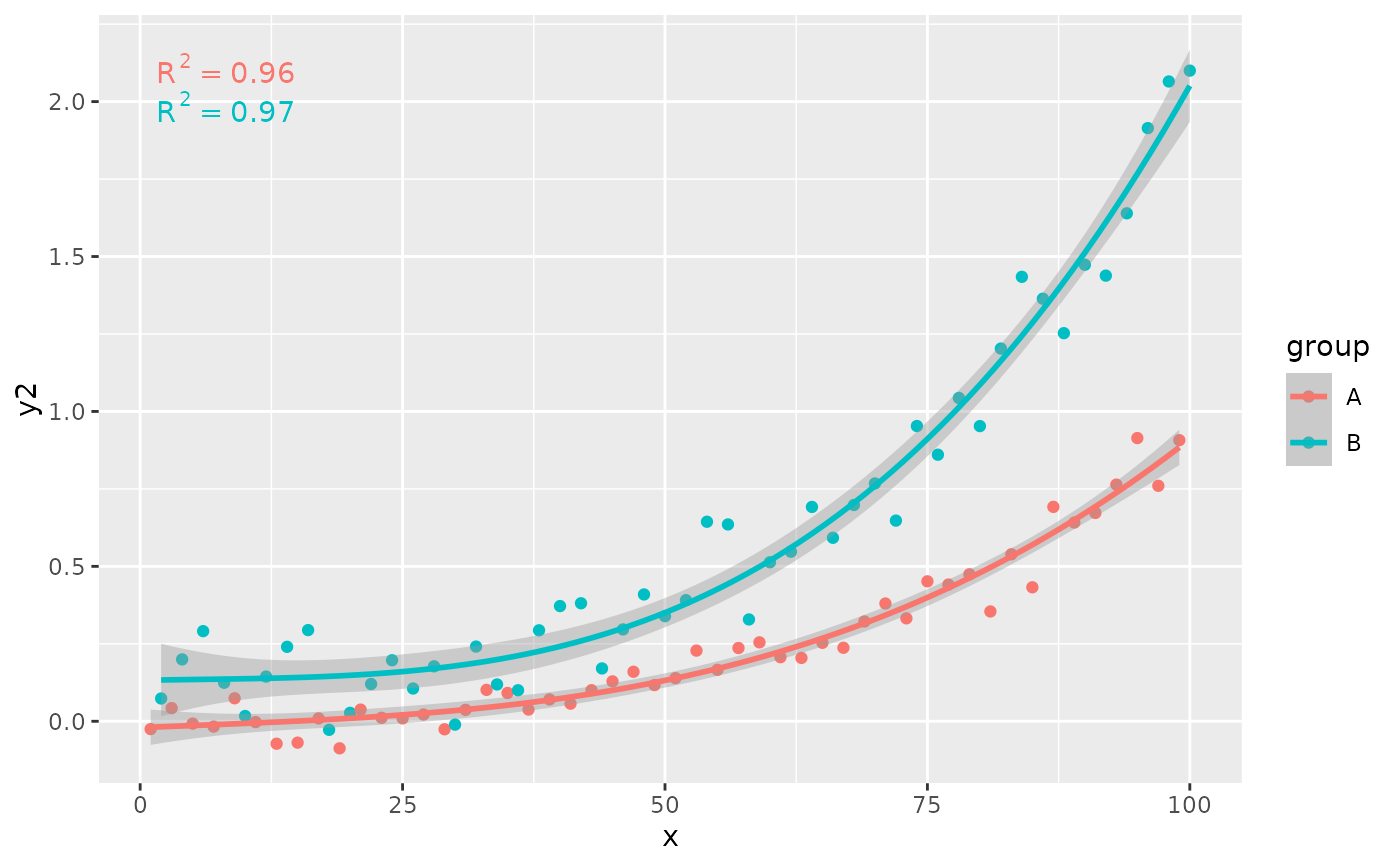
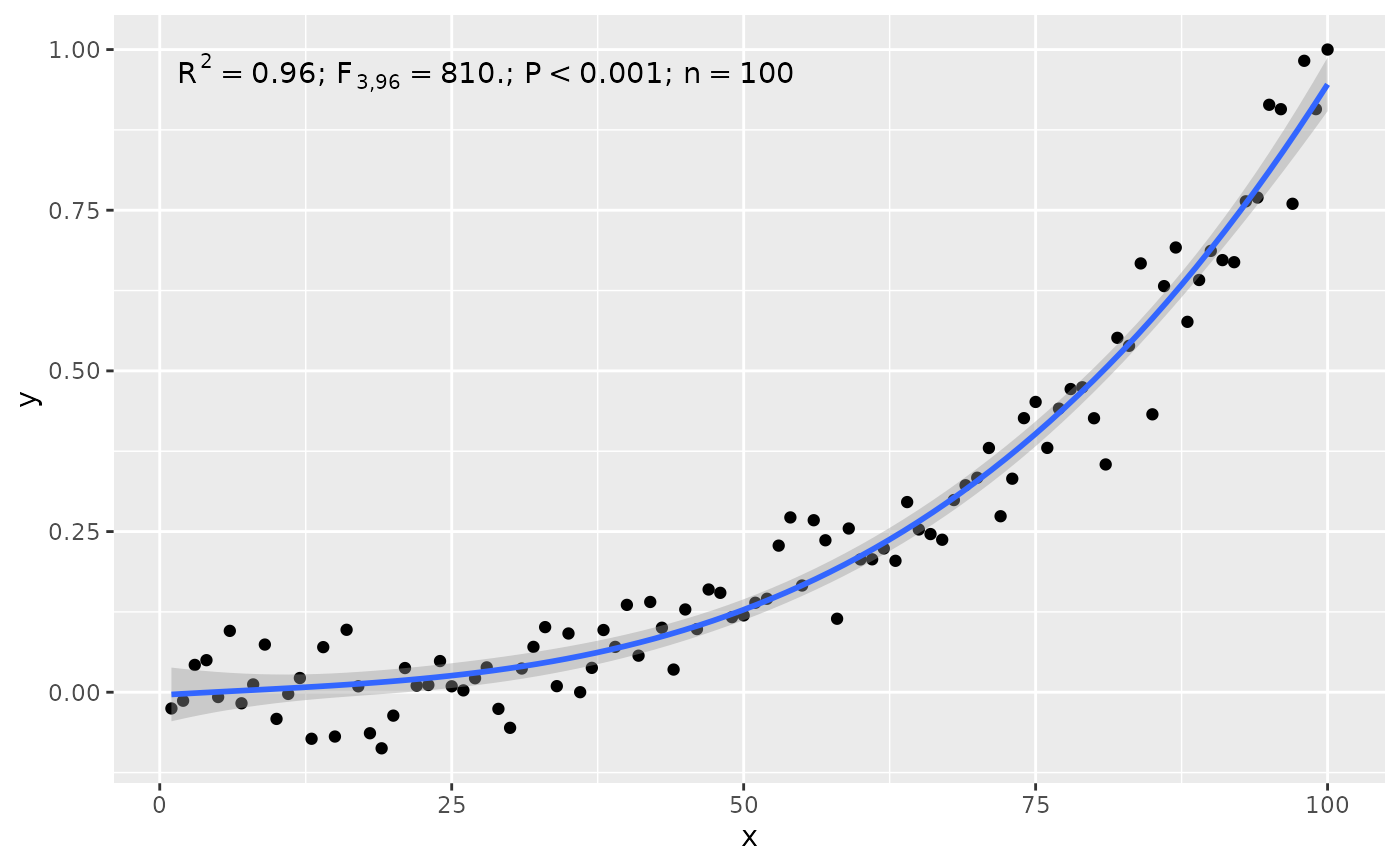 # grouping
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
# grouping
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
 # rotation
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, angle = 90)
# rotation
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, angle = 90)
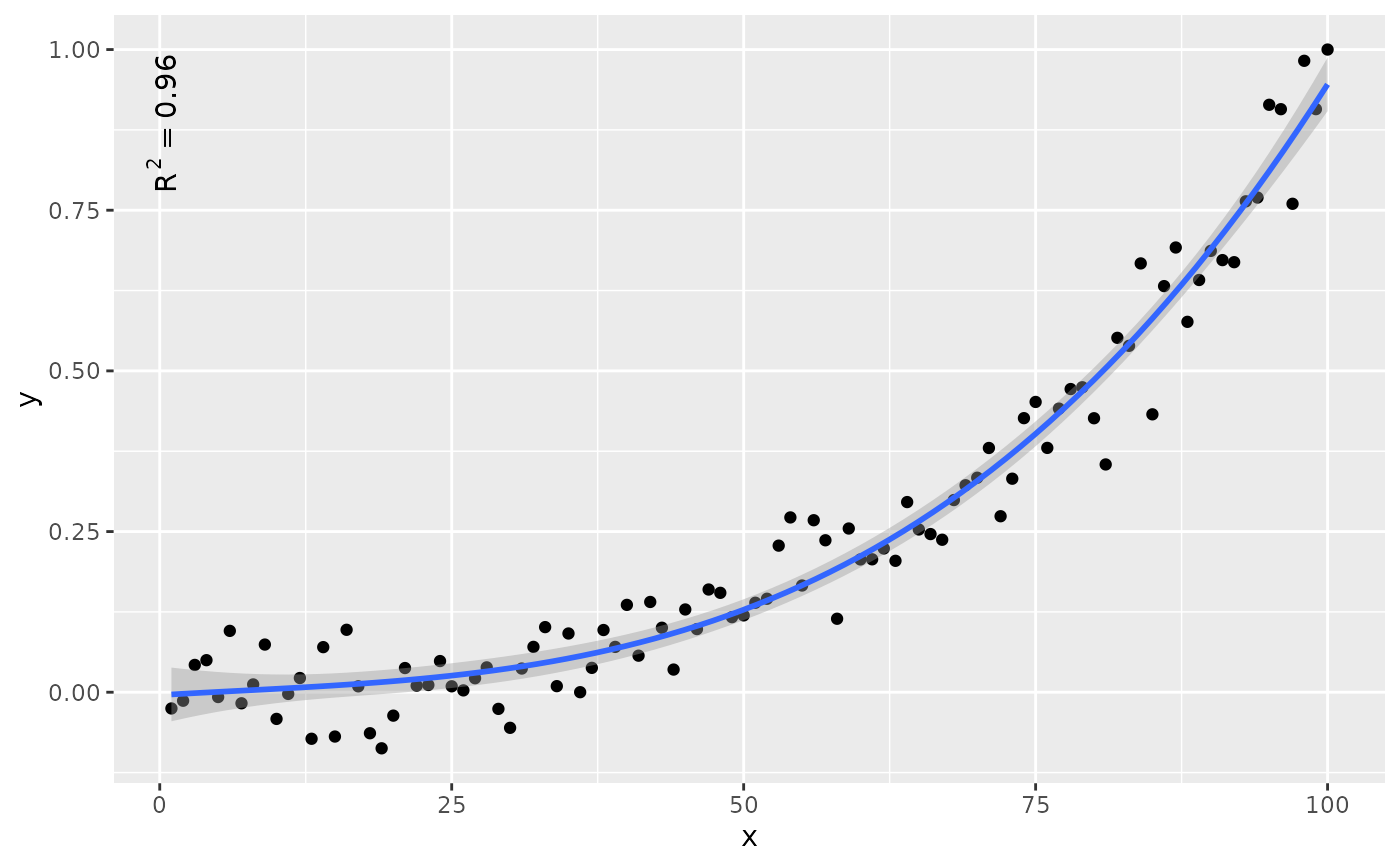 # label location
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = "bottom", label.x = "right")
# label location
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = "bottom", label.x = "right")
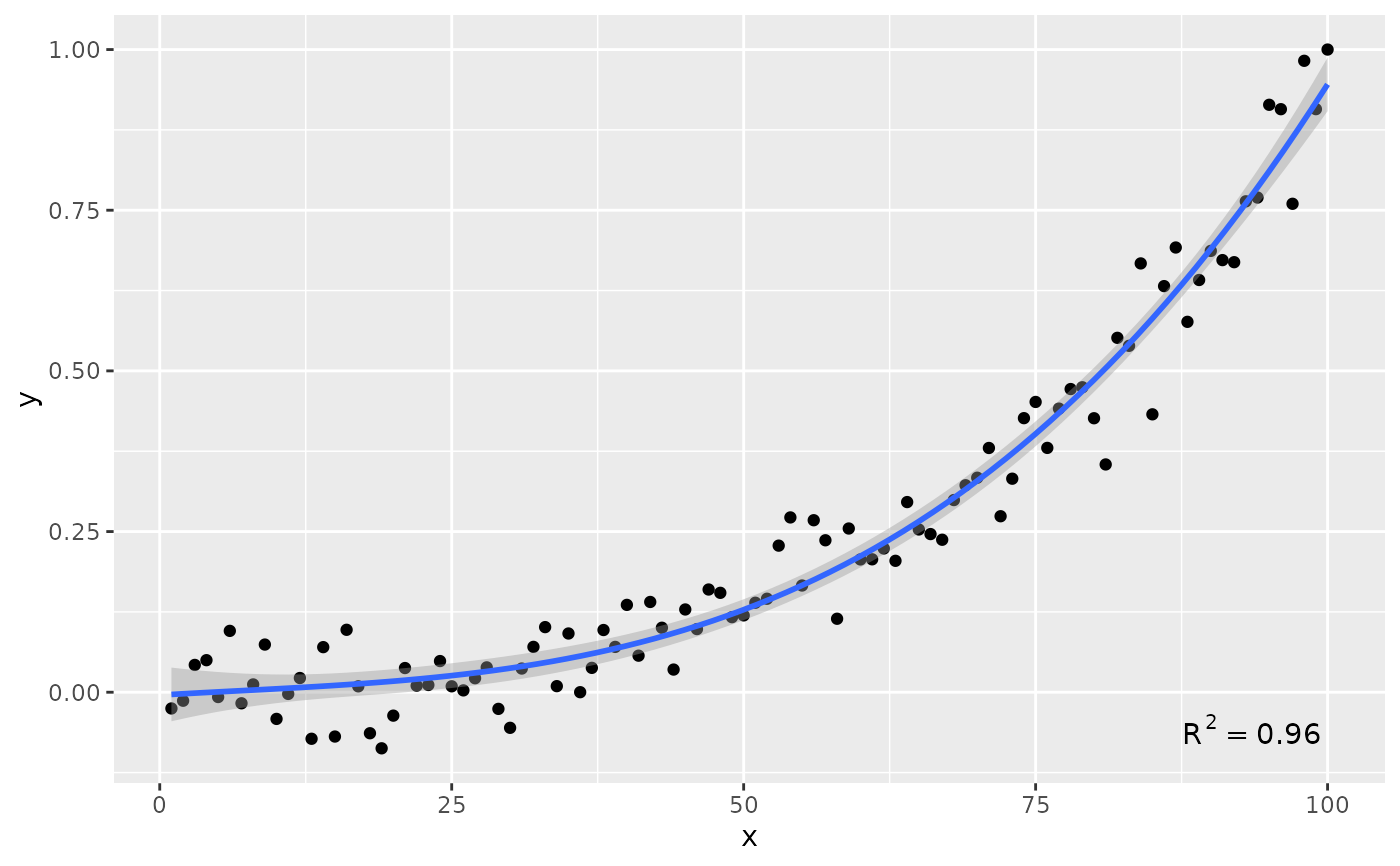 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = 0.1, label.x = 0.9)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = 0.1, label.x = 0.9)
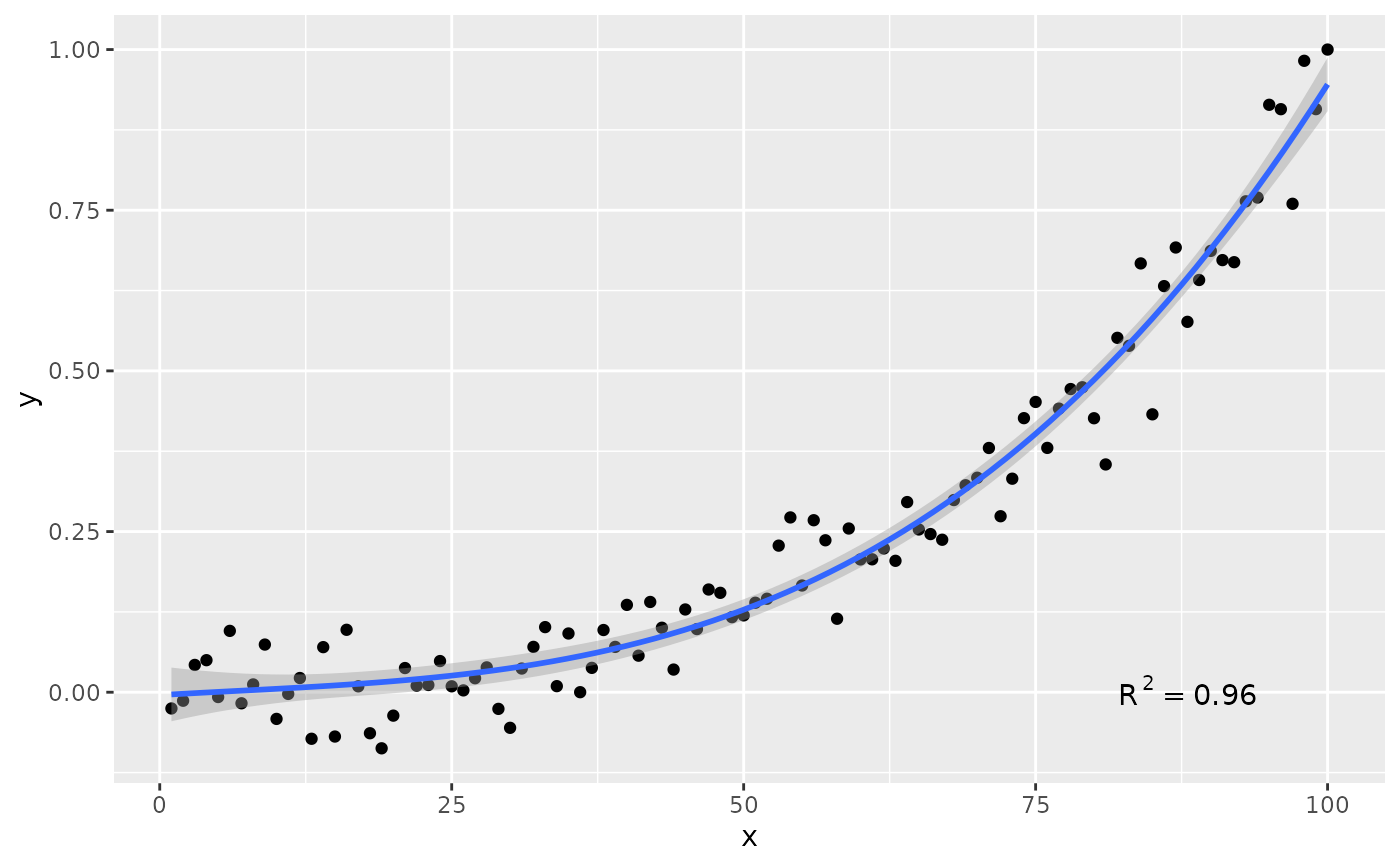 # modifying the explanatory variable within the model formula
# modifying the response variable within aes()
formula.trans <- y ~ I(x^2)
ggplot(my.data, aes(x, y + 1)) +
geom_point() +
stat_poly_line(formula = formula.trans) +
stat_poly_eq(use_label("eq"),
formula = formula.trans,
eq.x.rhs = "~x^2",
eq.with.lhs = "y + 1~~`=`~~")
#> Warning: 'formula' not an increasing polynomial: 'eq.label' is NA!
#> Warning: Removed 1 row containing missing values or values outside the scale range
#> (`geom_text_npc()`).
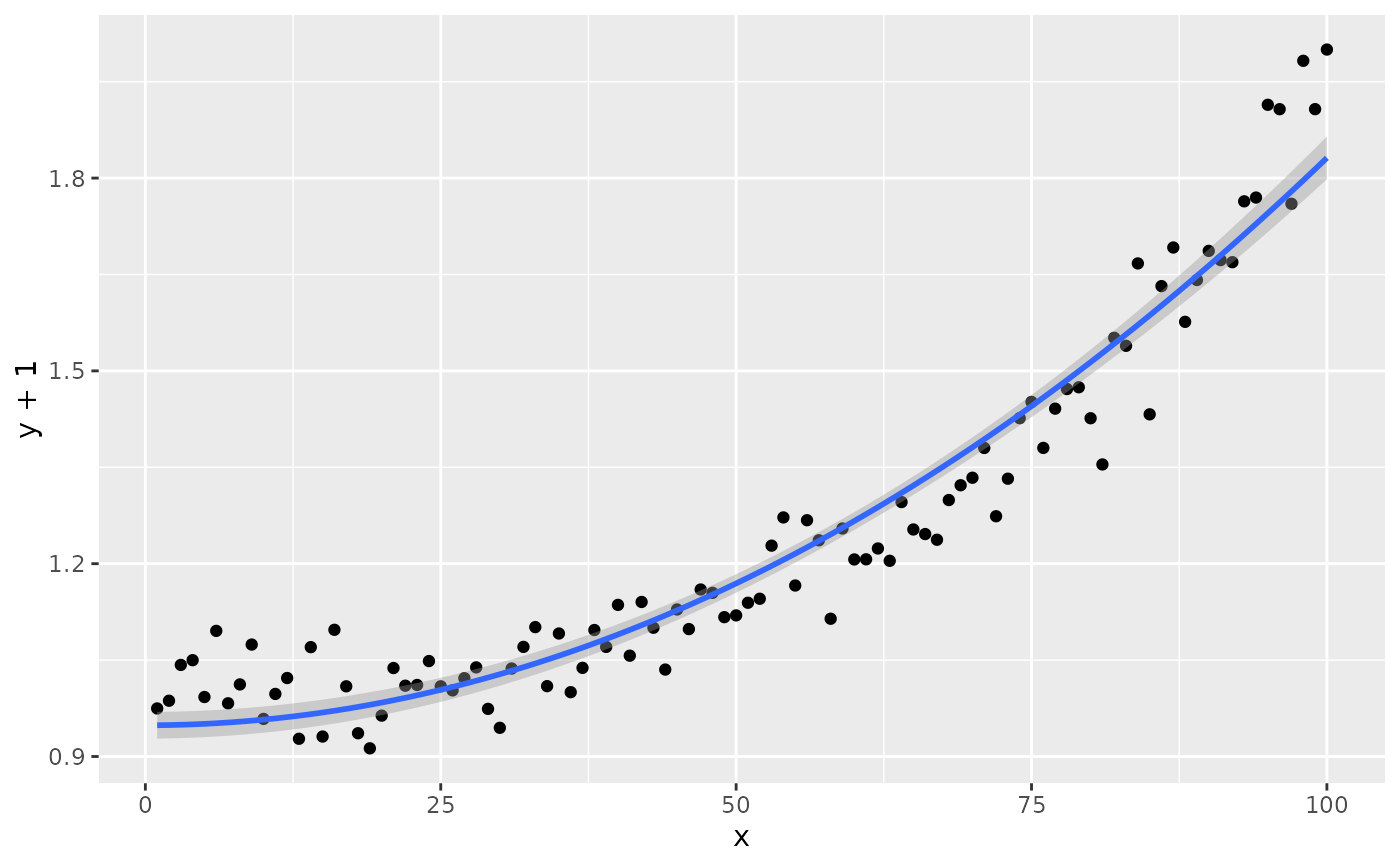
# modifying the explanatory variable within the model formula
# modifying the response variable within aes()
formula.trans <- y ~ I(x^2)
ggplot(my.data, aes(x, y + 1)) +
geom_point() +
stat_poly_line(formula = formula.trans) +
stat_poly_eq(use_label("eq"),
formula = formula.trans,
eq.x.rhs = "~x^2",
eq.with.lhs = "y + 1~~`=`~~")
#> Warning: 'formula' not an increasing polynomial: 'eq.label' is NA!
#> Warning: Removed 1 row containing missing values or values outside the scale range
#> (`geom_text_npc()`).
 # using weights
ggplot(my.data, aes(x, y, weight = w)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
# using weights
ggplot(my.data, aes(x, y, weight = w)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
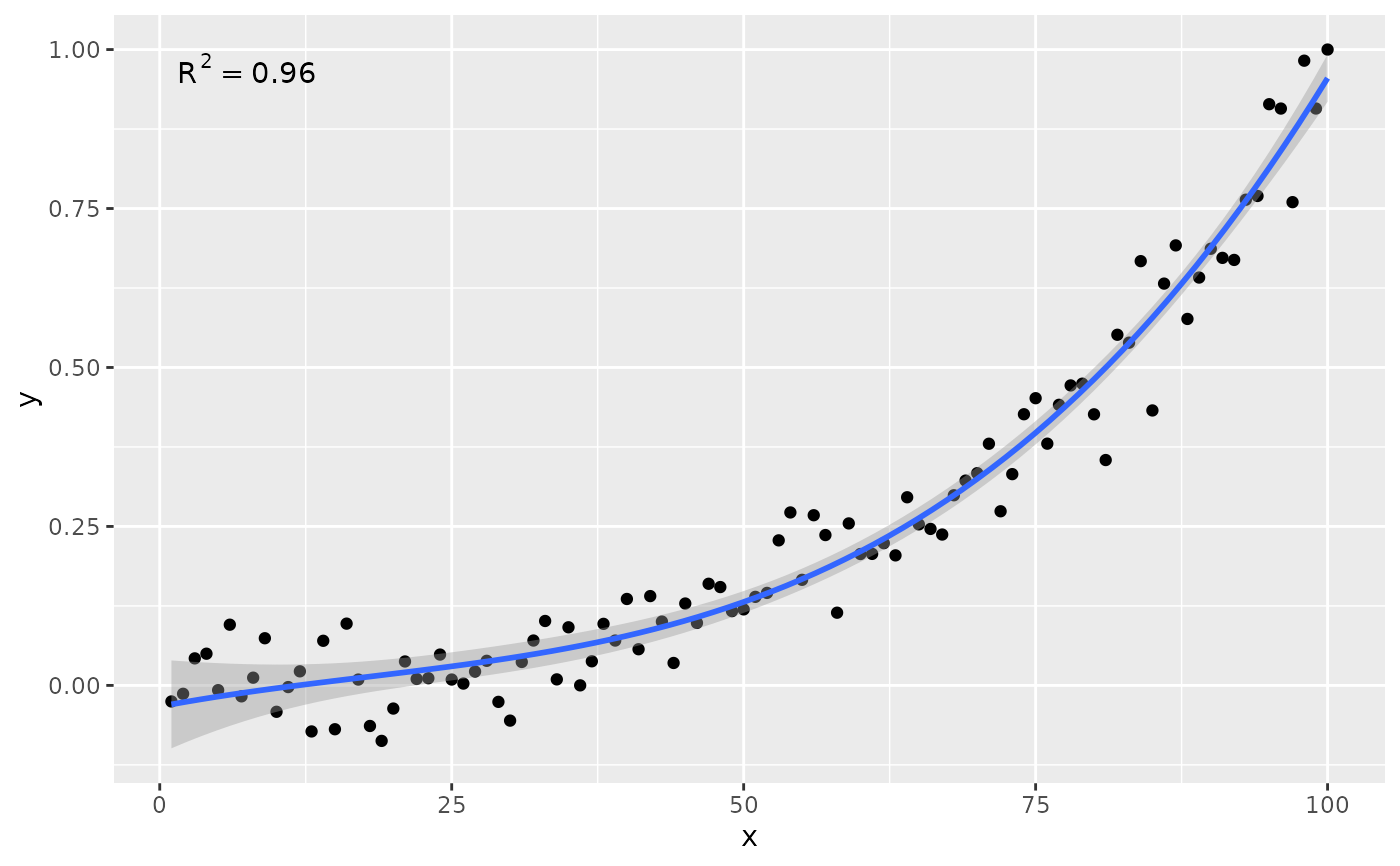 # no weights, 4 digits for R square
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, rr.digits = 4)
# no weights, 4 digits for R square
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, rr.digits = 4)
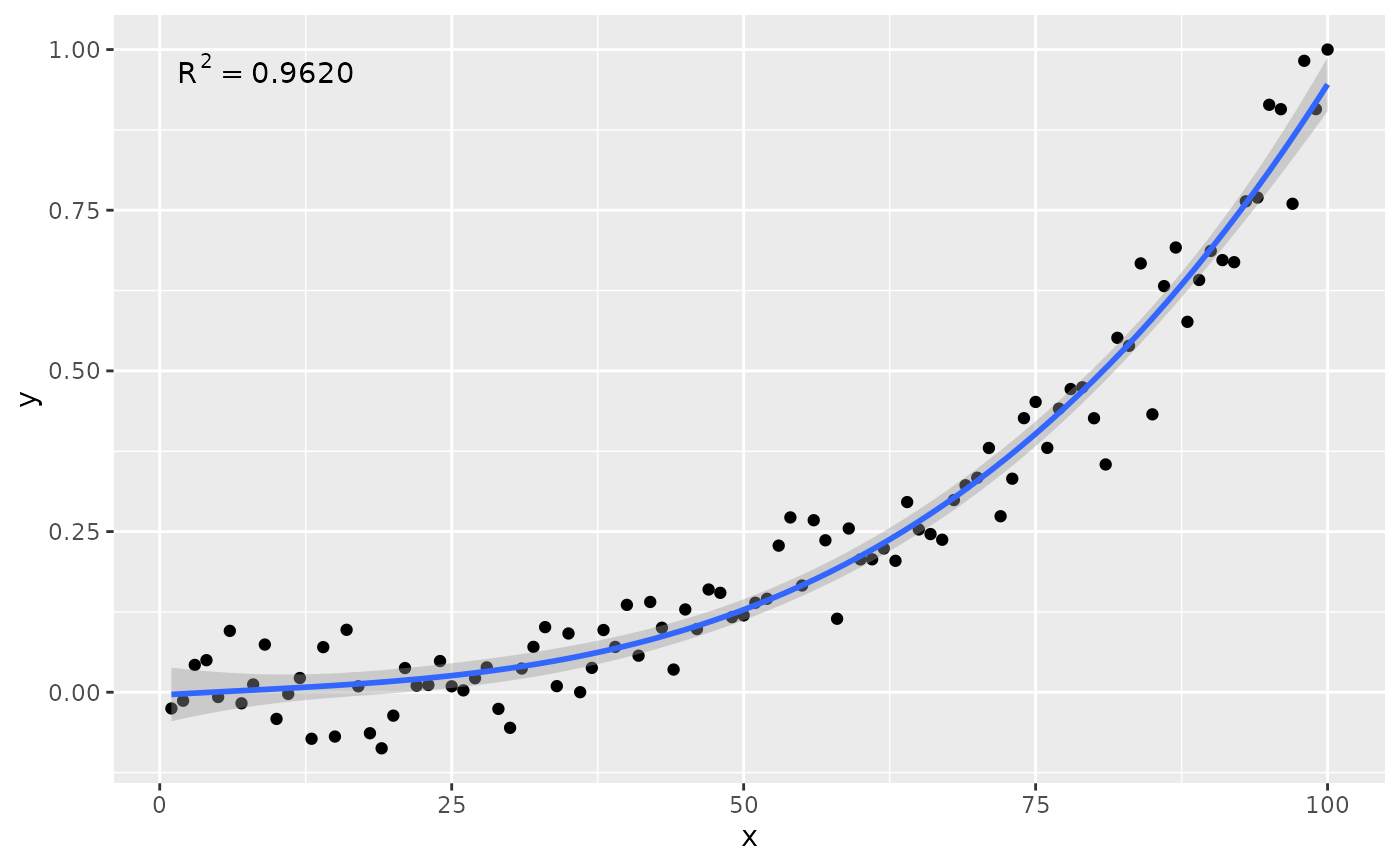 # manually assemble and map a specific label using paste() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = paste(after_stat(rr.label),
after_stat(n.label), sep = "*\", \"*")),
formula = formula)
# manually assemble and map a specific label using paste() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = paste(after_stat(rr.label),
after_stat(n.label), sep = "*\", \"*")),
formula = formula)
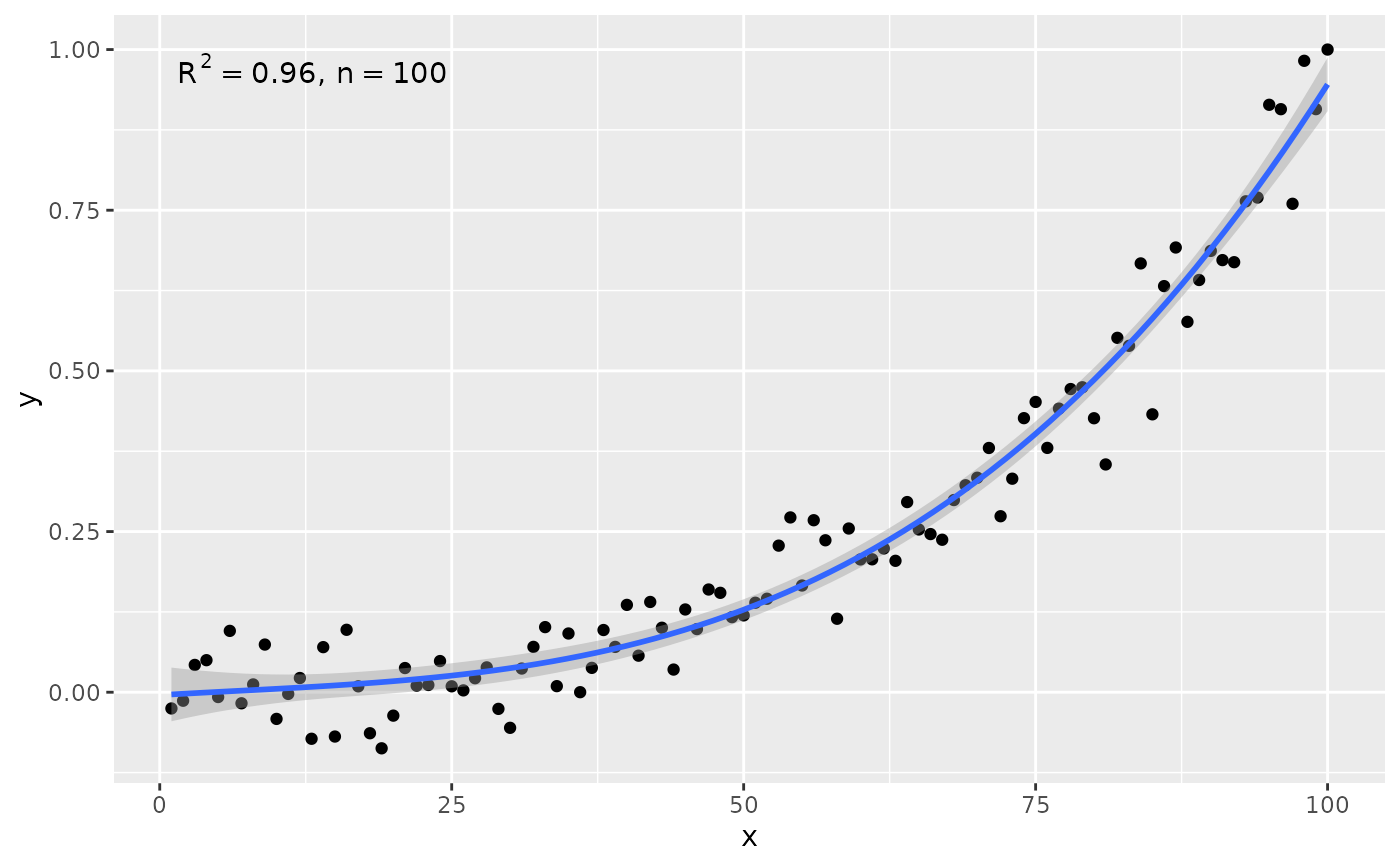 # manually assemble and map a specific label using sprintf() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = sprintf("%s*\" with \"*%s*\" and \"*%s",
after_stat(rr.label),
after_stat(f.value.label),
after_stat(p.value.label))),
formula = formula)
# manually assemble and map a specific label using sprintf() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = sprintf("%s*\" with \"*%s*\" and \"*%s",
after_stat(rr.label),
after_stat(f.value.label),
after_stat(p.value.label))),
formula = formula)
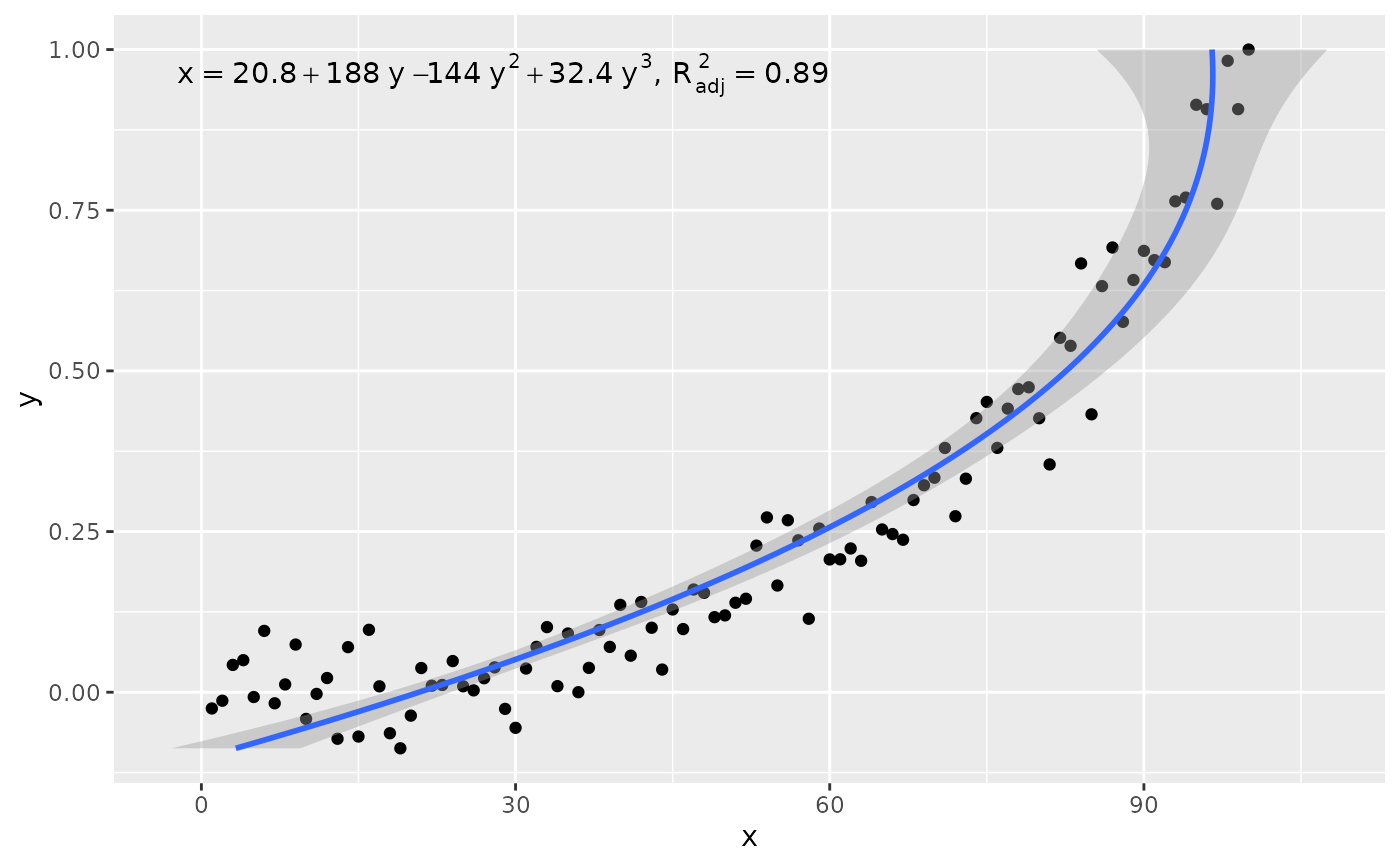
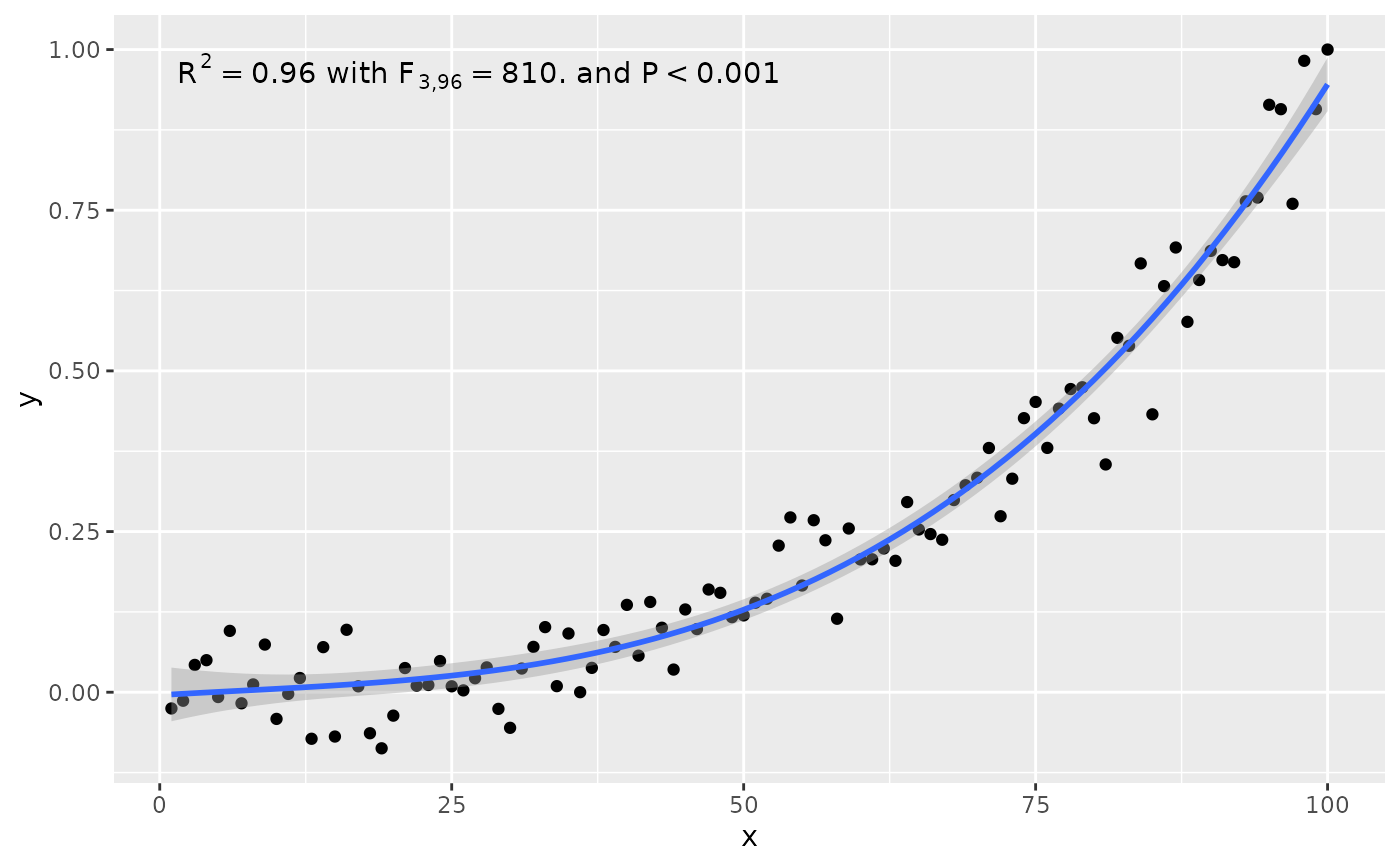 # x on y regression
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula, orientation = "y") +
stat_poly_eq(use_label("eq", "adj.R2"),
formula = x ~ poly(y, 3, raw = TRUE))
# x on y regression
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula, orientation = "y") +
stat_poly_eq(use_label("eq", "adj.R2"),
formula = x ~ poly(y, 3, raw = TRUE))
 # conditional user specified label
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = ifelse(after_stat(adj.r.squared) > 0.96,
paste(after_stat(adj.rr.label),
after_stat(eq.label),
sep = "*\", \"*"),
after_stat(adj.rr.label))),
rr.digits = 3,
formula = formula)
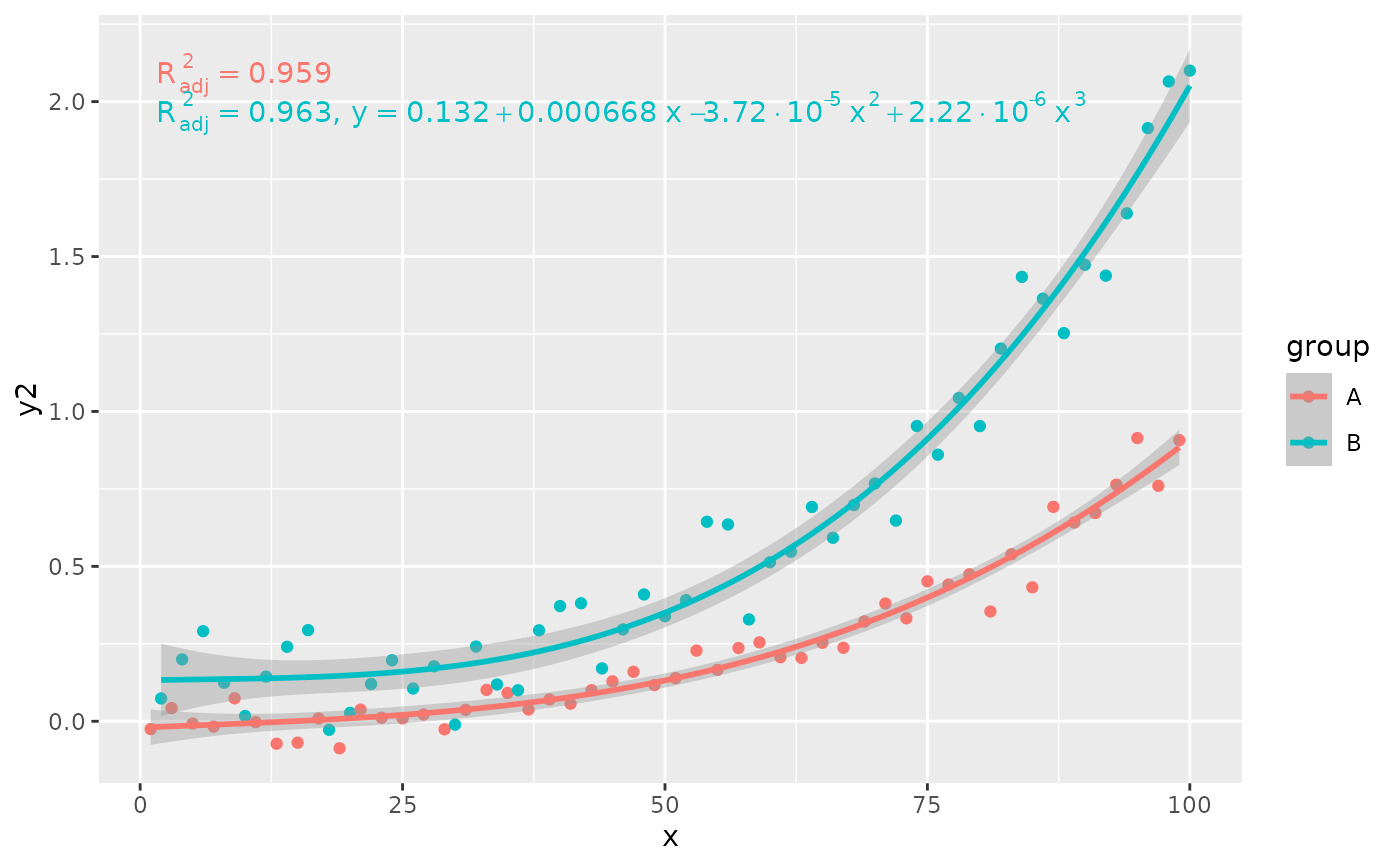
# conditional user specified label
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = ifelse(after_stat(adj.r.squared) > 0.96,
paste(after_stat(adj.rr.label),
after_stat(eq.label),
sep = "*\", \"*"),
after_stat(adj.rr.label))),
rr.digits = 3,
formula = formula)
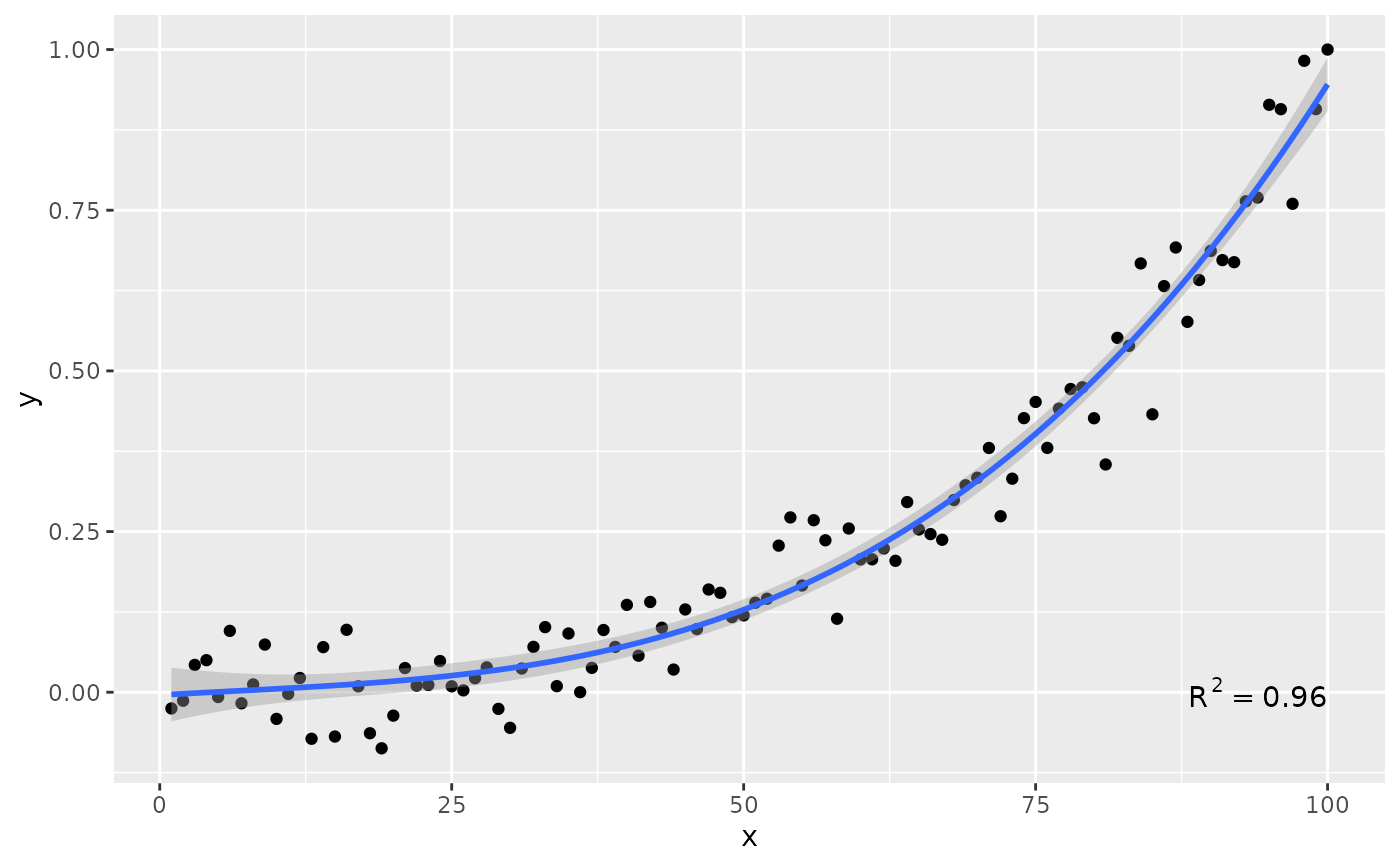
 # geom = "text"
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(geom = "text", label.x = 100, label.y = 0, hjust = 1,
formula = formula)
# geom = "text"
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(geom = "text", label.x = 100, label.y = 0, hjust = 1,
formula = formula)
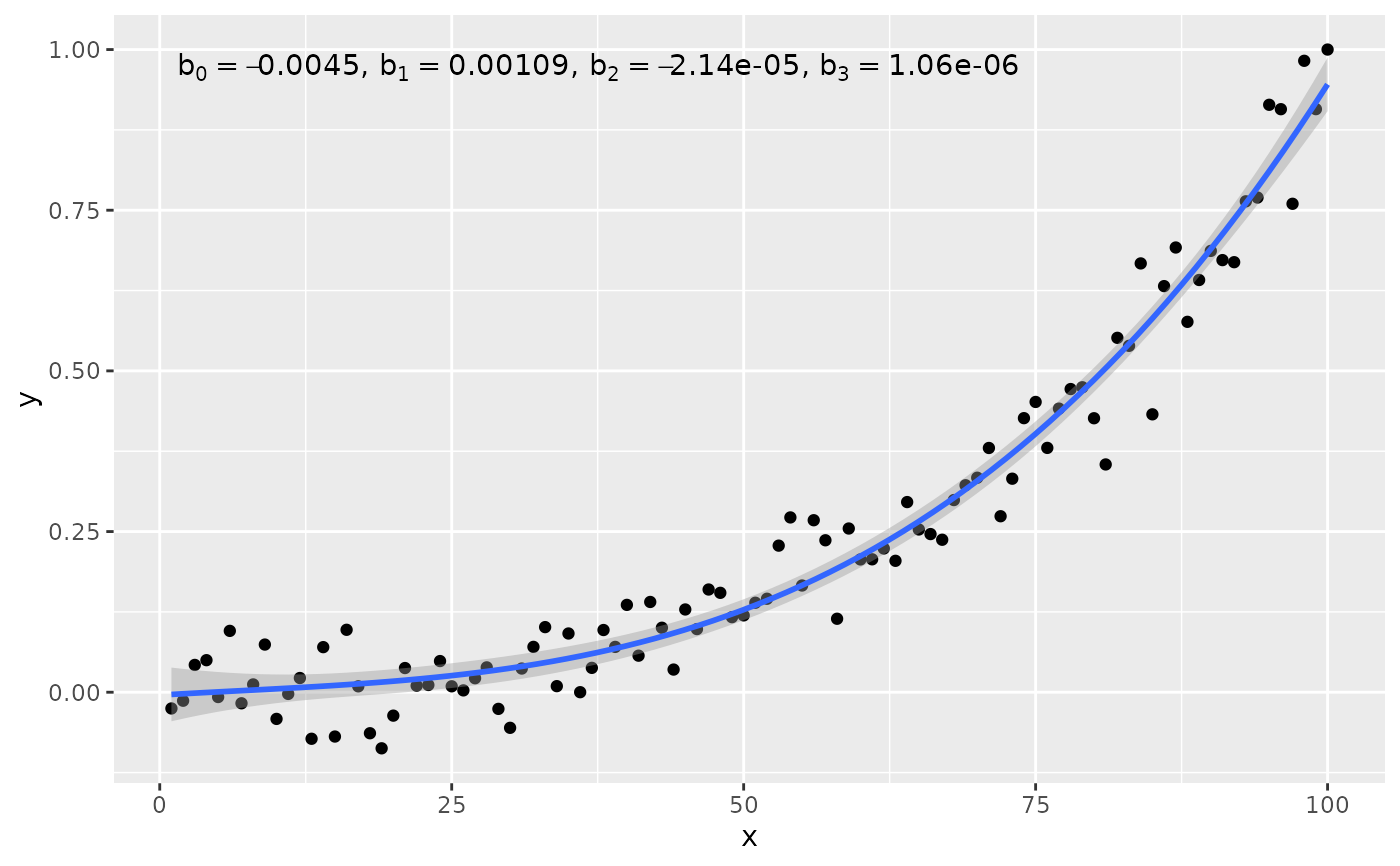
 # using numeric values
# Here we use columns b_0 ... b_3 for the coefficient estimates
my.format <-
"b[0]~`=`~%.3g*\", \"*b[1]~`=`~%.3g*\", \"*b[2]~`=`~%.3g*\", \"*b[3]~`=`~%.3g"
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
output.type = "numeric",
parse = TRUE,
mapping =
aes(label = sprintf(my.format,
after_stat(b_0), after_stat(b_1),
after_stat(b_2), after_stat(b_3))))
# using numeric values
# Here we use columns b_0 ... b_3 for the coefficient estimates
my.format <-
"b[0]~`=`~%.3g*\", \"*b[1]~`=`~%.3g*\", \"*b[2]~`=`~%.3g*\", \"*b[3]~`=`~%.3g"
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
output.type = "numeric",
parse = TRUE,
mapping =
aes(label = sprintf(my.format,
after_stat(b_0), after_stat(b_1),
after_stat(b_2), after_stat(b_3))))
 # Inspecting the returned data using geom_debug()
# This provides a quick way of finding out the names of the variables that
# are available for mapping to aesthetics with after_stat().
gginnards.installed <- requireNamespace("gginnards", quietly = TRUE)
if (gginnards.installed)
library(gginnards)
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug")
# Inspecting the returned data using geom_debug()
# This provides a quick way of finding out the names of the variables that
# are available for mapping to aesthetics with after_stat().
gginnards.installed <- requireNamespace("gginnards", quietly = TRUE)
if (gginnards.installed)
library(gginnards)
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA italic(R)^2~`=`~"0.96"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
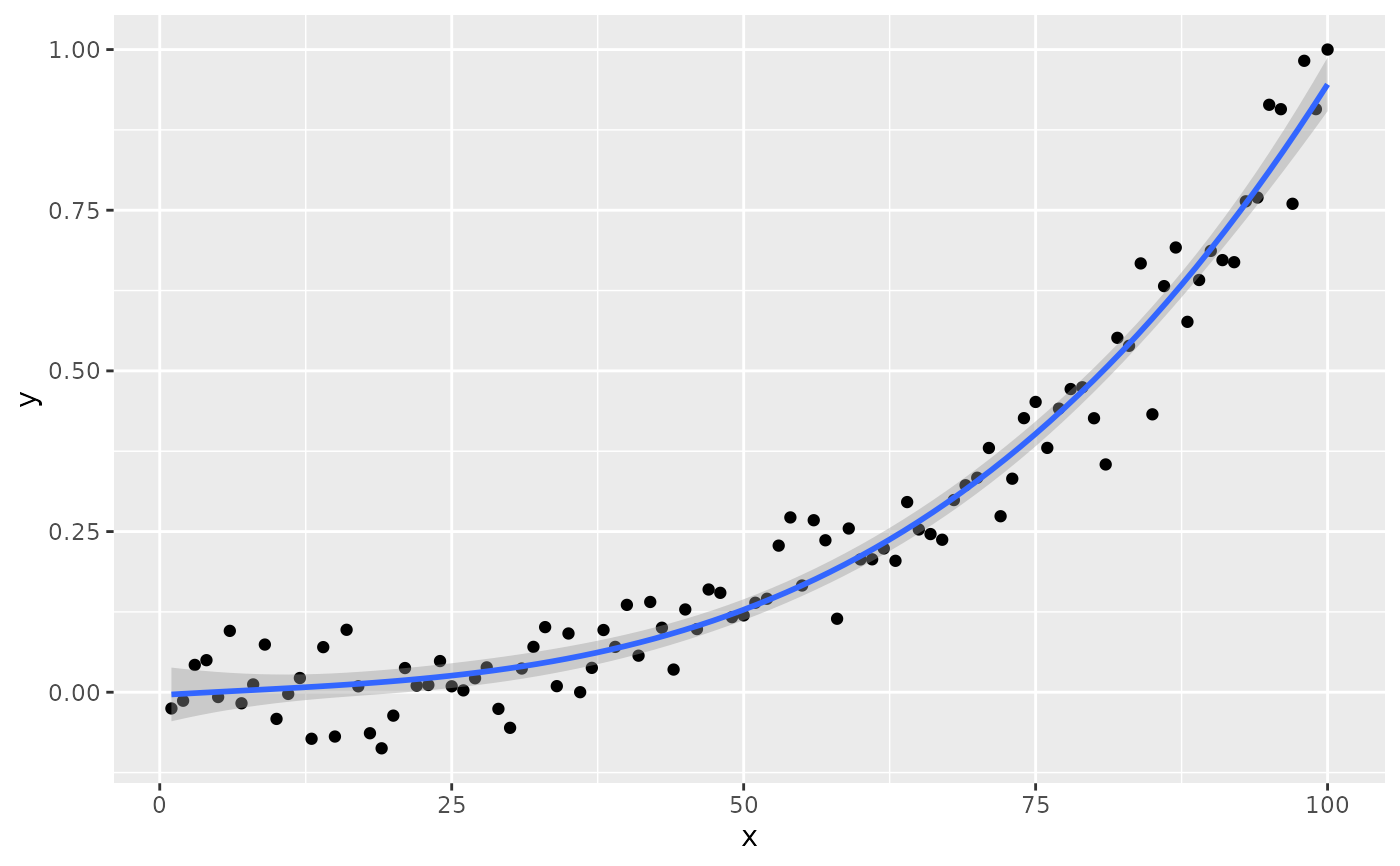
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug", output.type = "numeric")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA italic(R)^2~`=`~"0.96"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
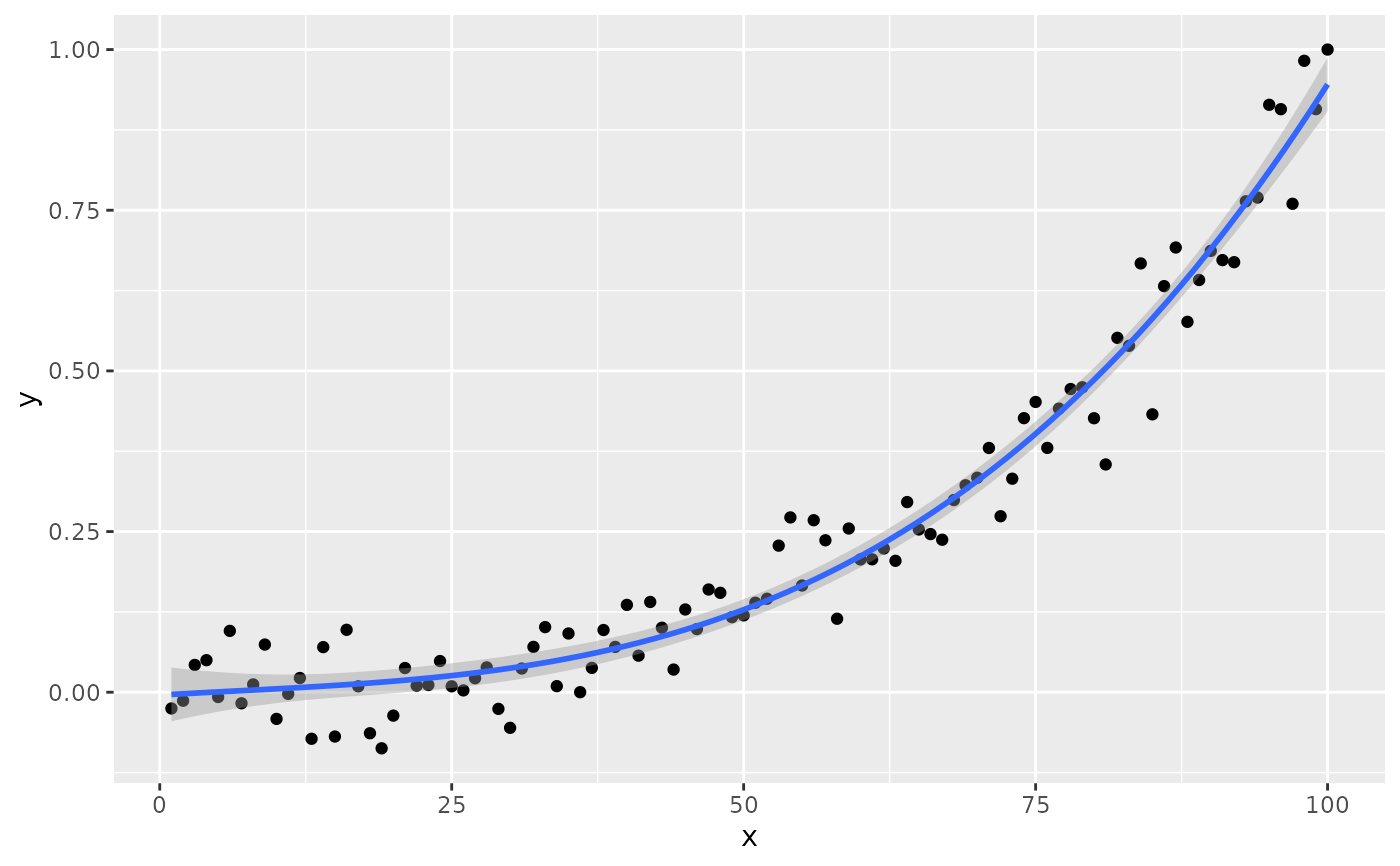
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug", output.type = "numeric")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA
#> coef.ls
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06, 2.268244e-02, 1.935252e-03, 4.440449e-05, 2.890821e-07, -1.981987e-01, 5.630500e-01, -4.819189e-01, 3.650820e+00, 8.433087e-01, 5.747135e-01, 6.309604e-01, 4.255076e-04
#> coefs r.squared
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06 0.9620171
#> rr.confint.level rr.confint.low rr.confint.high adj.r.squared f.value f.df1
#> 1 0.95 0.9464423 0.9696371 0.9608301 810.484 3
#> f.df2 p.value AIC BIC n rr.label b_0.constant b_0
#> 1 96 5.110349e-68 -291.8639 -278.838 100 FALSE -0.004495631
#> b_1 b_2 b_3 fm.method fm.class
#> 1 0.001089644 -2.139937e-05 1.055387e-06 lm:qr lm
#> fm.formula fm.formula.chr x y PANEL group
#> 1 y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1 -1
#> orientation
#> 1 x
# names of the variables
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
summary.fun = colnames)
#> Warning: Ignoring unknown parameters: `summary.fun`
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA
#> coef.ls
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06, 2.268244e-02, 1.935252e-03, 4.440449e-05, 2.890821e-07, -1.981987e-01, 5.630500e-01, -4.819189e-01, 3.650820e+00, 8.433087e-01, 5.747135e-01, 6.309604e-01, 4.255076e-04
#> coefs r.squared
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06 0.9620171
#> rr.confint.level rr.confint.low rr.confint.high adj.r.squared f.value f.df1
#> 1 0.95 0.9464423 0.9696371 0.9608301 810.484 3
#> f.df2 p.value AIC BIC n rr.label b_0.constant b_0
#> 1 96 5.110349e-68 -291.8639 -278.838 100 FALSE -0.004495631
#> b_1 b_2 b_3 fm.method fm.class
#> 1 0.001089644 -2.139937e-05 1.055387e-06 lm:qr lm
#> fm.formula fm.formula.chr x y PANEL group
#> 1 y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1 -1
#> orientation
#> 1 x
# names of the variables
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
summary.fun = colnames)
#> Warning: Ignoring unknown parameters: `summary.fun`
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA italic(R)^2~`=`~"0.96"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
output.type = "expression",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA italic(R)^2~`=`~"0.96"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
output.type = "expression",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA italic(R)^2~`=`~"0.96"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = after_stat(eq.label)),
formula = formula, geom = "debug",
output.type = "markdown",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA italic(R)^2~`=`~"0.96"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = after_stat(eq.label)),
formula = formula, geom = "debug",
output.type = "markdown",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy
#> 1 NA NA
#> label
#> 1 _y_ = -0.00450+0.00109 _x_-2.14×10<sup>-5</sup> _x_<sup>2</sup>+1.06×10<sup>-6</sup> _x_<sup>3</sup>
#> eq.label
#> 1 _y_ = -0.00450+0.00109 _x_-2.14×10<sup>-5</sup> _x_<sup>2</sup>+1.06×10<sup>-6</sup> _x_<sup>3</sup>
#> rr.label adj.rr.label
#> 1 _R_<sup>2</sup> = 0.96 _R_<sup>2</sup><sub>adj</sub> = 0.96
#> rr.confint.label AIC.label BIC.label f.value.label
#> 1 95% CI [0.95, 0.97] AIC = -291.9 BIC = -278.8 _F_<sub>3,96</sub> = 810.
#> p.value.label n.label grp.label method.label r.squared adj.r.squared
#> 1 _P_ < 0.001 _n_ = 100 -1 "method: lm:qr" 0.9620171 0.9608301
#> p.value n fm.method fm.class fm.formula
#> 1 5.110349e-68 100 lm:qr lm y ~ poly(x, 3, raw = TRUE)
#> fm.formula.chr x y PANEL group orientation
#> 1 y ~ poly(x, 3, raw = TRUE) 1 1 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
output.type = "latex",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy
#> 1 NA NA
#> label
#> 1 _y_ = -0.00450+0.00109 _x_-2.14×10<sup>-5</sup> _x_<sup>2</sup>+1.06×10<sup>-6</sup> _x_<sup>3</sup>
#> eq.label
#> 1 _y_ = -0.00450+0.00109 _x_-2.14×10<sup>-5</sup> _x_<sup>2</sup>+1.06×10<sup>-6</sup> _x_<sup>3</sup>
#> rr.label adj.rr.label
#> 1 _R_<sup>2</sup> = 0.96 _R_<sup>2</sup><sub>adj</sub> = 0.96
#> rr.confint.label AIC.label BIC.label f.value.label
#> 1 95% CI [0.95, 0.97] AIC = -291.9 BIC = -278.8 _F_<sub>3,96</sub> = 810.
#> p.value.label n.label grp.label method.label r.squared adj.r.squared
#> 1 _P_ < 0.001 _n_ = 100 -1 "method: lm:qr" 0.9620171 0.9608301
#> p.value n fm.method fm.class fm.formula
#> 1 5.110349e-68 100 lm:qr lm y ~ poly(x, 3, raw = TRUE)
#> fm.formula.chr x y PANEL group orientation
#> 1 y ~ poly(x, 3, raw = TRUE) 1 1 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
output.type = "latex",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA R^2 = 0.96
#> eq.label
#> 1 y = -0.00450 + 0.00109 x - 2.14\\times{} 10^{-05} x^2 + 1.06\\times{} 10^{-06} x^3
#> rr.label adj.rr.label rr.confint.label AIC.label
#> 1 R^2 = 0.96 R_{adj}^2 = 0.96 95% CI [0.95, 0.97] \\mathrm{AIC} = -291.9
#> BIC.label f.value.label p.value.label n.label
#> 1 \\mathrm{BIC} = -278.8 F_{3,96} = 810. P < 0.001 \\mathit{n} = 100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
output.type = "text",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA R^2 = 0.96
#> eq.label
#> 1 y = -0.00450 + 0.00109 x - 2.14\\times{} 10^{-05} x^2 + 1.06\\times{} 10^{-06} x^3
#> rr.label adj.rr.label rr.confint.label AIC.label
#> 1 R^2 = 0.96 R_{adj}^2 = 0.96 95% CI [0.95, 0.97] \\mathrm{AIC} = -291.9
#> BIC.label f.value.label p.value.label n.label
#> 1 \\mathrm{BIC} = -278.8 F_{3,96} = 810. P < 0.001 \\mathit{n} = 100
#> grp.label method.label r.squared adj.r.squared p.value n fm.method
#> 1 -1 "method: lm:qr" 0.9620171 0.9608301 5.110349e-68 100 lm:qr
#> fm.class fm.formula fm.formula.chr x y PANEL
#> 1 lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1
#> group orientation
#> 1 -1 x
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug",
output.type = "text",
summary.fun = function(x) {x[["eq.label"]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA R^2 = 0.96
#> eq.label rr.label
#> 1 y = -0.00450 + 0.00109 x - 2.14 10^-05 x^2 + 1.06 10^-06 x^3 R^2 = 0.96
#> adj.rr.label rr.confint.label AIC.label BIC.label f.value.label
#> 1 R_{adj}^2 = 0.96 95% CI [0.95, 0.97] AIC = -291.9 BIC = -278.8 F(3,96) = 810.
#> p.value.label n.label grp.label method.label r.squared adj.r.squared
#> 1 P < 0.001 n = 100 -1 "method: lm:qr" 0.9620171 0.9608301
#> p.value n fm.method fm.class fm.formula
#> 1 5.110349e-68 100 lm:qr lm y ~ poly(x, 3, raw = TRUE)
#> fm.formula.chr x y PANEL group orientation
#> 1 y ~ poly(x, 3, raw = TRUE) 1 1 1 -1 x
# show the content of a list column
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug", output.type = "numeric",
summary.fun = function(x) {x[["coef.ls"]][[1]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA R^2 = 0.96
#> eq.label rr.label
#> 1 y = -0.00450 + 0.00109 x - 2.14 10^-05 x^2 + 1.06 10^-06 x^3 R^2 = 0.96
#> adj.rr.label rr.confint.label AIC.label BIC.label f.value.label
#> 1 R_{adj}^2 = 0.96 95% CI [0.95, 0.97] AIC = -291.9 BIC = -278.8 F(3,96) = 810.
#> p.value.label n.label grp.label method.label r.squared adj.r.squared
#> 1 P < 0.001 n = 100 -1 "method: lm:qr" 0.9620171 0.9608301
#> p.value n fm.method fm.class fm.formula
#> 1 5.110349e-68 100 lm:qr lm y ~ poly(x, 3, raw = TRUE)
#> fm.formula.chr x y PANEL group orientation
#> 1 y ~ poly(x, 3, raw = TRUE) 1 1 1 -1 x
# show the content of a list column
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, geom = "debug", output.type = "numeric",
summary.fun = function(x) {x[["coef.ls"]][[1]]})
#> Warning: Ignoring unknown parameters: `summary.fun`
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA
#> coef.ls
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06, 2.268244e-02, 1.935252e-03, 4.440449e-05, 2.890821e-07, -1.981987e-01, 5.630500e-01, -4.819189e-01, 3.650820e+00, 8.433087e-01, 5.747135e-01, 6.309604e-01, 4.255076e-04
#> coefs r.squared
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06 0.9620171
#> rr.confint.level rr.confint.low rr.confint.high adj.r.squared f.value f.df1
#> 1 0.95 0.9464423 0.9696371 0.9608301 810.484 3
#> f.df2 p.value AIC BIC n rr.label b_0.constant b_0
#> 1 96 5.110349e-68 -291.8639 -278.838 100 FALSE -0.004495631
#> b_1 b_2 b_3 fm.method fm.class
#> 1 0.001089644 -2.139937e-05 1.055387e-06 lm:qr lm
#> fm.formula fm.formula.chr x y PANEL group
#> 1 y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1 -1
#> orientation
#> 1 x
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label
#> 1 NA NA
#> coef.ls
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06, 2.268244e-02, 1.935252e-03, 4.440449e-05, 2.890821e-07, -1.981987e-01, 5.630500e-01, -4.819189e-01, 3.650820e+00, 8.433087e-01, 5.747135e-01, 6.309604e-01, 4.255076e-04
#> coefs r.squared
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06 0.9620171
#> rr.confint.level rr.confint.low rr.confint.high adj.r.squared f.value f.df1
#> 1 0.95 0.9464423 0.9696371 0.9608301 810.484 3
#> f.df2 p.value AIC BIC n rr.label b_0.constant b_0
#> 1 96 5.110349e-68 -291.8639 -278.838 100 FALSE -0.004495631
#> b_1 b_2 b_3 fm.method fm.class
#> 1 0.001089644 -2.139937e-05 1.055387e-06 lm:qr lm
#> fm.formula fm.formula.chr x y PANEL group
#> 1 y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1 1 1 -1
#> orientation
#> 1 x