stat_correlation() applies stats::cor.test()
respecting grouping with method = "pearson" default but
alternatively using "kendall" or "spearman" methods. It
generates labels for correlation coefficients and p-value, coefficient of
determination (R^2) for method "pearson" and number of observations.
stat_correlation(
mapping = NULL,
data = NULL,
geom = "text_npc",
position = "identity",
...,
method = "pearson",
n.min = 2L,
alternative = "two.sided",
exact = NULL,
r.conf.level = ifelse(method == "pearson", 0.95, NA),
continuity = FALSE,
small.r = getOption("ggpmisc.small.r", default = FALSE),
small.p = getOption("ggpmisc.small.p", default = FALSE),
coef.keep.zeros = TRUE,
r.digits = 2,
t.digits = 3,
p.digits = 3,
CI.brackets = c("[", "]"),
label.x = "left",
label.y = "top",
hstep = 0,
vstep = NULL,
output.type = NULL,
boot.R = ifelse(method == "pearson", 0, 999),
na.rm = FALSE,
parse = NULL,
show.legend = FALSE,
inherit.aes = TRUE
)Arguments
- mapping
The aesthetic mapping, usually constructed with
aes. Only needs to be set at the layer level if you are overriding the plot defaults.- data
A layer specific dataset, only needed if you want to override the plot defaults.
- geom
The geometric object to use display the data
- position
The position adjustment to use for overlapping points on this layer
- ...
other arguments passed on to
layer. This can include aesthetics whose values you want to set, not map. Seelayerfor more details.- method
character One of "pearson", "kendall" or "spearman".
- n.min
integer Minimum number of distinct values in the variables for fitting to the attempted.
- alternative
character One of "two.sided", "less" or "greater".
- exact
logical Whether an exact p-value should be computed. Used for Kendall's tau and Spearman's rho.
- r.conf.level
numeric Confidence level for the returned confidence interval. If set to
NAcomputation of CI is skipped.- continuity
logical If TRUE , a continuity correction is used for Kendall's tau and Spearman's rho when not computed exactly.
- small.r, small.p
logical Flags to switch use of lower case r and p for coefficient of correlation (only for
method = "pearson") and p-value.- coef.keep.zeros
logical Keep or drop trailing zeros when formatting the correlation coefficients and t-value, z-value or S-value (see note below).
- r.digits, t.digits, p.digits
integer Number of digits after the decimal point to use for R, r.squared, tau or rho and P-value in labels. If
Inf, use exponential notation with three decimal places.- CI.brackets
character vector of length 2. The opening and closing brackets used for the CI label.
- label.x, label.y
numericwith range 0..1 "normalized parent coordinates" (npc units) or character if usinggeom_text_npc()orgeom_label_npc(). If usinggeom_text()orgeom_label()numeric in native data units. If too short they will be recycled.- hstep, vstep
numeric in npc units, the horizontal and vertical displacement step-size used between labels for different groups.
- output.type
character One of "expression", "LaTeX", "text", "markdown" or "numeric".
- boot.R
interger The number of bootstrap resamples. Set to zero for no bootstrap estimates for the CI.
- na.rm
a logical indicating whether NA values should be stripped before the computation proceeds.
- parse
logical Passed to the geom. If
TRUE, the labels will be parsed into expressions and displayed as described in?plotmath. Default isTRUEifoutput.type = "expression"andFALSEotherwise.- show.legend
logical. Should this layer be included in the legends?
NA, the default, includes if any aesthetics are mapped.FALSEnever includes, andTRUEalways includes.- inherit.aes
If
FALSE, overrides the default aesthetics, rather than combining with them. This is most useful for helper functions that define both data and aesthetics and shouldn't inherit behaviour from the default plot specification, e.g.borders.
Details
This statistic can be used to annotate a plot with the correlation coefficient and the outcome of its test of significance. It supports Pearson, Kendall and Spearman methods to compute correlation. This statistic generates labels as R expressions by default but LaTeX (use TikZ device), markdown (use package 'ggtext') and plain text are also supported, as well as numeric values for user-generated text labels. The character labels include the symbol describing the quantity together with the numeric value. For the confidence interval (CI) the default is to follow the APA recommendation of using square brackets.
The value of parse is set automatically based on output-type,
but if you assemble labels that need parsing from numeric output,
the default needs to be overridden. By default the value of
output.type is guessed from the name of the geometry.
A ggplot statistic receives as data a data frame that is not the one
passed as argument by the user, but instead a data frame with the variables
mapped to aesthetics. cor.test() is always applied to the variables
mapped to the x and y aesthetics, so the scales used for
x and y should both be continuous scales rather than
discrete.
Note
Currently coef.keep.zeros is ignored, with trailing zeros always
retained in the labels but not protected from being dropped by R when
character strings are parsed into expressions.
Aesthetics
stat_correaltion() requires x and
y. In addition, the aesthetics understood by the geom
("text" is the default) are understood and grouping respected.
Computed variables
If output.type is "numeric" the returned
tibble contains the columns listed below with variations depending on the
method. If the model fit function used does not return a value, the
variable is set to NA_real_.
- x,npcx
x position
- y,npcy
y position
- r, and cor, tau or rho
numeric values for correlation coefficient estimates
- t.value and its df, z.value or S.value
numeric values for statistic estimates
- p.value, n
numeric values.
- r.conf.level
numeric value, as fraction of one.
- r.confint.low
Confidence interval limit for
r.- r.confint.high
Confidence interval limit for
r.- grp.label
Set according to mapping in
aes.- method.label
Set according
methodused.- method, test
character values
If output.type different from "numeric" the returned tibble contains
in addition to the columns listed above those listed below. If the numeric
value is missing the label is set to character(0L).
- r.label, and cor.label, tau.label or rho.label
Correlation coefficient as a character string.
- t.value.label, z.value.label or S.value.label
t-value and degrees of freedom, z-value or S-value as a character string.
- p.value.label
P-value for test against zero, as a character string.
- r.confint.label, and cor.conint.label, tau.confint.label or rho.confint.label
Confidence interval for
r(only withmethod = "pearson").- n.label
Number of observations used in the fit, as a character string.
- grp.label
Set according to mapping in
aes, as a character string.
To explore the computed values returned for a given input we suggest the use
of geom_debug as shown in the last examples below.
See also
cor.test for details on the computations.
Examples
# generate artificial data
set.seed(4321)
x <- (1:100) / 10
y <- x + rnorm(length(x))
my.data <- data.frame(x = x,
y = y,
y.desc = - y,
group = c("A", "B"))
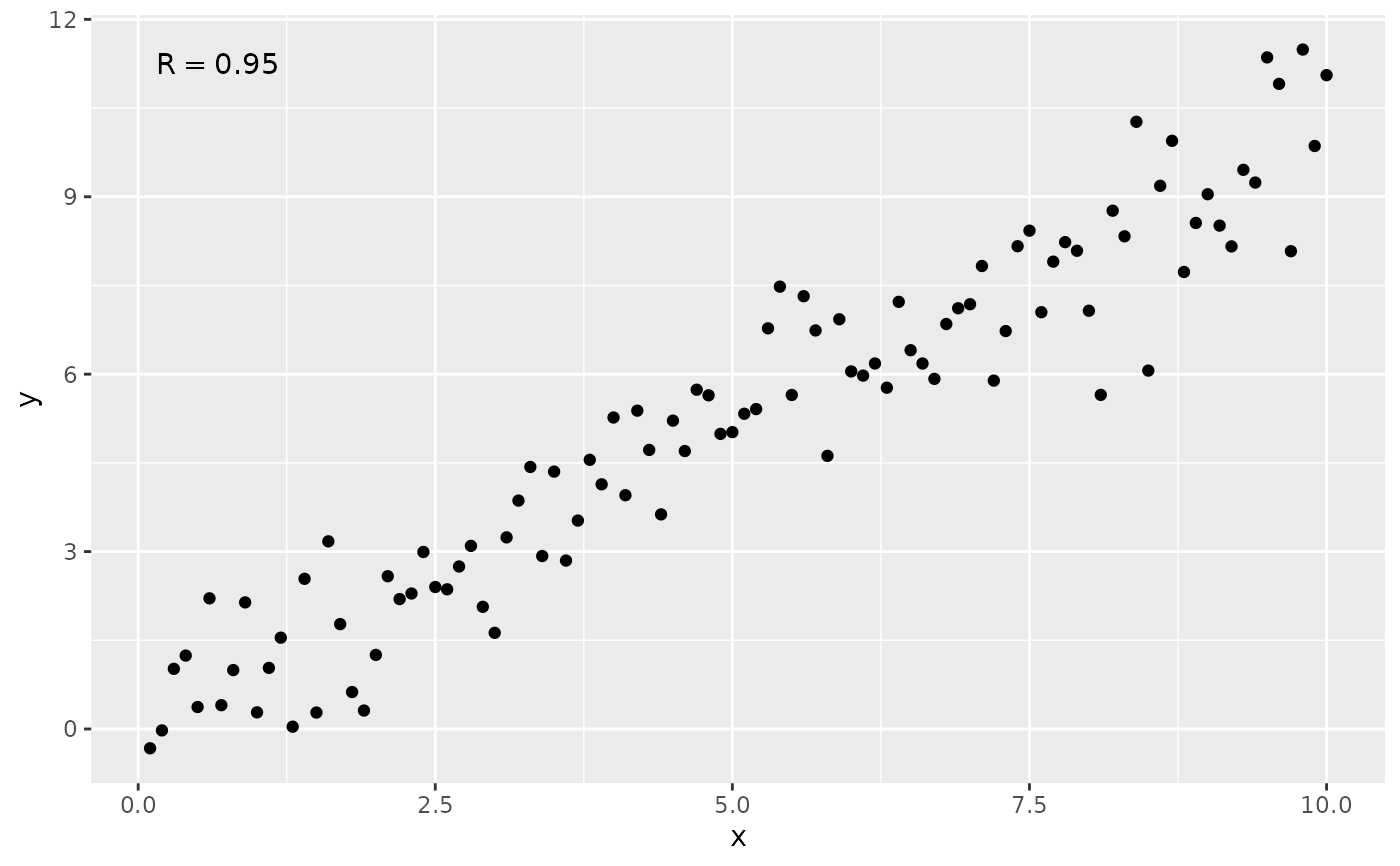
# by default only R is displayed
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation()
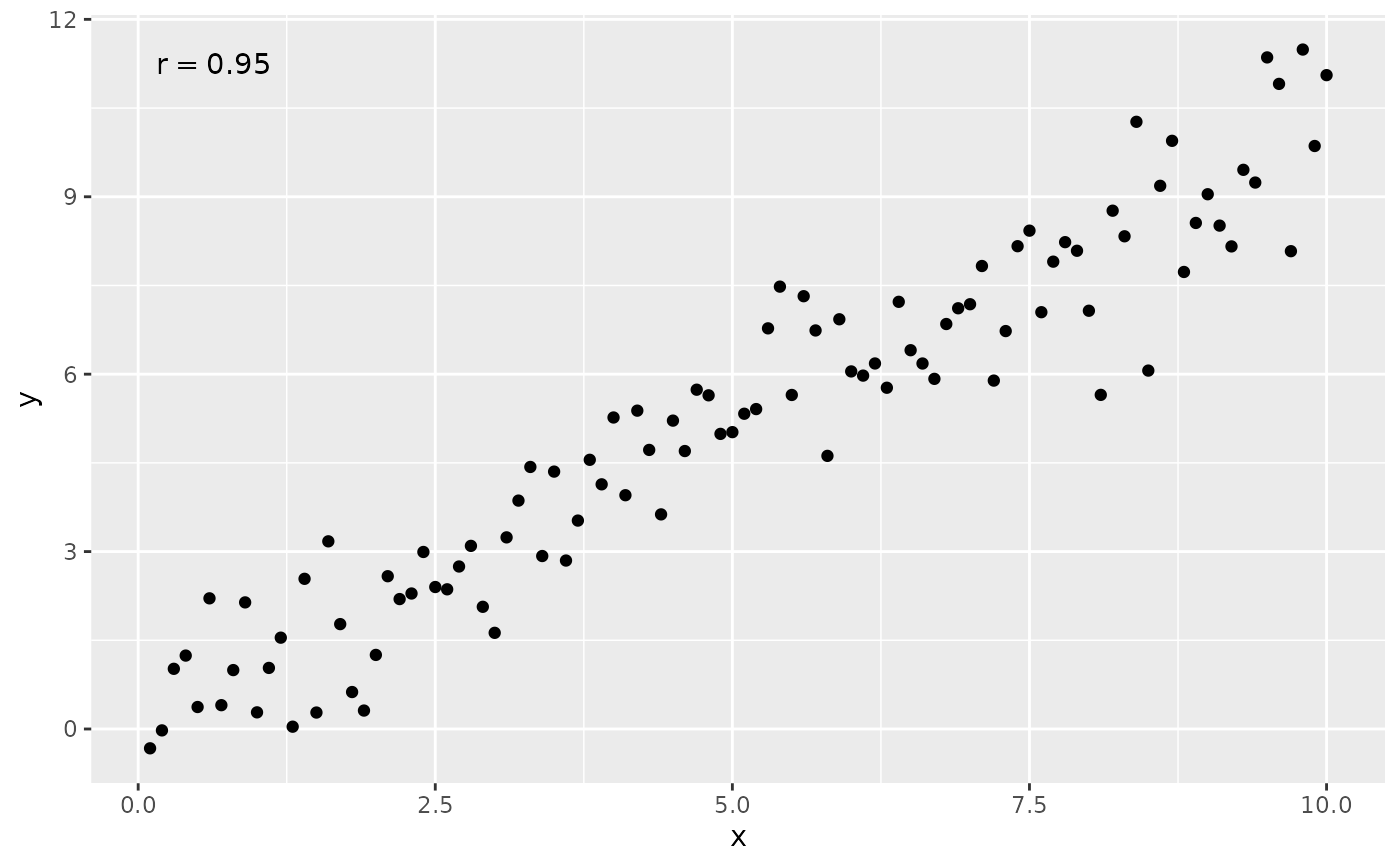
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(small.r = TRUE)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(small.r = TRUE)
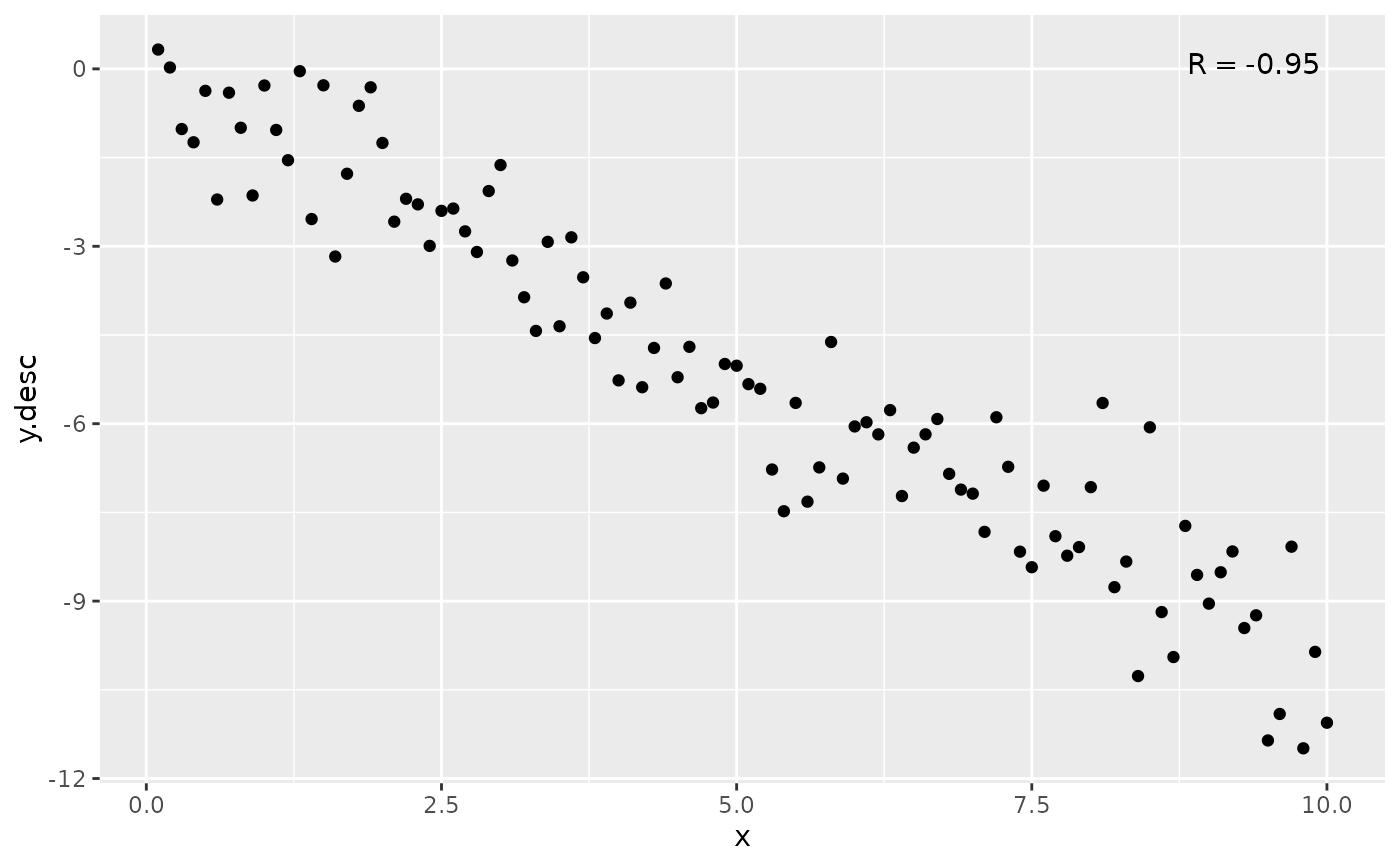
 ggplot(my.data, aes(x, y.desc)) +
geom_point() +
stat_correlation(label.x = "right")
ggplot(my.data, aes(x, y.desc)) +
geom_point() +
stat_correlation(label.x = "right")
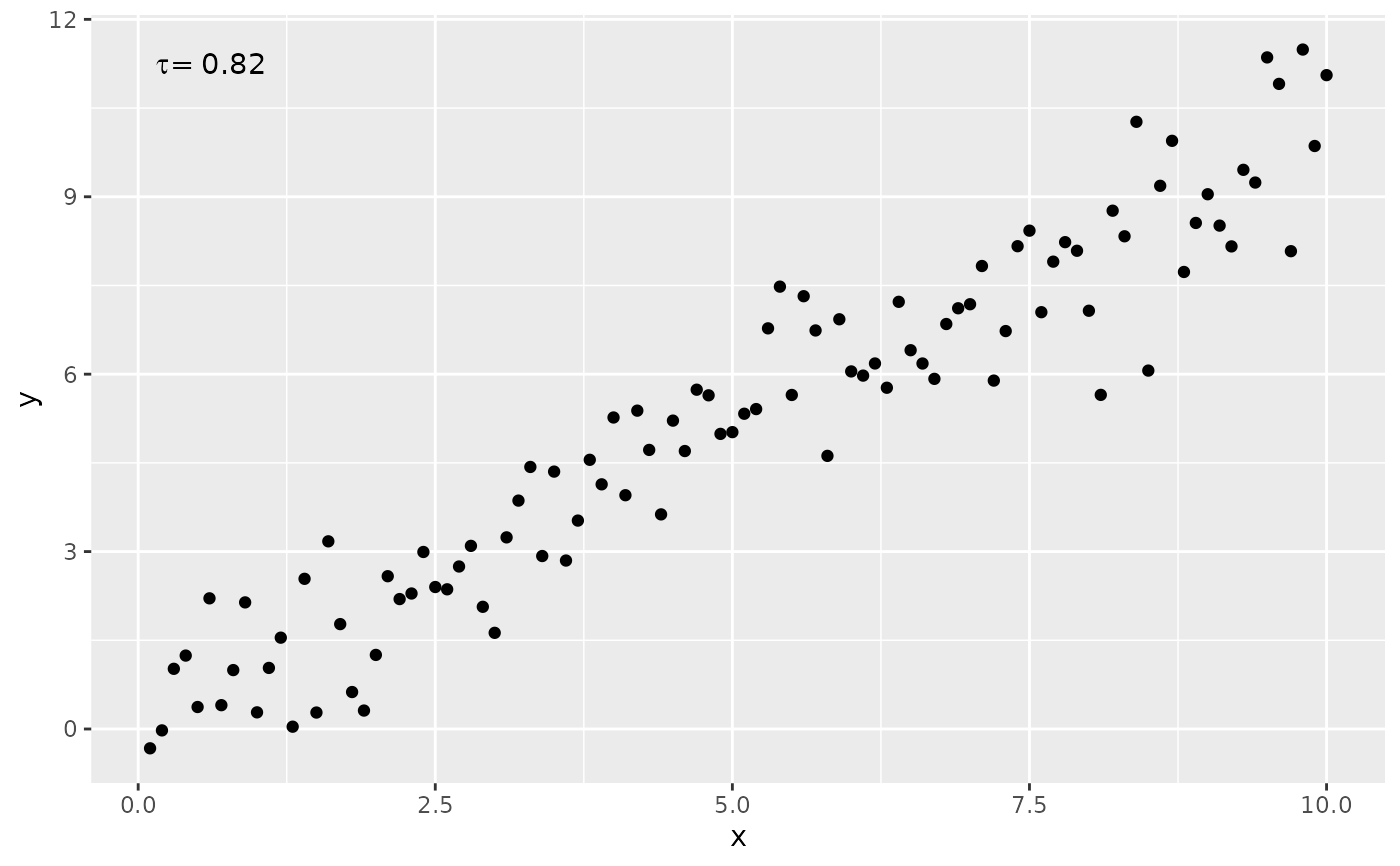
 # non-default methods
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(method = "kendall")
# non-default methods
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(method = "kendall")
 ggplot(my.data, aes(x, y)) +
geom_point() +
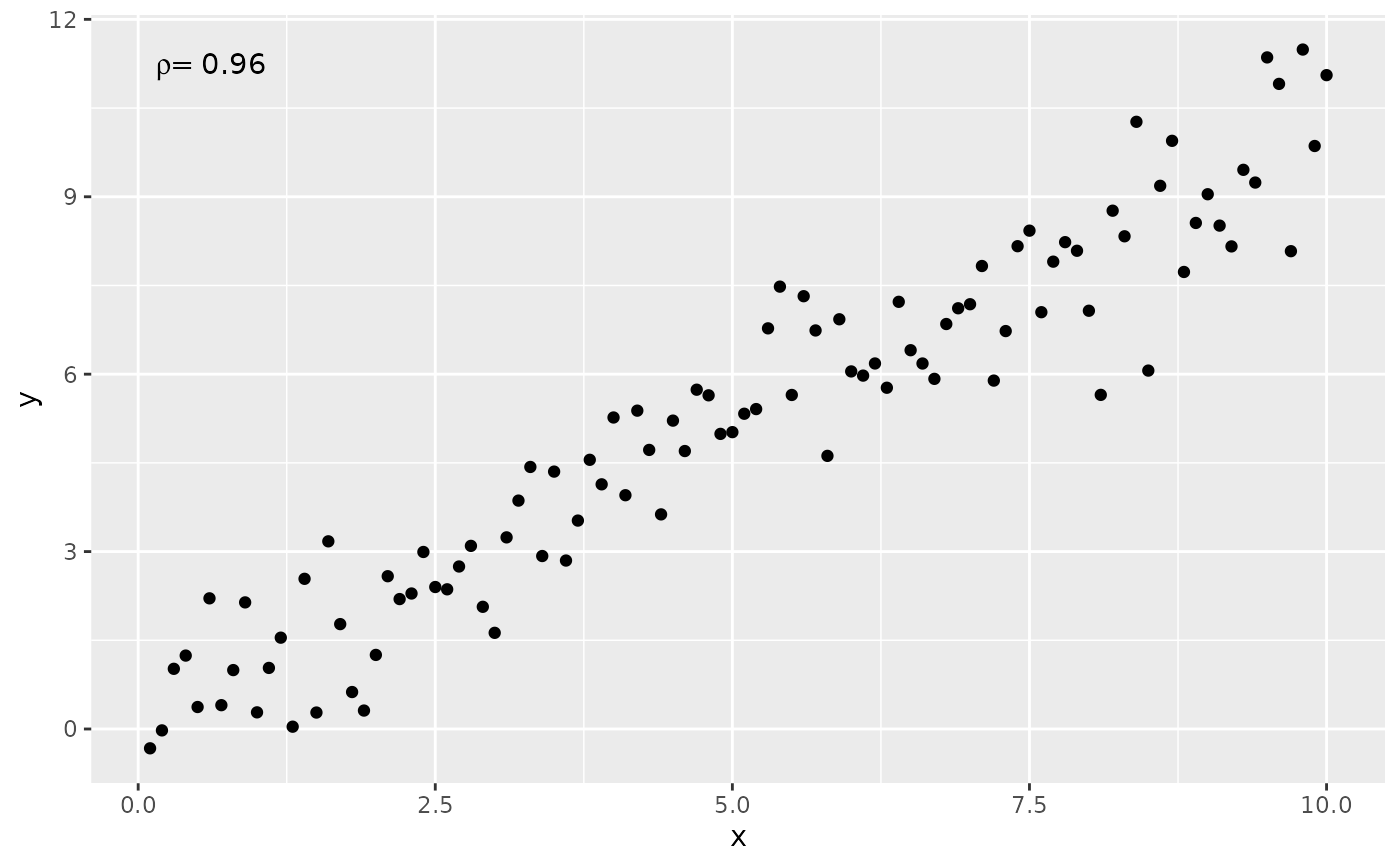
stat_correlation(method = "spearman")
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(method = "spearman")
 # use_label() can map a user selected label
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R2"))
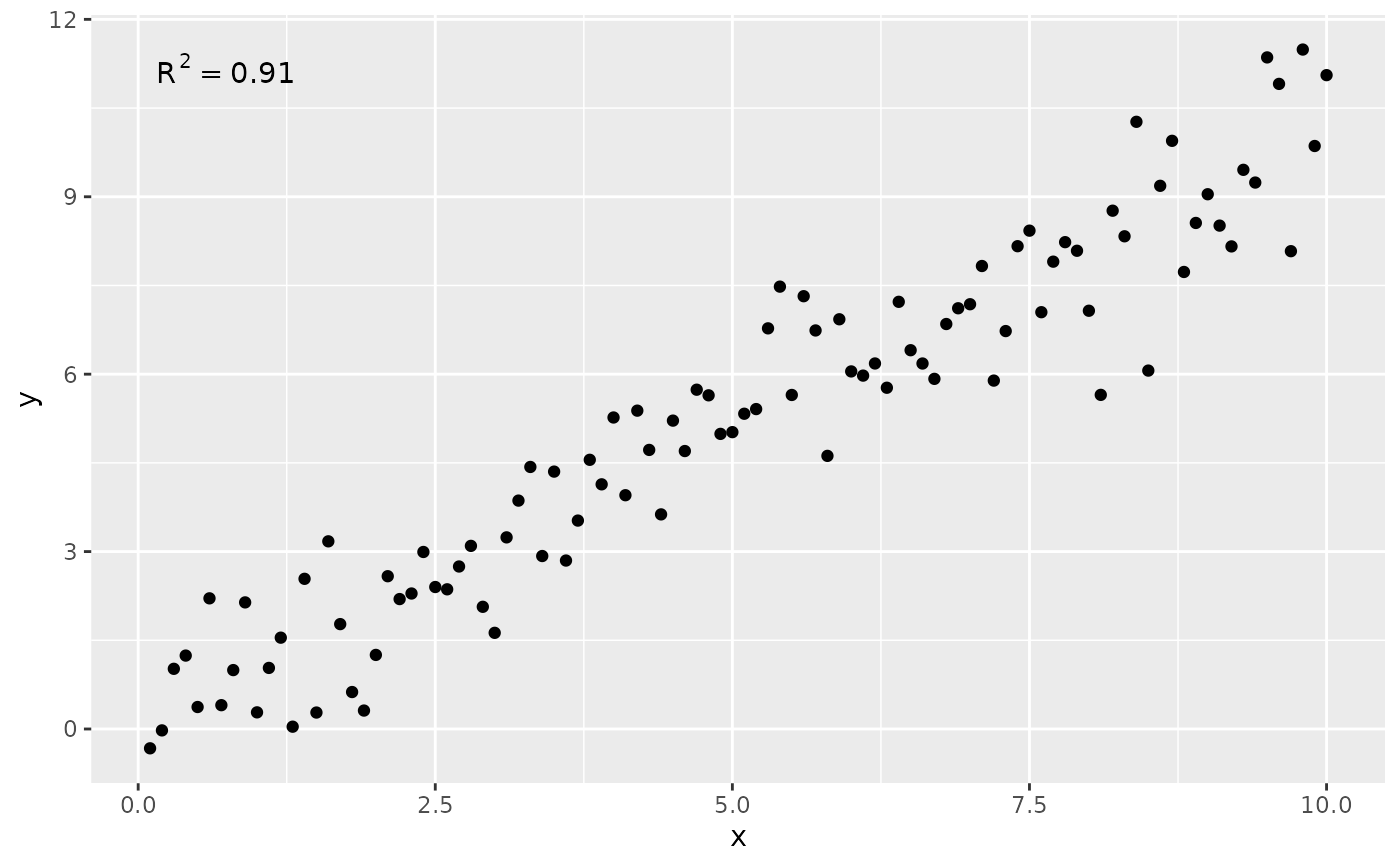
# use_label() can map a user selected label
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R2"))
 # use_label() can assemble and map a combined label
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "P", "n", "method"))
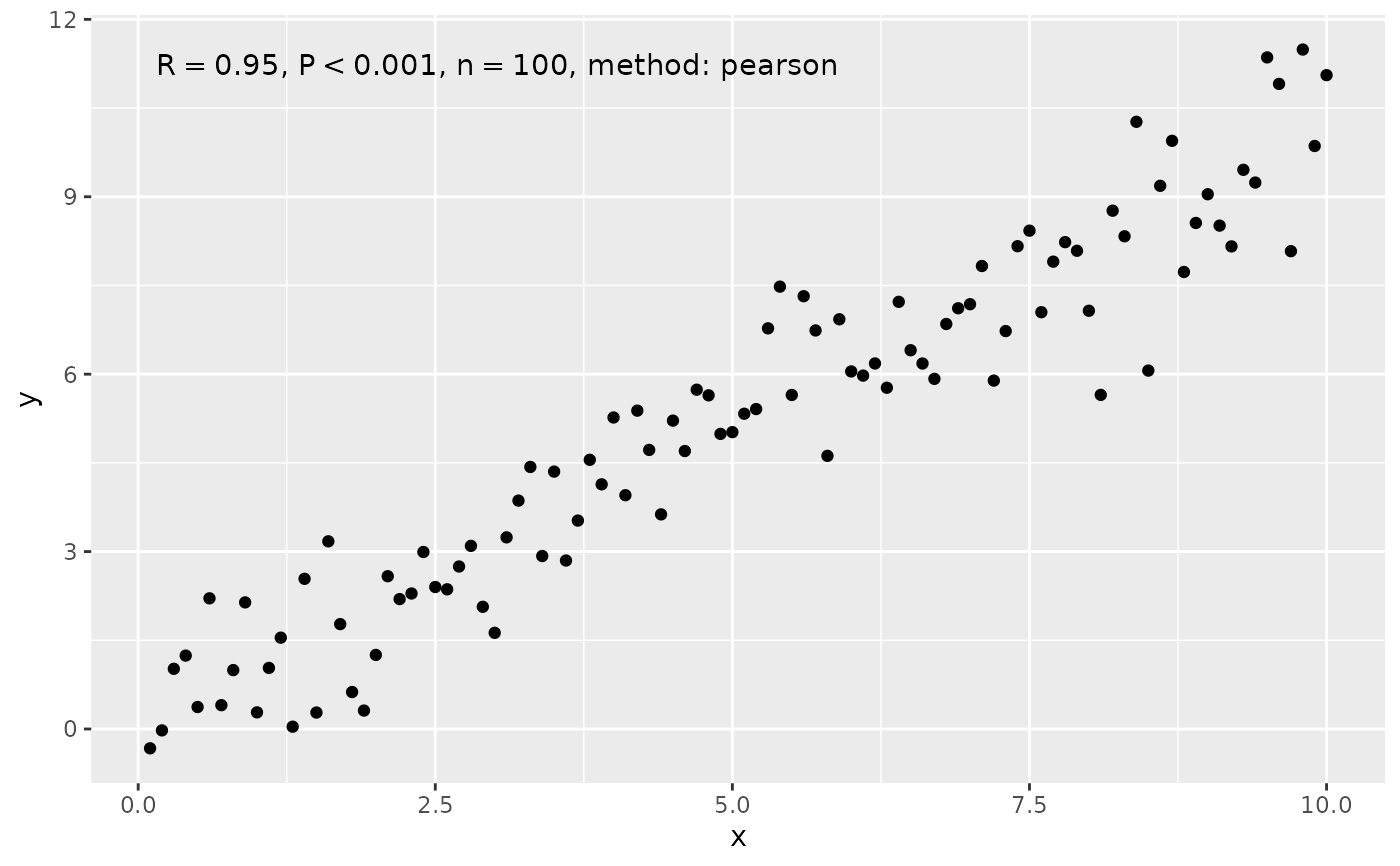
# use_label() can assemble and map a combined label
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "P", "n", "method"))
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"))
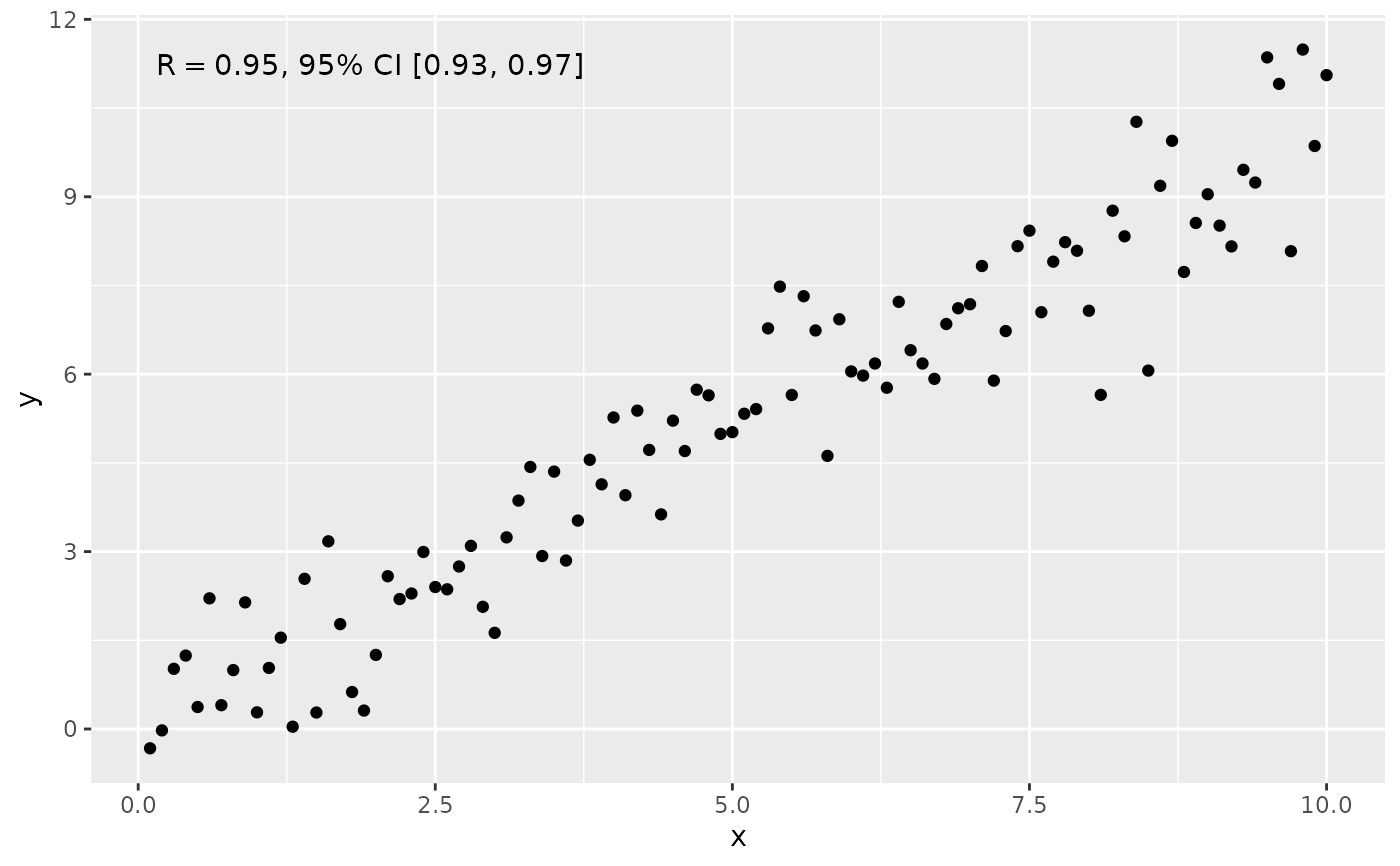
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"))
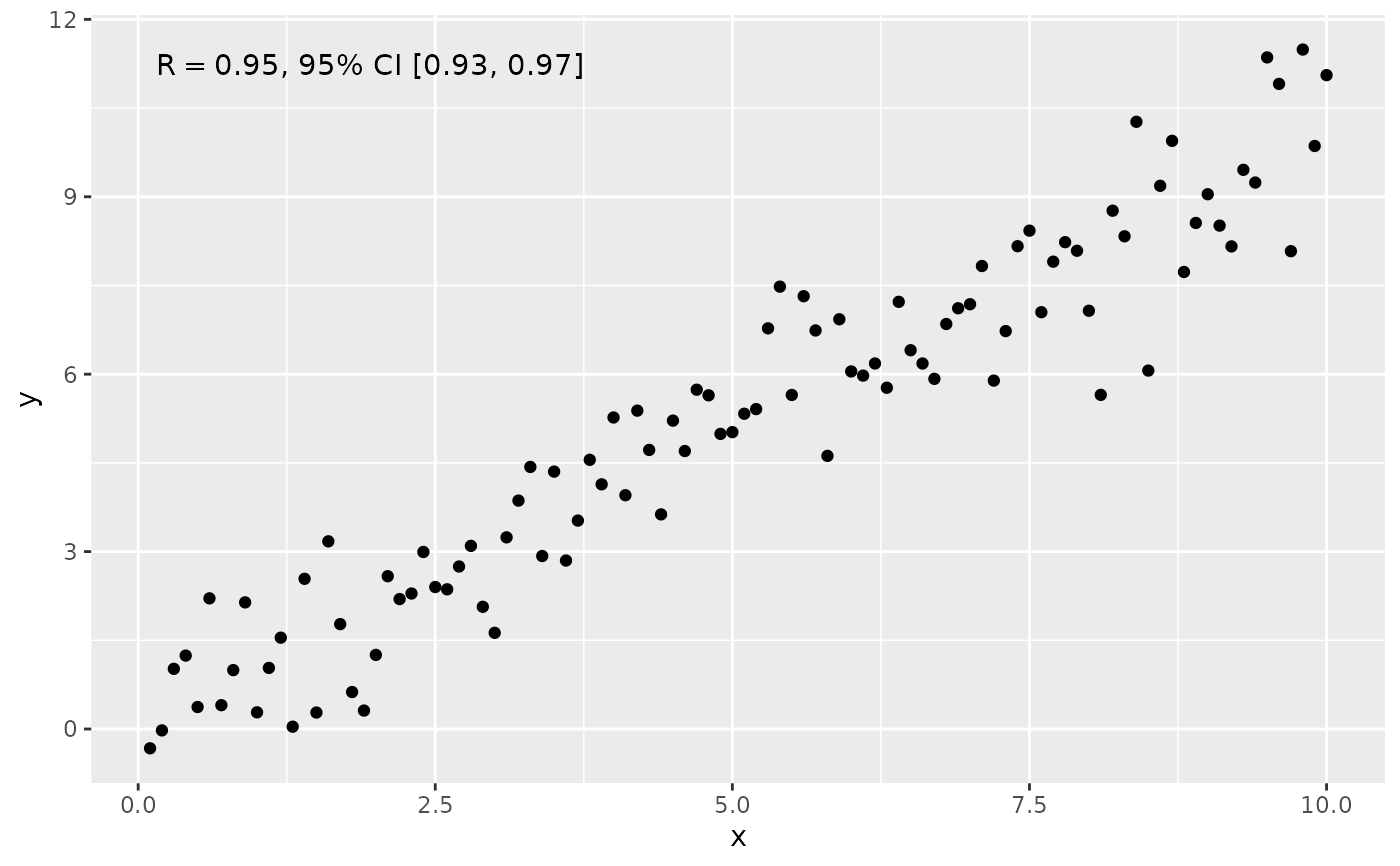
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"),
r.conf.level = 0.95)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"),
r.conf.level = 0.95)
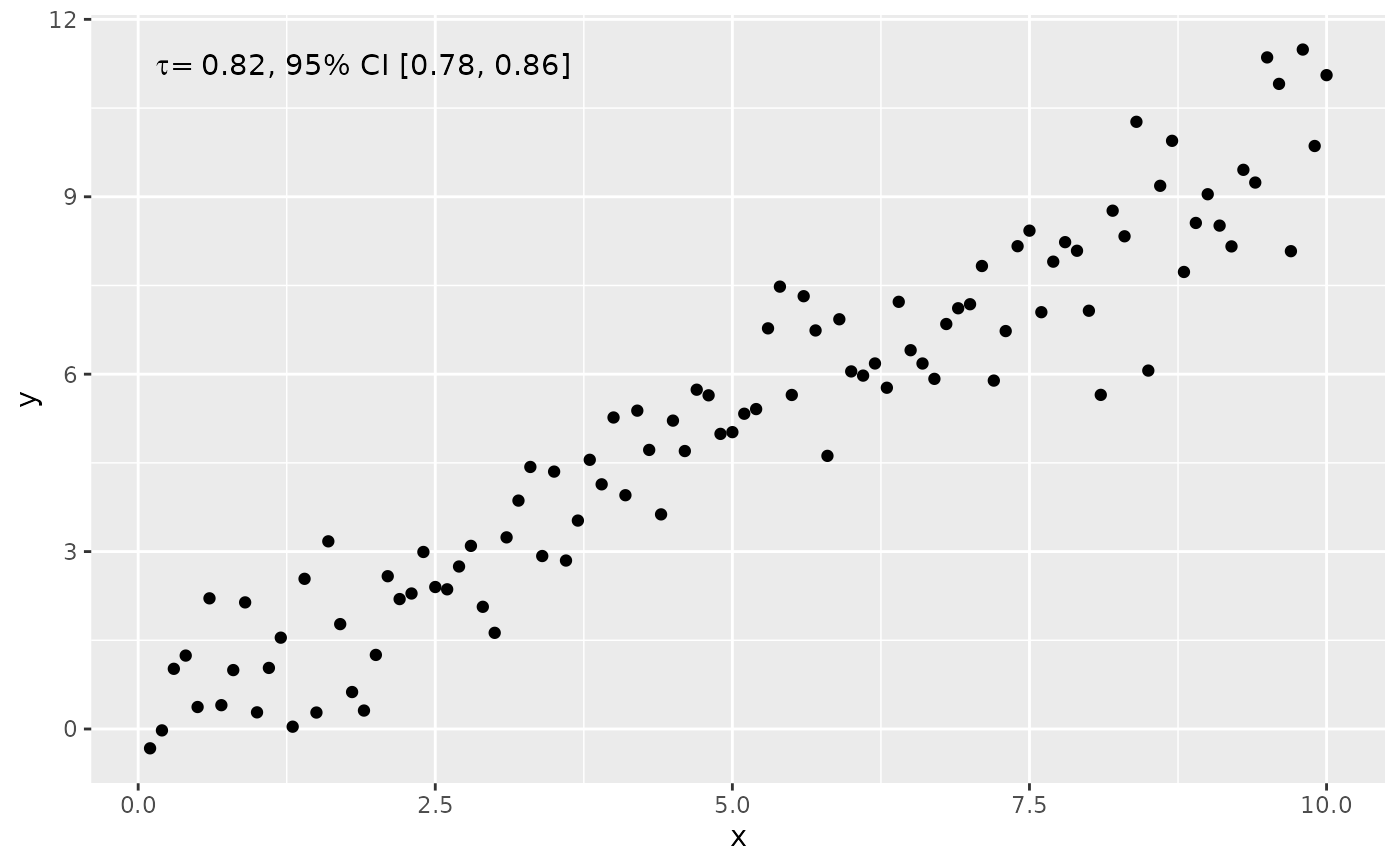
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"),
method = "kendall",
r.conf.level = 0.95)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"),
method = "kendall",
r.conf.level = 0.95)
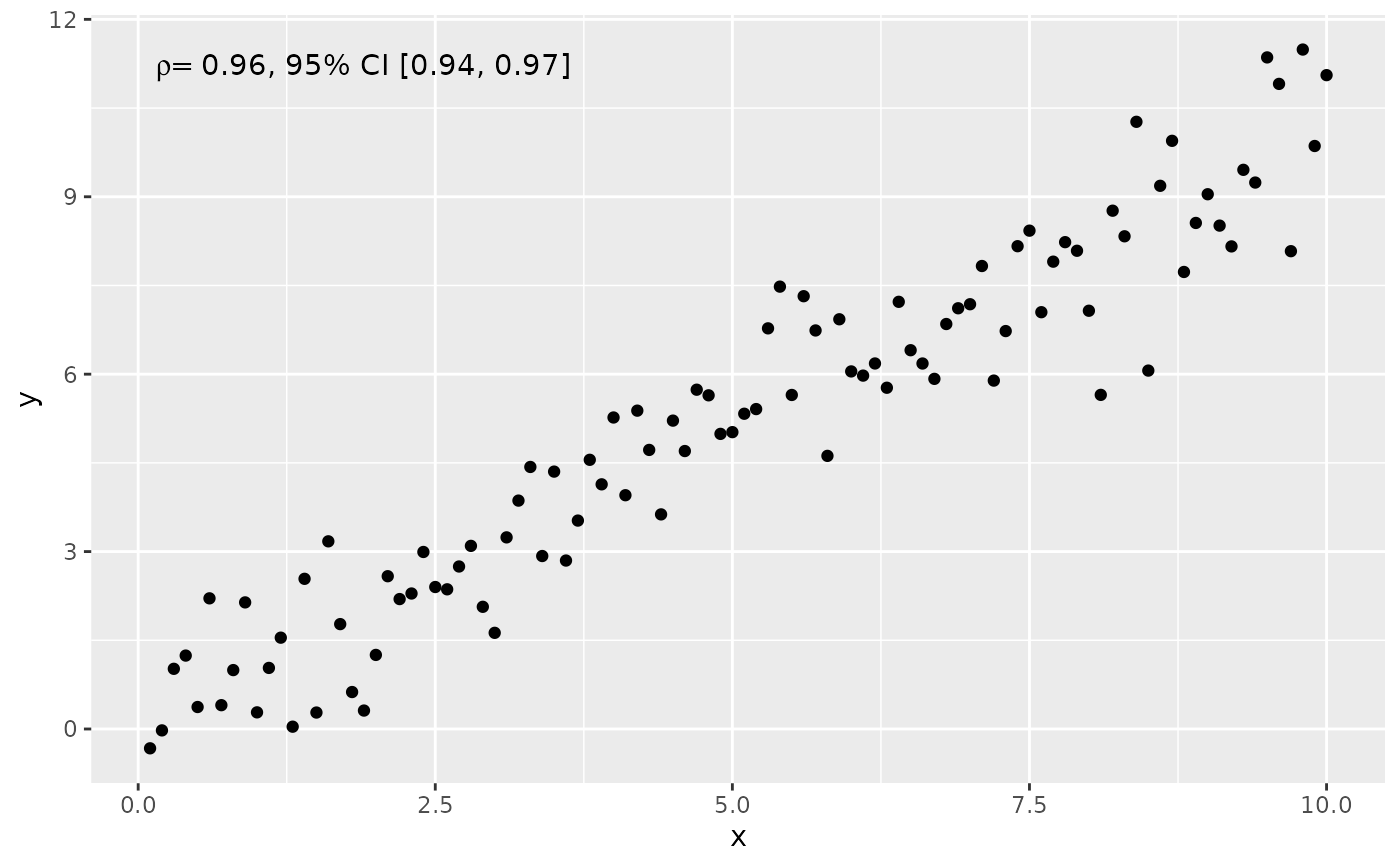
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"),
method = "spearman",
r.conf.level = 0.95)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(use_label("R", "R.CI"),
method = "spearman",
r.conf.level = 0.95)
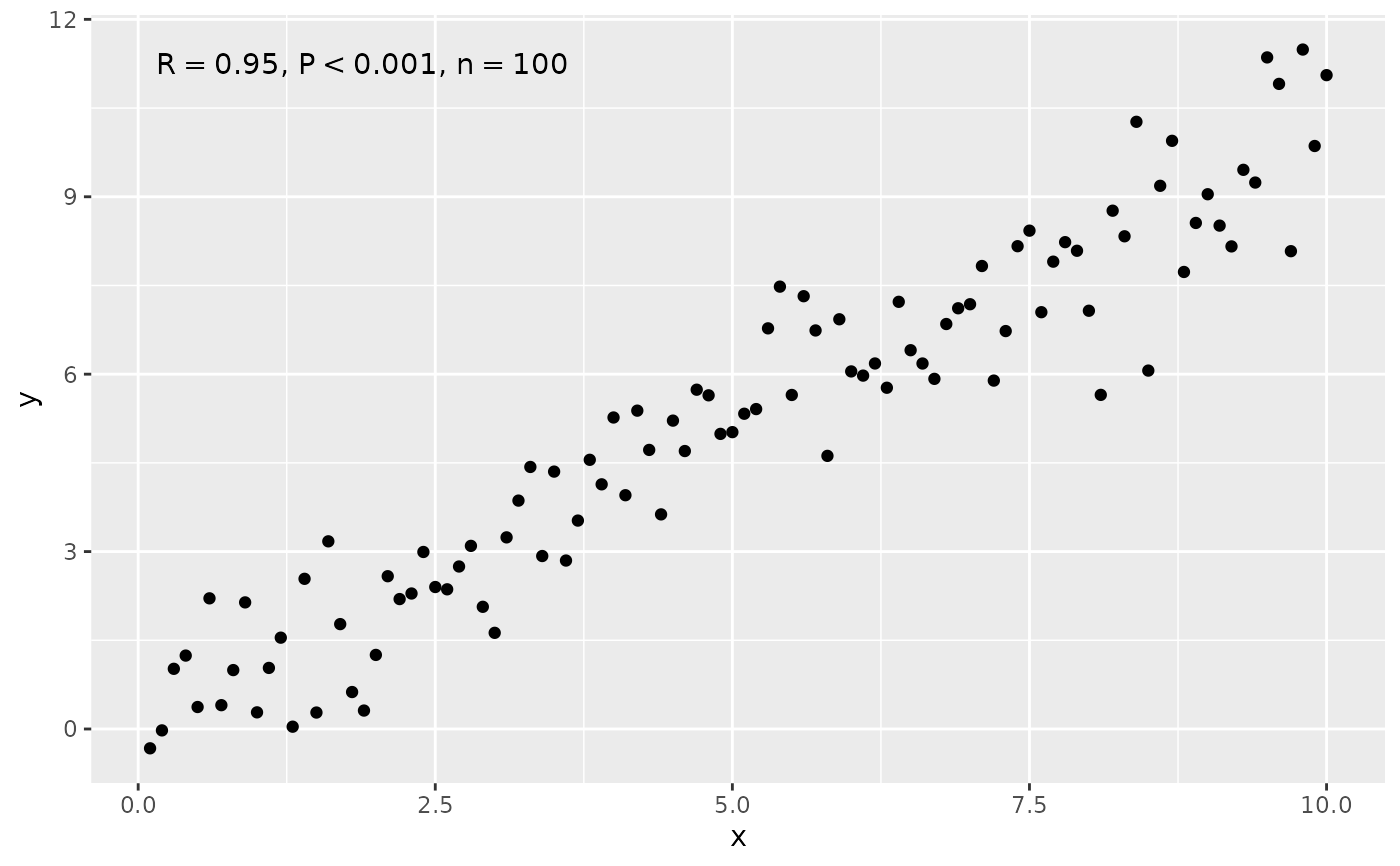
 # manually assemble and map a specific label using paste() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(aes(label = paste(after_stat(r.label),
after_stat(p.value.label),
after_stat(n.label),
sep = "*\", \"*")))
# manually assemble and map a specific label using paste() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(aes(label = paste(after_stat(r.label),
after_stat(p.value.label),
after_stat(n.label),
sep = "*\", \"*")))
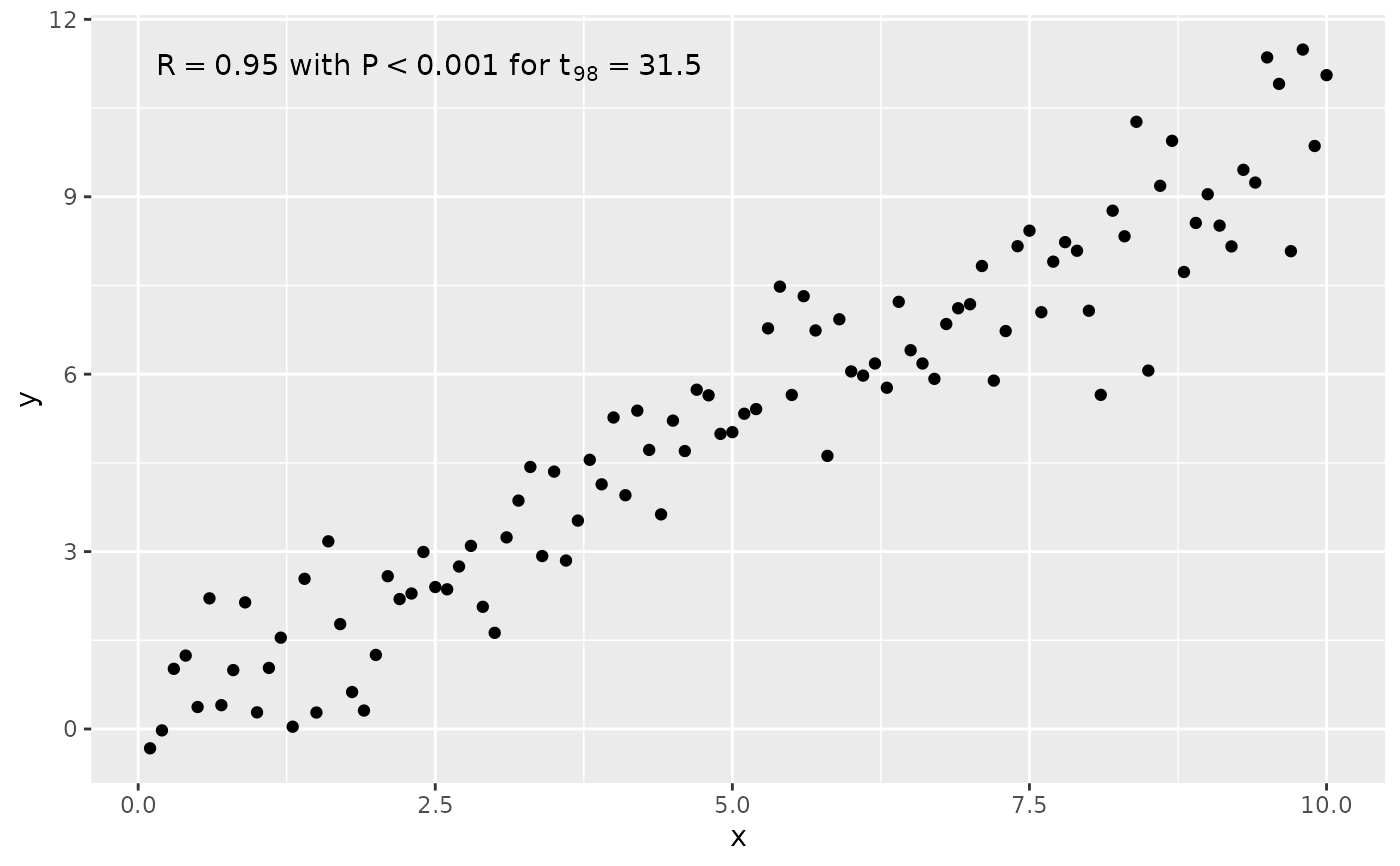
 # manually format and map a specific label using sprintf() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(aes(label = sprintf("%s*\" with \"*%s*\" for \"*%s",
after_stat(r.label),
after_stat(p.value.label),
after_stat(t.value.label))))
# manually format and map a specific label using sprintf() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(aes(label = sprintf("%s*\" with \"*%s*\" for \"*%s",
after_stat(r.label),
after_stat(p.value.label),
after_stat(t.value.label))))
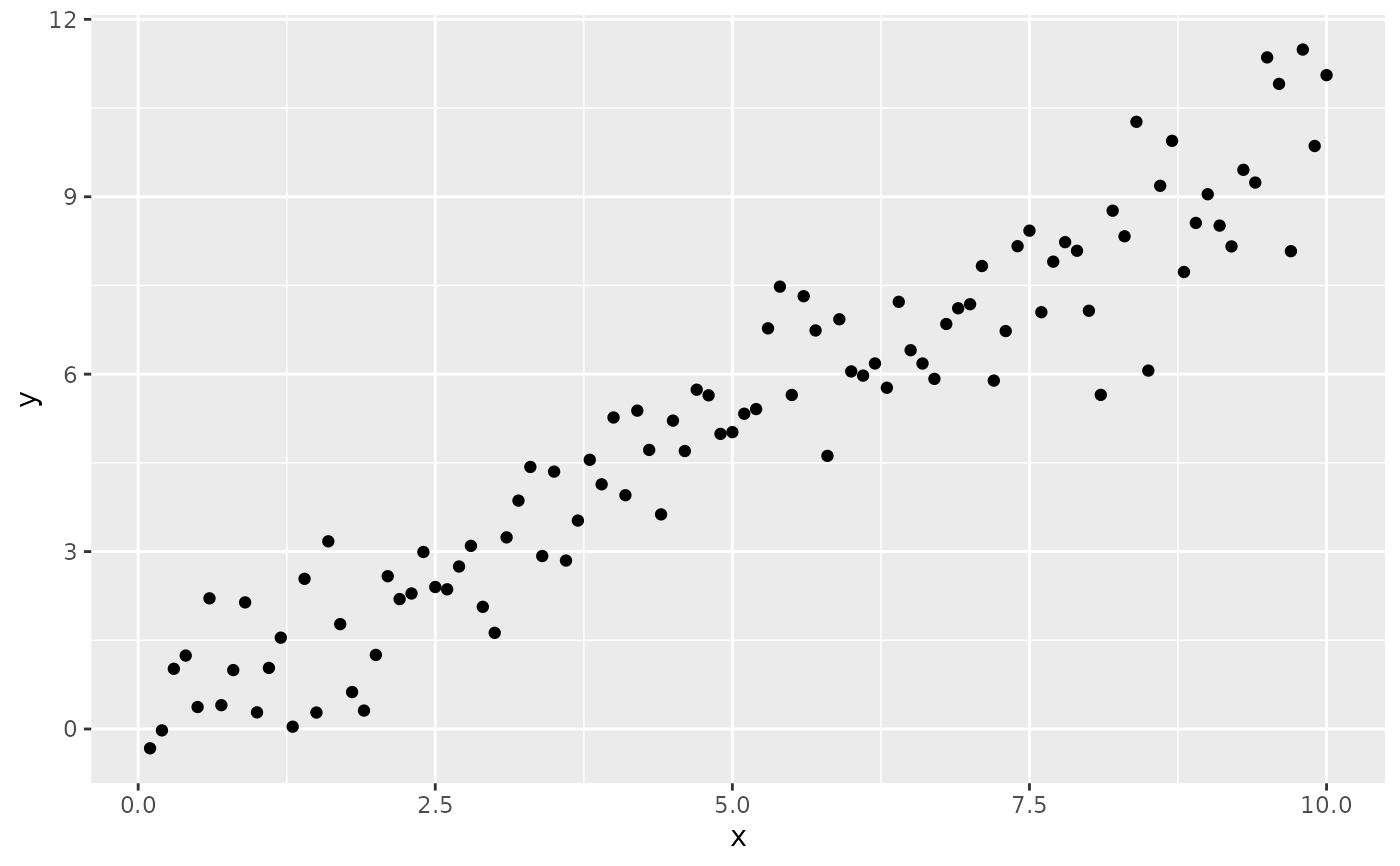
 # Inspecting the returned data using geom_debug()
# This provides a quick way of finding out the names of the variables that
# are available for mapping to aesthetics with after_stat().
gginnards.installed <- requireNamespace("gginnards", quietly = TRUE)
if (gginnards.installed)
library(gginnards)
# the whole of computed data
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug")
# Inspecting the returned data using geom_debug()
# This provides a quick way of finding out the names of the variables that
# are available for mapping to aesthetics with after_stat().
gginnards.installed <- requireNamespace("gginnards", quietly = TRUE)
if (gginnards.installed)
library(gginnards)
# the whole of computed data
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test
#> 1 NA NA italic(R)~`=`~"0.95" 31.54257 98 4.019532e-53 0.9541138 two.sided
#> n method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 100 pearson 0.95 0.9324397 0.9689463 italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(R)~`=`~"0.95" "95% CI [0.93, 0.97]"
#> method.label cor.label rr.label
#> 1 "method: pearson" italic(R)~`=`~"0.95" italic(R)^2~`=`~"0.91"
#> t.value.label x y PANEL group
#> 1 italic(t)[98]~`=`~"31.5" 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", method = "pearson")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test
#> 1 NA NA italic(R)~`=`~"0.95" 31.54257 98 4.019532e-53 0.9541138 two.sided
#> n method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 100 pearson 0.95 0.9324397 0.9689463 italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(R)~`=`~"0.95" "95% CI [0.93, 0.97]"
#> method.label cor.label rr.label
#> 1 "method: pearson" italic(R)~`=`~"0.95" italic(R)^2~`=`~"0.91"
#> t.value.label x y PANEL group
#> 1 italic(t)[98]~`=`~"31.5" 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", method = "pearson")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test
#> 1 NA NA italic(R)~`=`~"0.95" 31.54257 98 4.019532e-53 0.9541138 two.sided
#> n method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 100 pearson 0.95 0.9324397 0.9689463 italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(R)~`=`~"0.95" "95% CI [0.93, 0.97]"
#> method.label cor.label rr.label
#> 1 "method: pearson" italic(R)~`=`~"0.95" italic(R)^2~`=`~"0.91"
#> t.value.label x y PANEL group
#> 1 italic(t)[98]~`=`~"31.5" 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", method = "kendall")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test
#> 1 NA NA italic(R)~`=`~"0.95" 31.54257 98 4.019532e-53 0.9541138 two.sided
#> n method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 100 pearson 0.95 0.9324397 0.9689463 italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(R)~`=`~"0.95" "95% CI [0.93, 0.97]"
#> method.label cor.label rr.label
#> 1 "method: pearson" italic(R)~`=`~"0.95" italic(R)^2~`=`~"0.91"
#> t.value.label x y PANEL group
#> 1 italic(t)[98]~`=`~"31.5" 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", method = "kendall")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label z.value p.value tau test
#> 1 NA NA italic(tau)~`=`~"0.82" 12.13285 7.074566e-34 0.8230303 two.sided
#> n method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 100 kendall 0 NA NA italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(tau)~`=`~"0.82" <NA>
#> method.label tau.label tau.confint.label
#> 1 "method: kendall" italic(tau)~`=`~"0.82" <NA>
#> z.value.label x y PANEL group
#> 1 italic(z)~`=`~"12.1" 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", method = "spearman")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label z.value p.value tau test
#> 1 NA NA italic(tau)~`=`~"0.82" 12.13285 7.074566e-34 0.8230303 two.sided
#> n method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 100 kendall 0 NA NA italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(tau)~`=`~"0.82" <NA>
#> method.label tau.label tau.confint.label
#> 1 "method: kendall" italic(tau)~`=`~"0.82" <NA>
#> z.value.label x y PANEL group
#> 1 italic(z)~`=`~"12.1" 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", method = "spearman")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label S.value p.value rho test n
#> 1 NA NA italic(rho)~`=`~"0.96" 6974 0 0.9581518 two.sided 100
#> method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 spearman 0 NA NA italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(rho)~`=`~"0.96" <NA>
#> method.label rho.label rho.confint.label
#> 1 "method: spearman" italic(rho)~`=`~"0.96" <NA>
#> S.value.label x y PANEL group
#> 1 italic(S)~`=`~"6.97" %*% 10^{"+03"} 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", output.type = "numeric")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label S.value p.value rho test n
#> 1 NA NA italic(rho)~`=`~"0.96" 6974 0 0.9581518 two.sided 100
#> method r.conf.level r.confint.low r.confint.high p.value.label
#> 1 spearman 0 NA NA italic(P)~`<`~"0.001"
#> n.label grp.label r.label r.confint.label
#> 1 italic(n)~`=`~100 -1 italic(rho)~`=`~"0.96" <NA>
#> method.label rho.label rho.confint.label
#> 1 "method: spearman" italic(rho)~`=`~"0.96" <NA>
#> S.value.label x y PANEL group
#> 1 italic(S)~`=`~"6.97" %*% 10^{"+03"} 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", output.type = "numeric")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test n method
#> 1 NA NA <NA> 31.54257 98 4.019532e-53 0.9541138 two.sided 100 pearson
#> r.conf.level r.confint.low r.confint.high r.label x y PANEL group
#> 1 0.95 0.9324397 0.9689463 <NA> 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", output.type = "markdown")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test n method
#> 1 NA NA <NA> 31.54257 98 4.019532e-53 0.9541138 two.sided 100 pearson
#> r.conf.level r.confint.low r.confint.high r.label x y PANEL group
#> 1 0.95 0.9324397 0.9689463 <NA> 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", output.type = "markdown")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test n method
#> 1 NA NA _R_ = 0.95 31.54257 98 4.019532e-53 0.9541138 two.sided 100 pearson
#> r.conf.level r.confint.low r.confint.high p.value.label n.label grp.label
#> 1 0.95 0.9324397 0.9689463 _P_ < 0.001 _n_ = 100 -1
#> r.label r.confint.label method.label cor.label
#> 1 _R_ = 0.95 95% CI [0.93, 0.97] "method: pearson" _R_ = 0.95
#> rr.label t.value.label x y PANEL group
#> 1 _R_<sup>2</sup> = 0.91 _t_<sub>98</sub> = 31.5 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", output.type = "LaTeX")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test n method
#> 1 NA NA _R_ = 0.95 31.54257 98 4.019532e-53 0.9541138 two.sided 100 pearson
#> r.conf.level r.confint.low r.confint.high p.value.label n.label grp.label
#> 1 0.95 0.9324397 0.9689463 _P_ < 0.001 _n_ = 100 -1
#> r.label r.confint.label method.label cor.label
#> 1 _R_ = 0.95 95% CI [0.93, 0.97] "method: pearson" _R_ = 0.95
#> rr.label t.value.label x y PANEL group
#> 1 _R_<sup>2</sup> = 0.91 _t_<sub>98</sub> = 31.5 0.1 11.48941 1 -1
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_correlation(geom = "debug", output.type = "LaTeX")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test n method
#> 1 NA NA R = 0.95 31.54257 98 4.019532e-53 0.9541138 two.sided 100 pearson
#> r.conf.level r.confint.low r.confint.high p.value.label n.label
#> 1 0.95 0.9324397 0.9689463 P < 0.001 \\mathit{n} = 100
#> grp.label r.label r.confint.label method.label cor.label rr.label
#> 1 -1 R = 0.95 95% CI [0.93, 0.97] "method: pearson" R = 0.95 R^2 = 0.91
#> t.value.label x y PANEL group
#> 1 t_{98} = 31.5 0.1 11.48941 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> npcx npcy label t.value df p.value cor test n method
#> 1 NA NA R = 0.95 31.54257 98 4.019532e-53 0.9541138 two.sided 100 pearson
#> r.conf.level r.confint.low r.confint.high p.value.label n.label
#> 1 0.95 0.9324397 0.9689463 P < 0.001 \\mathit{n} = 100
#> grp.label r.label r.confint.label method.label cor.label rr.label
#> 1 -1 R = 0.95 95% CI [0.93, 0.97] "method: pearson" R = 0.95 R^2 = 0.91
#> t.value.label x y PANEL group
#> 1 t_{98} = 31.5 0.1 11.48941 1 -1