Predict Method for Support Vector Machines
predict.svm.RdThis function predicts values based upon a model trained by svm.
Usage
# S3 method for class 'svm'
predict(object, newdata, decision.values = FALSE,
probability = FALSE, ..., na.action = na.omit)Arguments
- object
Object of class
"svm", created bysvm.- newdata
An object containing the new input data: either a matrix or a sparse matrix (object of class
Matrixprovided by the Matrix package, or of classmatrix.csrprovided by the SparseM package, or of classsimple_triplet_matrixprovided by the slam package). A vector will be transformed to a n x 1 matrix.- decision.values
Logical controlling whether the decision values of all binary classifiers computed in multiclass classification shall be computed and returned.
- probability
Logical indicating whether class probabilities should be computed and returned. Only possible if the model was fitted with the
probabilityoption enabled.- na.action
A function to specify the action to be taken if ‘NA’s are found. The default action is
na.omit, which leads to rejection of cases with missing values on any required variable. An alternative isna.fail, which causes an error ifNAcases are found. (NOTE: If given, this argument must be named.)- ...
Currently not used.
Value
A vector of predicted values (for classification: a vector of labels, for density
estimation: a logical vector). If decision.value is
TRUE, the vector gets a "decision.values" attribute
containing a n x c matrix (n number of predicted values, c number of
classifiers) of all c binary classifiers' decision values. There are k
* (k - 1) / 2 classifiers (k number of classes). The colnames of
the matrix indicate the labels of the two classes. If probability is
TRUE, the vector gets a "probabilities" attribute
containing a n x k matrix (n number of predicted values, k number of
classes) of the class probabilities.
Note
If the training set was scaled by svm (done by default), the
new data is scaled accordingly using scale and center of
the training data.
Author
David Meyer (based on C++-code by Chih-Chung Chang and Chih-Jen Lin)
David.Meyer@R-project.org
Examples
data(iris)
attach(iris)
## classification mode
# default with factor response:
model <- svm(Species ~ ., data = iris)
# alternatively the traditional interface:
x <- subset(iris, select = -Species)
y <- Species
model <- svm(x, y, probability = TRUE)
print(model)
#>
#> Call:
#> svm.default(x = x, y = y, probability = TRUE)
#>
#>
#> Parameters:
#> SVM-Type: C-classification
#> SVM-Kernel: radial
#> cost: 1
#>
#> Number of Support Vectors: 51
#>
summary(model)
#>
#> Call:
#> svm.default(x = x, y = y, probability = TRUE)
#>
#>
#> Parameters:
#> SVM-Type: C-classification
#> SVM-Kernel: radial
#> cost: 1
#>
#> Number of Support Vectors: 51
#>
#> ( 8 22 21 )
#>
#>
#> Number of Classes: 3
#>
#> Levels:
#> setosa versicolor virginica
#>
#>
#>
# test with train data
pred <- predict(model, x)
# (same as:)
pred <- fitted(model)
# compute decision values and probabilites
pred <- predict(model, x, decision.values = TRUE, probability = TRUE)
attr(pred, "decision.values")[1:4,]
#> setosa/versicolor setosa/virginica versicolor/virginica
#> 1 1.196152 1.091757 0.6708810
#> 2 1.064621 1.056185 0.8483518
#> 3 1.180842 1.074542 0.6439798
#> 4 1.110699 1.053012 0.6782041
attr(pred, "probabilities")[1:4,]
#> setosa versicolor virginica
#> 1 0.9799440 0.01105443 0.009001534
#> 2 0.9726238 0.01772298 0.009653187
#> 3 0.9786104 0.01166303 0.009726537
#> 4 0.9745697 0.01497637 0.010453974
## try regression mode on two dimensions
# create data
x <- seq(0.1, 5, by = 0.05)
y <- log(x) + rnorm(x, sd = 0.2)
# estimate model and predict input values
m <- svm(x, y)
new <- predict(m, x)
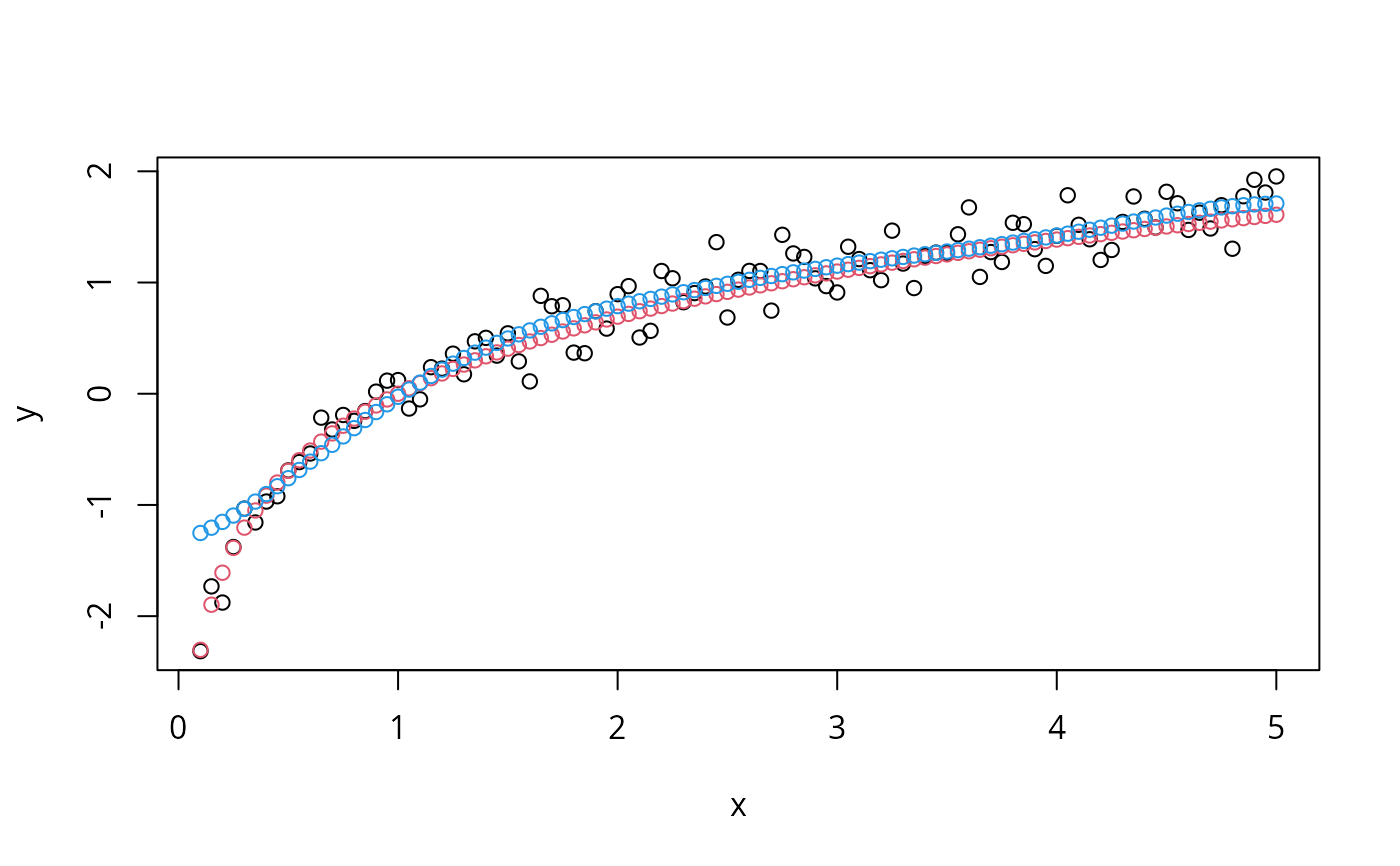
# visualize
plot (x, y)
points (x, log(x), col = 2)
points (x, new, col = 4)
 ## density-estimation
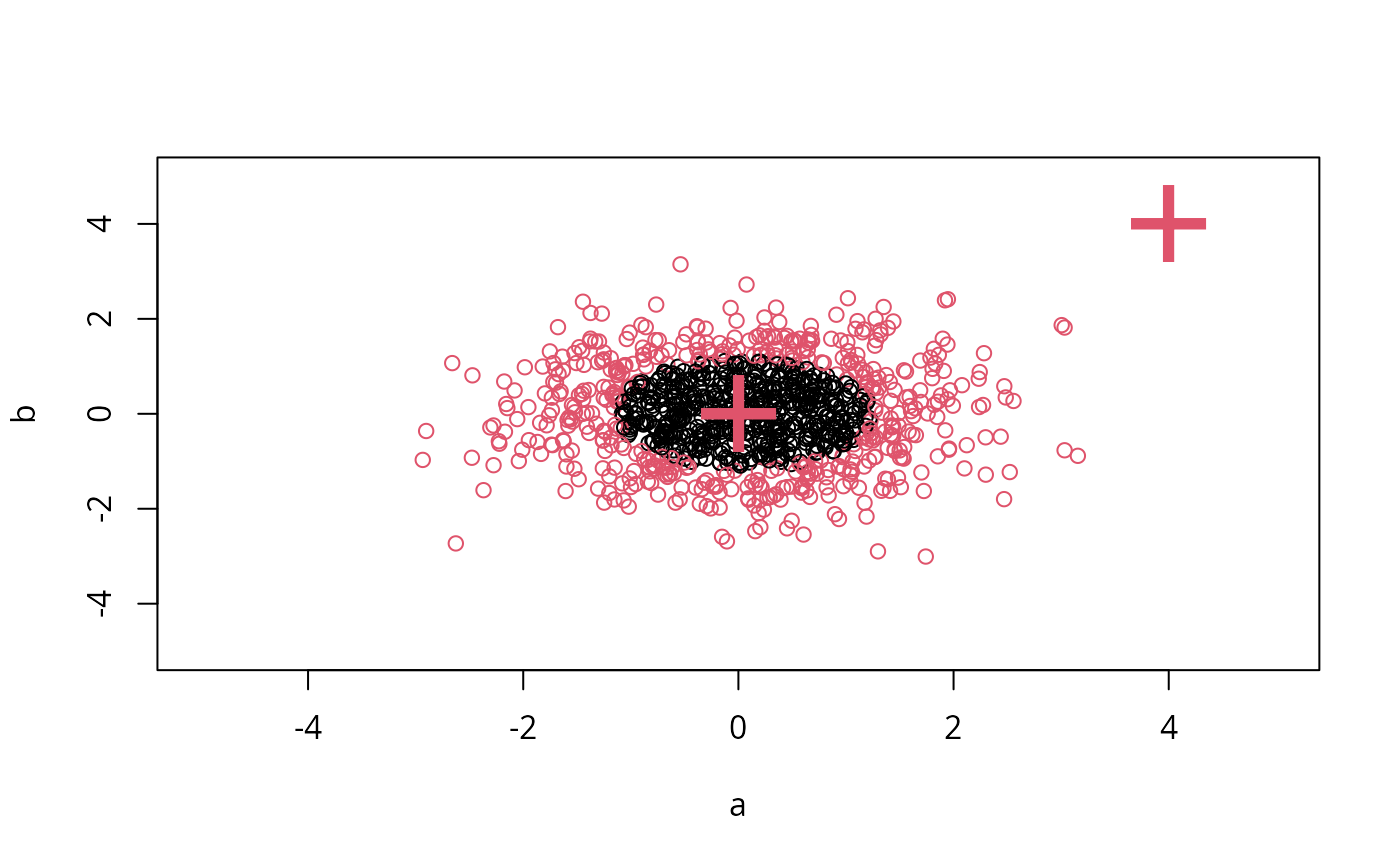
# create 2-dim. normal with rho=0:
X <- data.frame(a = rnorm(1000), b = rnorm(1000))
attach(X)
# traditional way:
m <- svm(X, gamma = 0.1)
# formula interface:
m <- svm(~., data = X, gamma = 0.1)
# or:
m <- svm(~ a + b, gamma = 0.1)
# test:
newdata <- data.frame(a = c(0, 4), b = c(0, 4))
predict (m, newdata)
#> 1 2
#> TRUE FALSE
# visualize:
plot(X, col = 1:1000 %in% m$index + 1, xlim = c(-5,5), ylim=c(-5,5))
points(newdata, pch = "+", col = 2, cex = 5)
## density-estimation
# create 2-dim. normal with rho=0:
X <- data.frame(a = rnorm(1000), b = rnorm(1000))
attach(X)
# traditional way:
m <- svm(X, gamma = 0.1)
# formula interface:
m <- svm(~., data = X, gamma = 0.1)
# or:
m <- svm(~ a + b, gamma = 0.1)
# test:
newdata <- data.frame(a = c(0, 4), b = c(0, 4))
predict (m, newdata)
#> 1 2
#> TRUE FALSE
# visualize:
plot(X, col = 1:1000 %in% m$index + 1, xlim = c(-5,5), ylim=c(-5,5))
points(newdata, pch = "+", col = 2, cex = 5)