Genetic Components of Alcoholism
alpha.RdLevels of expressed alpha synuclein mRNA in three groups of allele lengths of NACP-REP1.
alphaFormat
A data frame with 97 observations on 2 variables.
alengthallele length, a factor with levels
"short","intermediate"and"long".elevelexpression levels of alpha synuclein mRNA.
Details
Various studies have linked alcohol dependence phenotypes to chromosome 4. One candidate gene is NACP (non-amyloid component of plaques), coding for alpha synuclein. Bönsch et al. (2005) found longer alleles of NACP-REP1 in alcohol-dependent patients compared with healthy controls and reported that the allele lengths show some association with levels of expressed alpha synuclein mRNA.
Source
Bönsch, D., Lederer, T., Reulbach, U., Hothorn, T., Kornhuber, J. and Bleich, S. (2005). Joint analysis of the NACP-REP1 marker within the alpha synuclein gene concludes association with alcohol dependence. Human Molecular Genetics 14(7), 967–971. doi:10.1093/hmg/ddi090
References
Hothorn, T., Hornik, K., van de Wiel, M. A. and Zeileis, A. (2006). A Lego system for conditional inference. The American Statistician 60(3), 257–263. doi:10.1198/000313006X118430
Winell, H. and Lindbäck, J. (2018). A general score-independent test for order-restricted inference. Statistics in Medicine 37(21), 3078–3090. doi:10.1002/sim.7690
Examples
## Boxplots
boxplot(elevel ~ alength, data = alpha)
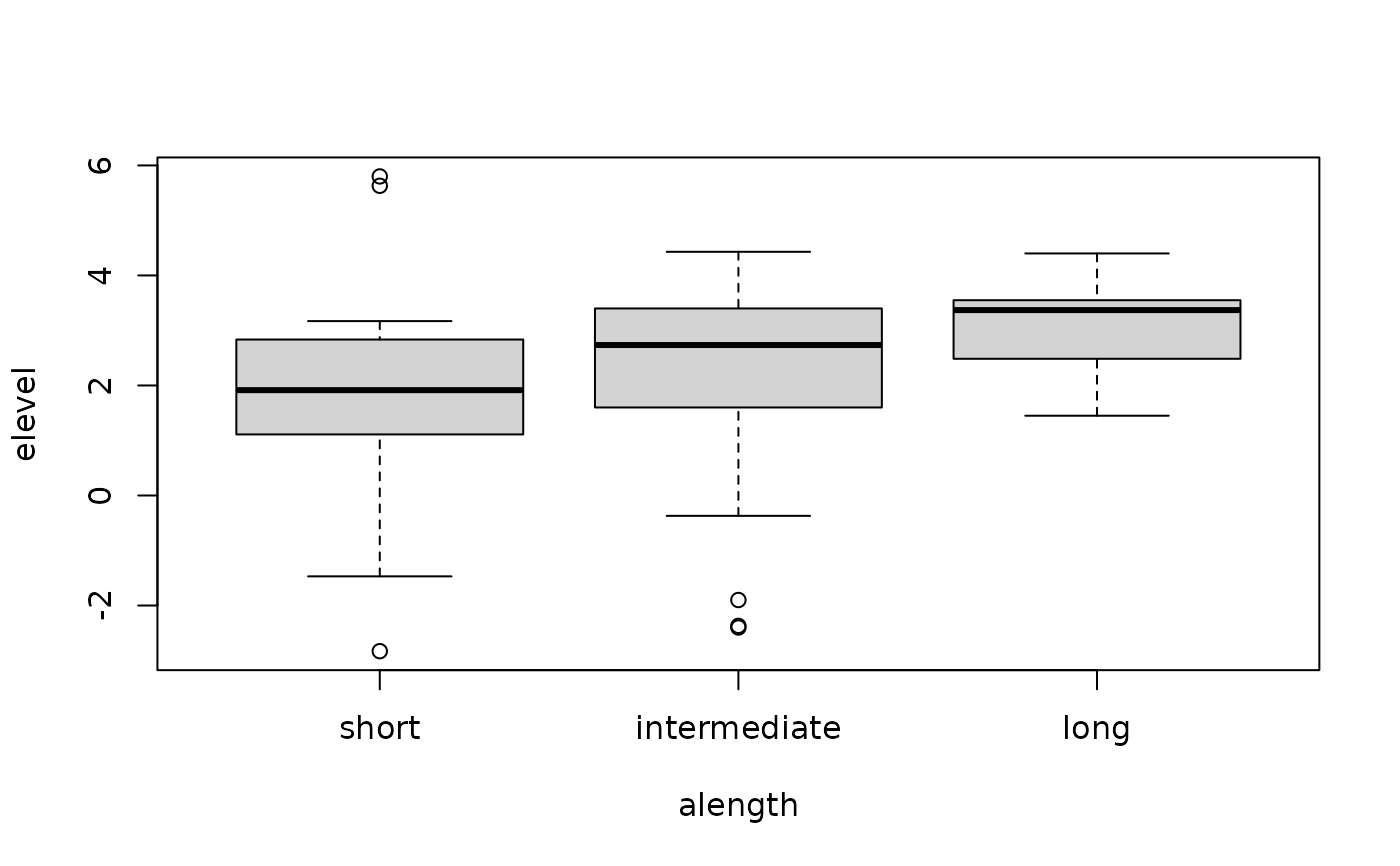 ## Asymptotic Kruskal-Wallis test
kruskal_test(elevel ~ alength, data = alpha)
#>
#> Asymptotic Kruskal-Wallis Test
#>
#> data: elevel by alength (short, intermediate, long)
#> chi-squared = 8.8302, df = 2, p-value = 0.01209
#>
## Asymptotic Kruskal-Wallis test using midpoint scores
kruskal_test(elevel ~ alength, data = alpha,
scores = list(alength = c(2, 7, 11)))
#>
#> Asymptotic Linear-by-Linear Association Test
#>
#> data: elevel by alength (short < intermediate < long)
#> Z = 2.9263, p-value = 0.00343
#> alternative hypothesis: two.sided
#>
## Asymptotic score-independent test
## Winell and Lindbaeck (2018)
(it <- independence_test(elevel ~ alength, data = alpha,
ytrafo = function(data)
trafo(data, numeric_trafo = rank_trafo),
xtrafo = function(data)
trafo(data, factor_trafo = function(x)
zheng_trafo(as.ordered(x)))))
#>
#> Asymptotic General Independence Test
#>
#> data: elevel by alength (short, intermediate, long)
#> maxT = 2.9651, p-value = 0.008198
#> alternative hypothesis: two.sided
#>
## Extract the "best" set of scores
ss <- statistic(it, type = "standardized")
idx <- which(abs(ss) == max(abs(ss)), arr.ind = TRUE)
ss[idx[1], idx[2], drop = FALSE]
#>
#> gamma = (0.0, 0.4, 1.0) 2.96508
## Asymptotic Kruskal-Wallis test
kruskal_test(elevel ~ alength, data = alpha)
#>
#> Asymptotic Kruskal-Wallis Test
#>
#> data: elevel by alength (short, intermediate, long)
#> chi-squared = 8.8302, df = 2, p-value = 0.01209
#>
## Asymptotic Kruskal-Wallis test using midpoint scores
kruskal_test(elevel ~ alength, data = alpha,
scores = list(alength = c(2, 7, 11)))
#>
#> Asymptotic Linear-by-Linear Association Test
#>
#> data: elevel by alength (short < intermediate < long)
#> Z = 2.9263, p-value = 0.00343
#> alternative hypothesis: two.sided
#>
## Asymptotic score-independent test
## Winell and Lindbaeck (2018)
(it <- independence_test(elevel ~ alength, data = alpha,
ytrafo = function(data)
trafo(data, numeric_trafo = rank_trafo),
xtrafo = function(data)
trafo(data, factor_trafo = function(x)
zheng_trafo(as.ordered(x)))))
#>
#> Asymptotic General Independence Test
#>
#> data: elevel by alength (short, intermediate, long)
#> maxT = 2.9651, p-value = 0.008198
#> alternative hypothesis: two.sided
#>
## Extract the "best" set of scores
ss <- statistic(it, type = "standardized")
idx <- which(abs(ss) == max(abs(ss)), arr.ind = TRUE)
ss[idx[1], idx[2], drop = FALSE]
#>
#> gamma = (0.0, 0.4, 1.0) 2.96508