Marginal Model Plotting
marginalModelPlot.RdFor a regression object, draw a plot of the response on the vertical axis versus
a linear combination \(u\) of regressors in the mean function on the horizontal
axis. Added to the plot are a smooth for the graph, along with
a smooth from the plot of the fitted values on \(u\). mmps is an alias
for marginalModelPlots, and mmp is an alias for marginalModelPlot.
Usage
marginalModelPlots(...)
mmps(model, terms= ~ ., fitted=TRUE, layout=NULL, ask,
main, groups, key=TRUE, ...)
marginalModelPlot(...)
mmp(model, ...)
# S3 method for class 'lm'
mmp(model, variable, sd = FALSE,
xlab = deparse(substitute(variable)),
smooth=TRUE, key=TRUE, pch, groups=NULL, ...)
# Default S3 method
mmp(model, variable, sd = FALSE,
xlab = deparse(substitute(variable)), ylab, smooth=TRUE,
key=TRUE, pch, groups=NULL,
col.line = carPalette()[c(2, 8)], col=carPalette()[1],
id=FALSE, grid=TRUE, ...)
# S3 method for class 'glm'
mmp(model, variable, sd = FALSE,
xlab = deparse(substitute(variable)), ylab,
smooth=TRUE, key=TRUE, pch, groups=NULL,
col.line = carPalette()[c(2, 8)], col=carPalette()[1],
id=FALSE, grid=TRUE, ...)Arguments
- model
A regression object, usually of class either
lmorglm, for which there is apredictmethod defined.- terms
A one-sided formula. A marginal model plot will be drawn for each term on the right-side of this formula that is not a factor. The default is
~ ., which specifies that all the terms informula(object)will be used. If a conditioning argument is given, egterms = ~. | a, then separate colors and smoothers are used for each unique non-missing value ofa. See examples below.- fitted
If
TRUE, the default, then a marginal model plot in the direction of the fitted values for a linear model or the linear predictor of a generalized linear model will be drawn.- layout
If set to a value like
c(1, 1)orc(4, 3), the layout of the graph will have this many rows and columns. If not set, the program will select an appropriate layout. If the number of graphs exceed nine, you must select the layout yourself, or you will get a maximum of nine per page. Iflayout=NA, the function does not set the layout and the user can use theparfunction to control the layout, for example to have plots from two models in the same graphics window.- ask
If
TRUE, ask before clearing the graph window to draw more plots.- main
Main title for the array of plots. Use
main=""to suppress the title; if missing, a title will be supplied.- ...
Additional arguments passed from
mmpstommpand then toplot. Users should generally usemmps, or equivalentlymarginalModelPlots.- variable
The quantity to be plotted on the horizontal axis. If this argument is missing, the horizontal variable is the linear predictor, returned by
predict(object)for models of classlm, with default label"Fitted values", or returned bypredict(object, type="link")for models of classglm, with default label"Linear predictor". It can be any other vector of length equal to the number of observations in the object. Thus themmpfunction can be used to get a marginal model plot versus any regressor or predictor while themmpsfunction can be used only to get marginal model plots for the first-order regressors in the formula. In particular, terms defined by a spline basis are skipped bymmps, but you can usemmpto get the plot for the variable used to define the splines.- sd
If
TRUE, display sd smooths. For a binomial regression with all sample sizes equal to one, this argument is ignored as the SD bounds don't make any sense.- xlab
label for horizontal axis.
- ylab
label for vertical axis, defaults to name of response.
- smooth
specifies the smoother to be used along with its arguments; if
FALSE, no smoother is shown; can be a list giving the smoother function and its named arguments;TRUE, the default, is equivalent tolist(smoother=loessLine, span=2/3)for linear models andlist(smoother=gamLine, k=3)for generalized linear models. SeeScatterplotSmoothersfor the smoothers supplied by the car package and their arguments; thespreadargument is not supported for marginal model plots.- groups
The name of a vector that specifies a grouping variable for separate colors/smoothers. This can also be specified as a conditioning argument on the
termsargument.- key
If
TRUE, include a key at the top of the plot, ifFALSEomit the key. If grouping is present, the key is only printed for the upper-left plot.- id
controls point identification; if
FALSE(the default), no points are identified; can be a list of named arguments to theshowLabelsfunction;TRUEis equivalent tolist(method="y", n=2, cex=1, col=carPalette()[1], location="lr"), which identifies the 2 points with the most unusual response (Y) values.- pch
plotting character to use if no grouping is present.
- col.line
colors for data and model smooth, respectively. The default is to use
carPalette,carPalette()[c(2, 8)], blue and red.- col
color(s) for the plotted points.
- grid
If TRUE, the default, a light-gray background grid is put on the graph
Details
mmp and marginalModelPlot draw one marginal model plot against
whatever is specified as the horizontal axis.
mmps and marginalModelPlots draws marginal model plots
versus each of the terms in the terms argument and versus fitted values.
mmps skips factors and interactions if they are specified in the
terms argument. Terms based on polynomials or on splines (or
potentially any term that is represented by a matrix of regressors) will
be used to form a marginal model plot by returning a linear combination of the
terms. For example, if you specify terms = ~ X1 + poly(X2, 3) and
poly(X2, 3) was part of the original model formula, the horizontal
axis of the marginal model plot for X2 will be the value of
predict(model, type="terms")[, "poly(X2, 3)"]). If the predict
method for the model you are using doesn't support type="terms",
then the polynomial/spline term is skipped. Adding a conditioning variable,
e.g., terms = ~ a + b | c, will produce marginal model plots for a
and b with different colors and smoothers for each unique non-missing
value of c.
For linear models, the default smoother is loess.
For generalized linear models, the default smoother uses gamLine, fitting
a generalized additive model with the same family, link and weights as the fit of the
model. SD smooths are not computed for for generalized linear models.
For generalized linear models the default number of elements in the spline basis is
k=3; this is done to allow fitting for predictors with just a few support
points. If you have many support points you may wish to set k to a higher
number, or k=-1 for the default used by gam.
References
Cook, R. D., & Weisberg, S. (1997). Graphics for assessing the adequacy of regression models. Journal of the American Statistical Association, 92(438), 490-499.
Fox, J. and Weisberg, S. (2019) An R Companion to Applied Regression, Third Edition. Sage.
Weisberg, S. (2005) Applied Linear Regression, Third Edition, Wiley, Section 8.4.
Author
Sanford Weisberg, sandy@umn.edu
Examples
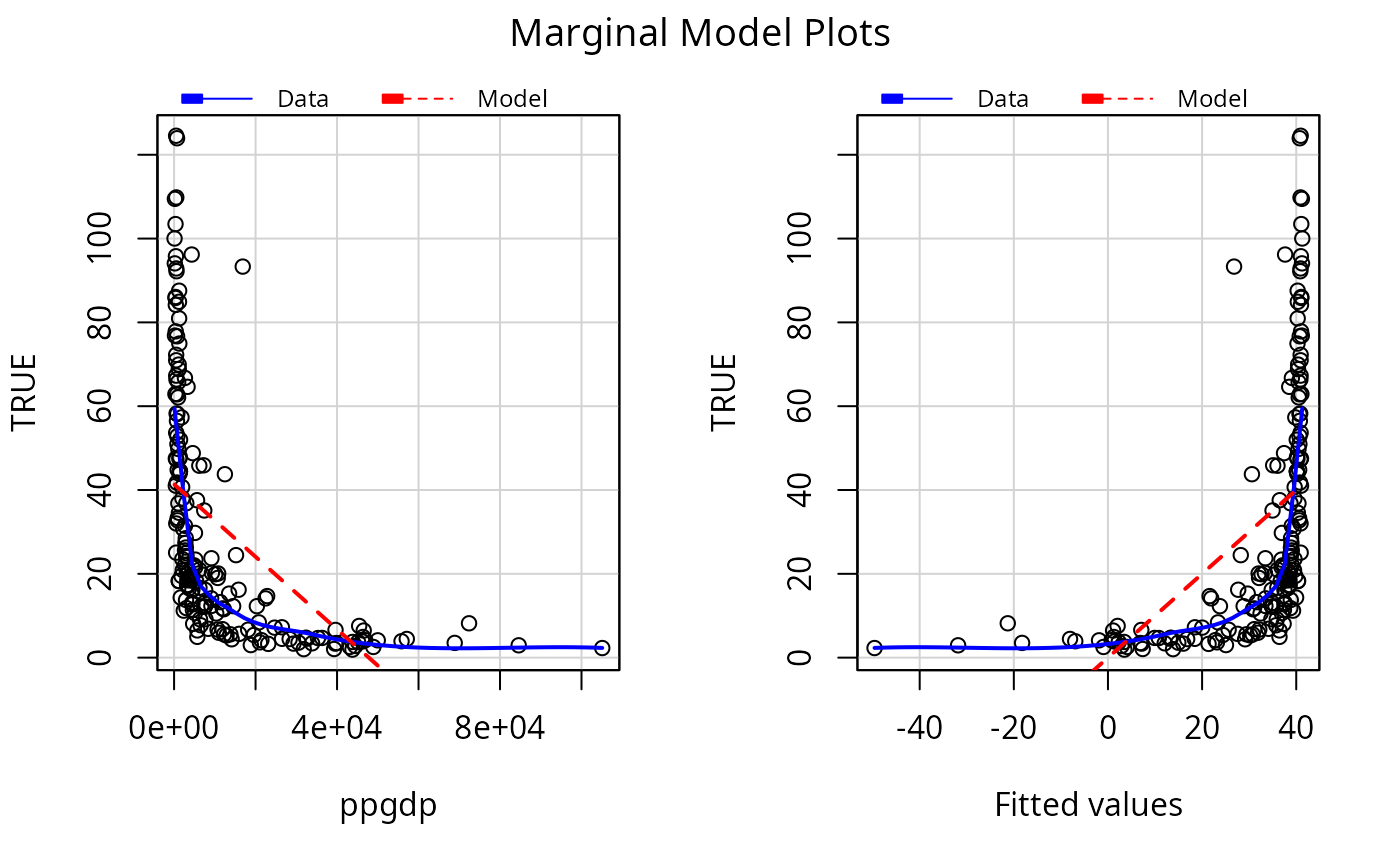
c1 <- lm(infantMortality ~ ppgdp, UN)
mmps(c1)
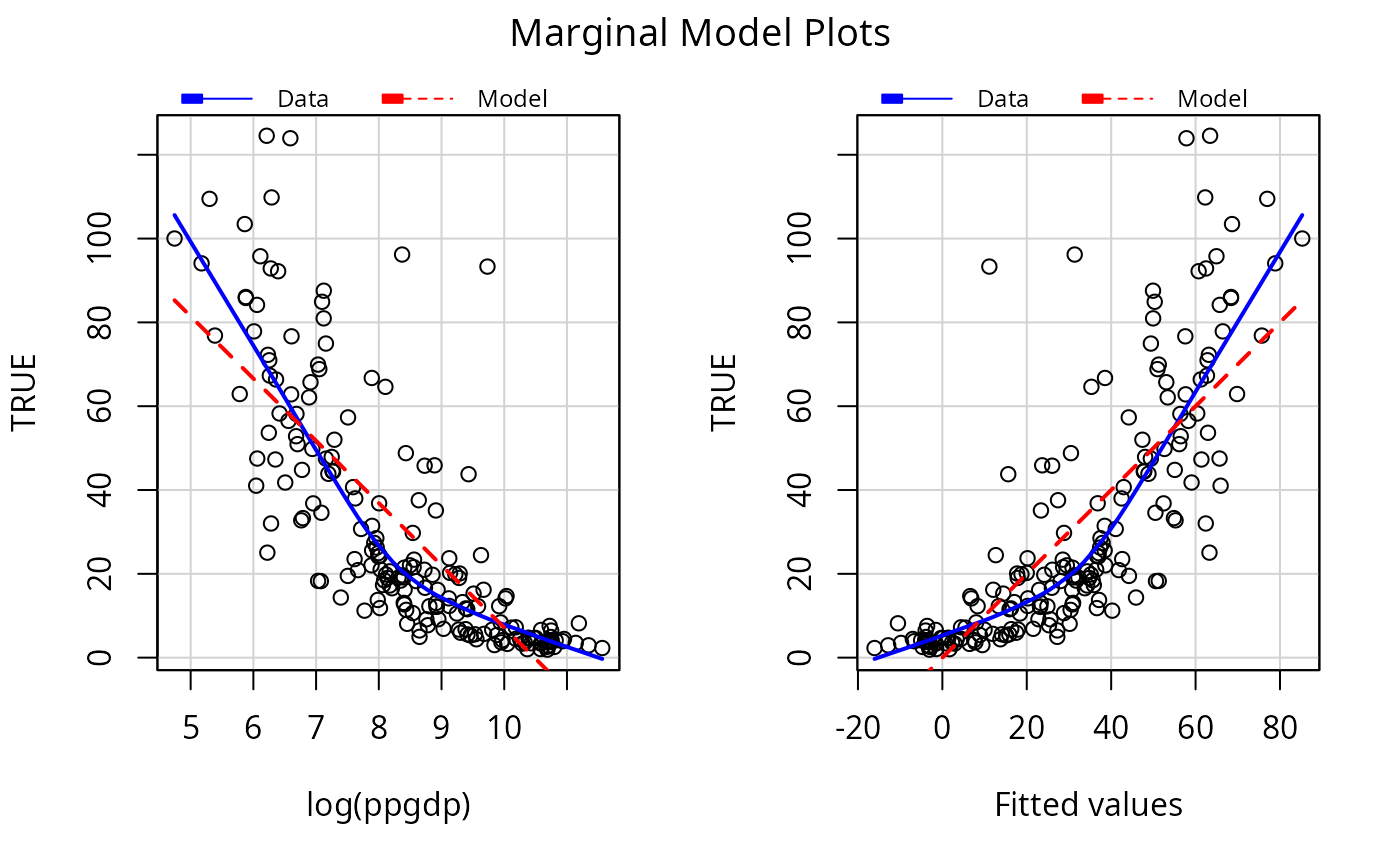
 c2 <- update(c1, ~ log(ppgdp))
mmps(c2)
c2 <- update(c1, ~ log(ppgdp))
mmps(c2)
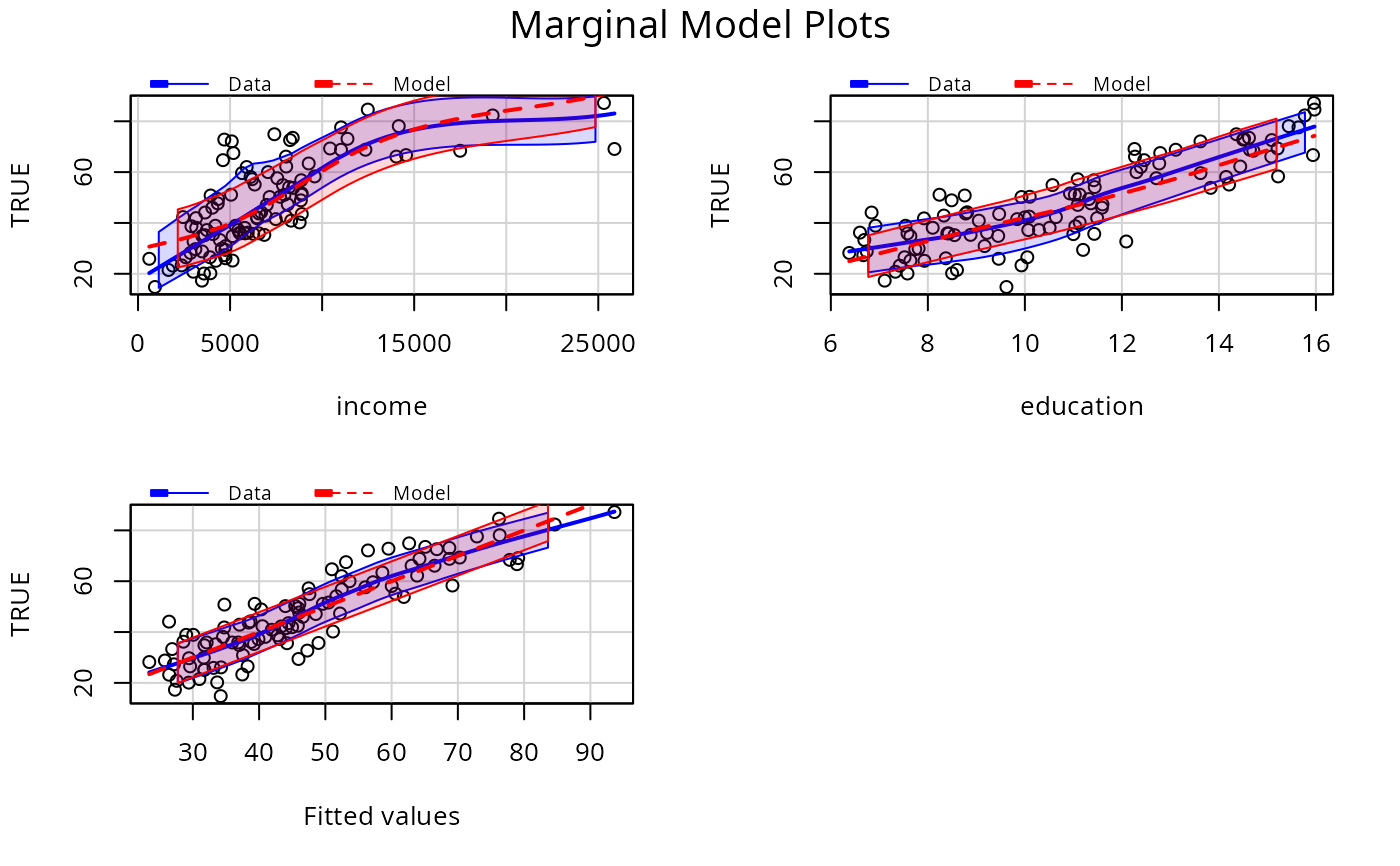
 # include SD lines
p1 <- lm(prestige ~ income + education, Prestige)
mmps(p1, sd=TRUE)
# include SD lines
p1 <- lm(prestige ~ income + education, Prestige)
mmps(p1, sd=TRUE)
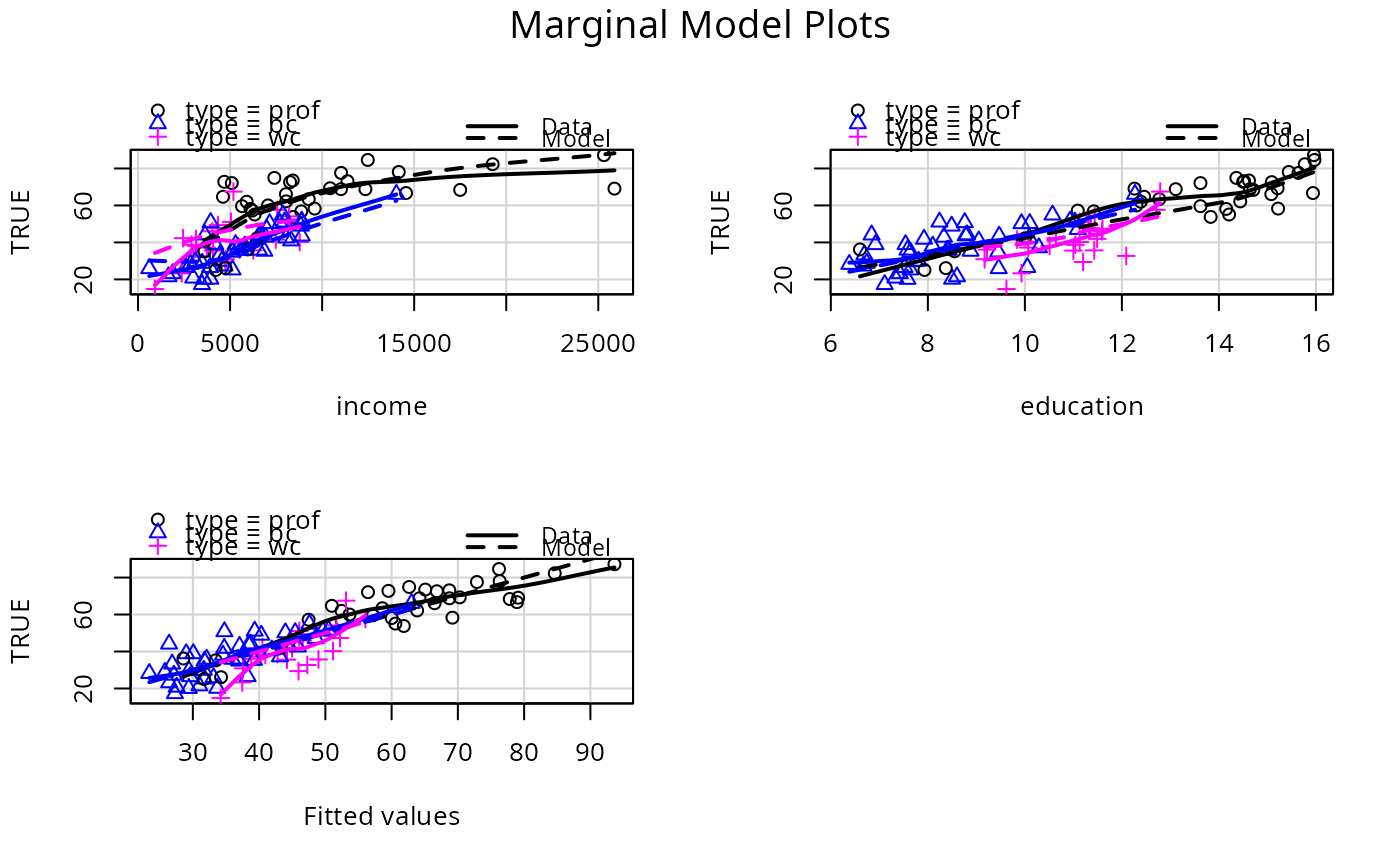
 # condition on type:
mmps(p1, ~. | type)
# condition on type:
mmps(p1, ~. | type)
 # logisitic regression example
# smoothers return warning messages.
# fit a separate smoother and color for each type of occupation.
m1 <- glm(lfp ~ ., family=binomial, data=Mroz)
mmps(m1)
# logisitic regression example
# smoothers return warning messages.
# fit a separate smoother and color for each type of occupation.
m1 <- glm(lfp ~ ., family=binomial, data=Mroz)
mmps(m1)
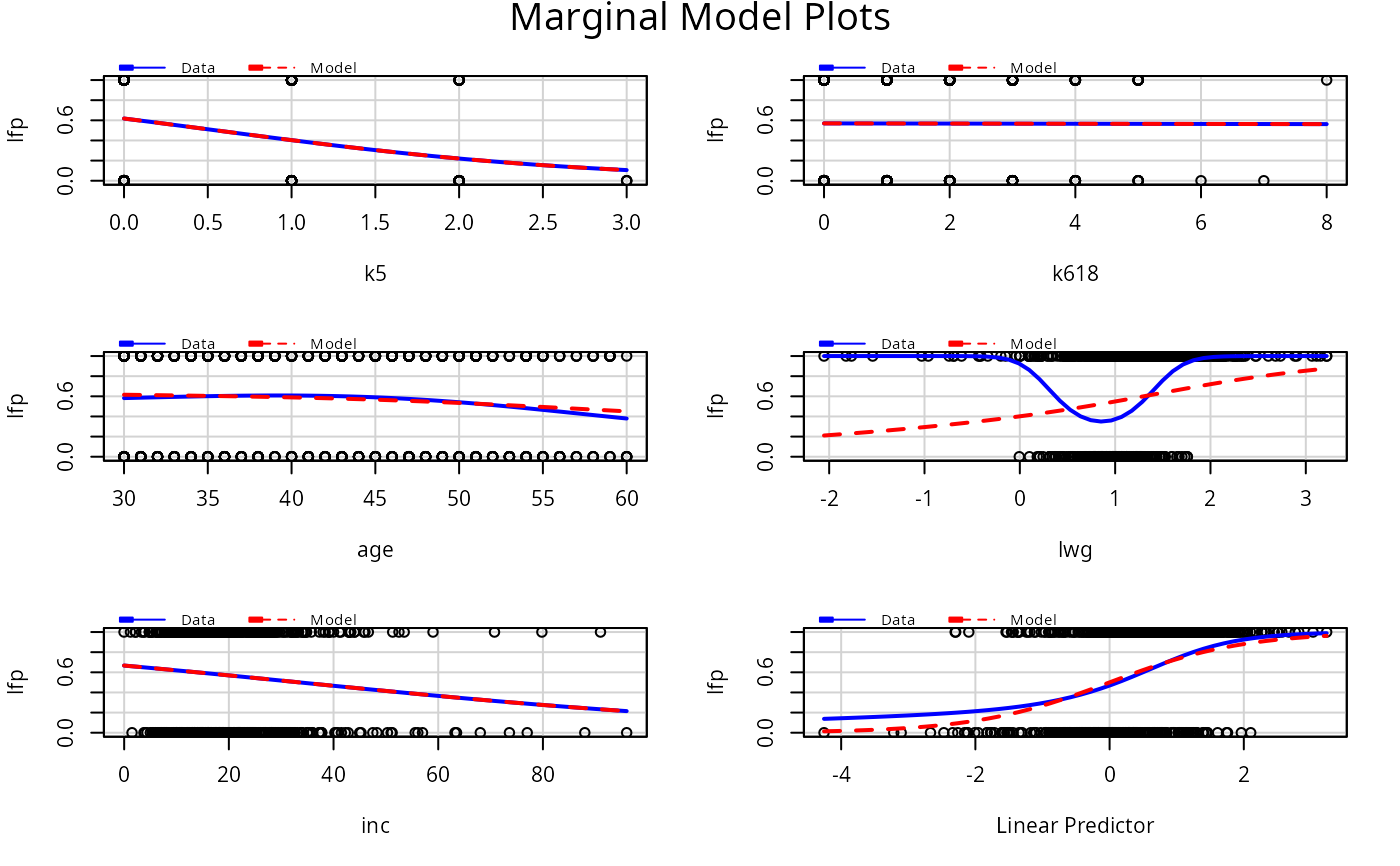 #> Warning: Interactions and/or factors skipped
#> Warning: Interactions and/or factors skipped