Tidying methods for MCMC (Stan, JAGS, etc.) fits
Source:R/MCMCglmm_tidiers.R, R/mcmc_tidiers.R
mcmc_tidiers.RdTidying methods for MCMC (Stan, JAGS, etc.) fits
Usage
# S3 method for class 'MCMCglmm'
tidy(x, effects = c("fixed", "ran_pars"), scales = NULL, ...)
tidyMCMC(
x,
pars,
robust = FALSE,
conf.int = FALSE,
conf.level = 0.95,
conf.method = c("quantile", "HPDinterval"),
drop.pars = c("lp__", "deviance"),
rhat = FALSE,
ess = FALSE,
index = FALSE,
...
)
# S3 method for class 'rjags'
tidy(
x,
robust = FALSE,
conf.int = FALSE,
conf.level = 0.95,
conf.method = "quantile",
...
)
# S3 method for class 'stanfit'
tidy(
x,
pars,
robust = FALSE,
conf.int = FALSE,
conf.level = 0.95,
conf.method = c("quantile", "HPDinterval"),
drop.pars = c("lp__", "deviance"),
rhat = FALSE,
ess = FALSE,
index = FALSE,
...
)
# S3 method for class 'mcmc'
tidy(
x,
pars,
robust = FALSE,
conf.int = FALSE,
conf.level = 0.95,
conf.method = c("quantile", "HPDinterval"),
drop.pars = c("lp__", "deviance"),
rhat = FALSE,
ess = FALSE,
index = FALSE,
...
)
# S3 method for class 'mcmc.list'
tidy(
x,
pars,
robust = FALSE,
conf.int = FALSE,
conf.level = 0.95,
conf.method = c("quantile", "HPDinterval"),
drop.pars = c("lp__", "deviance"),
rhat = FALSE,
ess = FALSE,
index = FALSE,
...
)Arguments
- x
a model fit to be converted to a data frame
- effects
which effects (fixed, random, etc.) to return
- scales
scales on which to report results
- ...
mostly unused; for
tidy.MCMCglmm, these represent options passed through totidy.mcmc(e.g.robust,conf.int,conf.method, ...)- pars
(character) specification of which parameters to include
- robust
use mean and standard deviation (if FALSE) or median and mean absolute deviation (if TRUE) to compute point estimates and uncertainty?
- conf.int
(logical) include confidence interval?
- conf.level
probability level for CI
- conf.method
method for computing confidence intervals ("quantile" or "HPDinterval")
- drop.pars
Parameters not to include in the output (such as log-probability information)
- rhat, ess
(logical) include Rhat and/or effective sample size estimates?
- index
Add index column, remove index from term. For example,
term a[13]becomesterm aandindex 13.
Examples
if (require("MCMCglmm")) {
## original model
if (FALSE) { # \dontrun{
mm0 <- MCMCglmm(Reaction ~ Days,
random = ~Subject, data = sleepstudy,
nitt=4000,
pr = TRUE
)
} # }
## load stored object
load(system.file("extdata","MCMCglmm_example.rda",
package="broom.mixed"))
tidy(mm0)
tidy(mm1)
tidy(mm2)
tail(tidy(mm0,effects="ran_vals"))
}
#> Loading required package: MCMCglmm
#> Loading required package: coda
#>
#> Attaching package: ‘coda’
#> The following object is masked from ‘package:rstan’:
#>
#> traceplot
#> Loading required package: ape
#>
#> Attaching package: ‘ape’
#> The following object is masked from ‘package:dplyr’:
#>
#> where
#>
#> Attaching package: ‘MCMCglmm’
#> The following object is masked from ‘package:brms’:
#>
#> me
#> # A tibble: 6 × 6
#> effect group level term estimate std.error
#> <chr> <chr> <chr> <chr> <dbl> <dbl>
#> 1 ran_vals Subject 351 (Intercept) -8.10 12.9
#> 2 ran_vals Subject 352 (Intercept) 36.1 14.6
#> 3 ran_vals Subject 369 (Intercept) 7.11 13.4
#> 4 ran_vals Subject 370 (Intercept) -6.54 12.2
#> 5 ran_vals Subject 371 (Intercept) -4.52 14.1
#> 6 ran_vals Subject 372 (Intercept) 17.6 14.3
# Using example from "RStan Getting Started"
# https://github.com/stan-dev/rstan/wiki/RStan-Getting-Started
model_file <- system.file("extdata", "8schools.stan", package = "broom.mixed")
schools_dat <- list(J = 8,
y = c(28, 8, -3, 7, -1, 1, 18, 12),
sigma = c(15, 10, 16, 11, 9, 11, 10, 18))
## original model
if (FALSE) { # \dontrun{
set.seed(2015)
rstan_example <- rstan::stan(file = model_file, data = schools_dat,
iter = 1000, chains = 2, save_dso = FALSE)
} # }
if (require(rstan)) {
## load stored object
rstan_example <- readRDS(system.file("extdata", "rstan_example.rds", package = "broom.mixed"))
tidy(rstan_example)
tidy(rstan_example, conf.int = TRUE, pars = "theta")
td_mean <- tidy(rstan_example, conf.int = TRUE)
td_median <- tidy(rstan_example, conf.int = TRUE, robust = TRUE)
if (require(dplyr) && require(ggplot2)) {
tds <- (dplyr::bind_rows(list(mean=td_mean, median=td_median), .id="method")
%>% mutate(type=ifelse(grepl("^theta",term),"theta",
ifelse(grepl("^eta",term),"eta",
"other")))
)
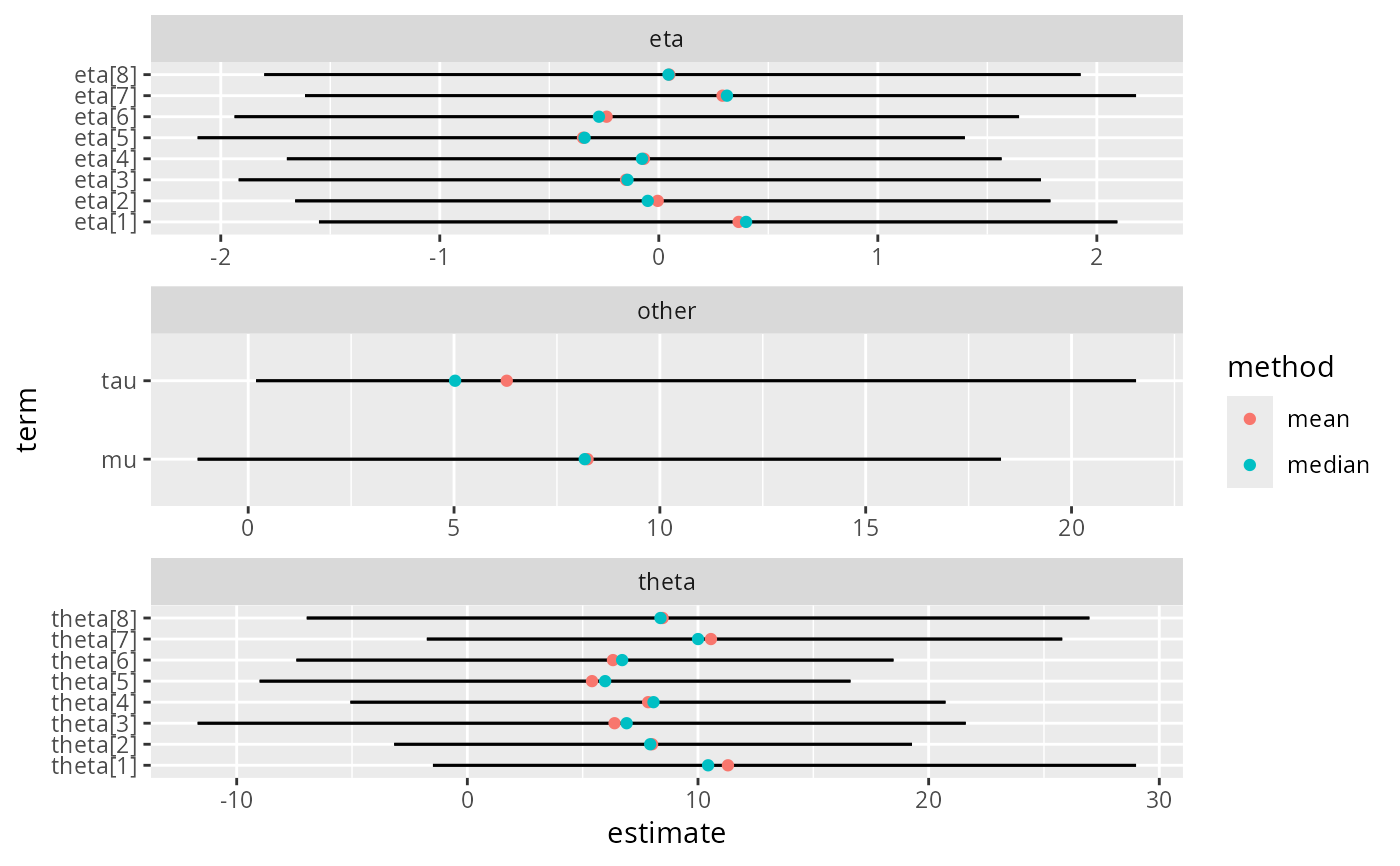
ggplot(tds, aes(estimate, term)) +
geom_errorbarh(aes(xmin = conf.low, xmax = conf.high),height=0) +
geom_point(aes(color = method))+
facet_wrap(~type,scale="free",ncol=1)
} ## require(dplyr,ggplot2)
} ## require(rstan)
#> `height` was translated to `width`.
 if (require(R2jags)) {
## see help("jags",package="R2jags")
## and example("jags",package="R2jags")
## for details
## load stored object
R2jags_example <- readRDS(system.file("extdata", "R2jags_example.rds", package = "broom.mixed"))
tidy(R2jags_example)
tidy(R2jags_example, conf.int=TRUE, conf.method="quantile")
}
#> Loading required package: R2jags
#> Loading required package: rjags
#> Linked to JAGS 4.3.2
#> Loaded modules: basemod,bugs
#>
#> Attaching package: ‘R2jags’
#> The following object is masked from ‘package:coda’:
#>
#> traceplot
#> The following object is masked from ‘package:rstan’:
#>
#> traceplot
#> # A tibble: 10 × 5
#> term estimate std.error conf.low conf.high
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 mu -0.321 1.37 -1.70 1.85
#> 2 sigma 0.710 0.535 0.0360 1.44
#> 3 theta[1] -0.00383 1.62 -1.68 2.92
#> 4 theta[2] -0.0727 1.65 -2.09 2.70
#> 5 theta[3] -0.613 1.17 -1.76 1.86
#> 6 theta[4] -0.719 1.24 -1.78 1.71
#> 7 theta[5] -0.439 1.38 -1.71 2.18
#> 8 theta[6] -0.0943 1.70 -1.71 2.84
#> 9 theta[7] -0.312 1.57 -1.72 2.41
#> 10 theta[8] -0.556 1.34 -2.46 1.58
if (require(R2jags)) {
## see help("jags",package="R2jags")
## and example("jags",package="R2jags")
## for details
## load stored object
R2jags_example <- readRDS(system.file("extdata", "R2jags_example.rds", package = "broom.mixed"))
tidy(R2jags_example)
tidy(R2jags_example, conf.int=TRUE, conf.method="quantile")
}
#> Loading required package: R2jags
#> Loading required package: rjags
#> Linked to JAGS 4.3.2
#> Loaded modules: basemod,bugs
#>
#> Attaching package: ‘R2jags’
#> The following object is masked from ‘package:coda’:
#>
#> traceplot
#> The following object is masked from ‘package:rstan’:
#>
#> traceplot
#> # A tibble: 10 × 5
#> term estimate std.error conf.low conf.high
#> <chr> <dbl> <dbl> <dbl> <dbl>
#> 1 mu -0.321 1.37 -1.70 1.85
#> 2 sigma 0.710 0.535 0.0360 1.44
#> 3 theta[1] -0.00383 1.62 -1.68 2.92
#> 4 theta[2] -0.0727 1.65 -2.09 2.70
#> 5 theta[3] -0.613 1.17 -1.76 1.86
#> 6 theta[4] -0.719 1.24 -1.78 1.71
#> 7 theta[5] -0.439 1.38 -1.71 2.18
#> 8 theta[6] -0.0943 1.70 -1.71 2.84
#> 9 theta[7] -0.312 1.57 -1.72 2.41
#> 10 theta[8] -0.556 1.34 -2.46 1.58