Introduction to `broom.mixed`
Ben Bolker
2026-03-05
Source:vignettes/broom_mixed_intro.rmd
broom_mixed_intro.rmdIntroduction
broom.mixed is a spinoff of the broom package. The
goal of broom is to bring the modeling process into a
“tidy”(TM) workflow, in particular by providing standardized verbs that
provide information on
-
tidy: estimates, standard errors, confidence intervals, etc. -
augment: residuals, fitted values, influence measures, etc. -
glance: whole-model summaries: AIC, R-squared, etc.
broom.mixed aims to provide these methods for as many
packages and model types in the R ecosystem as possible. These methods
have been separated from those in the main broom package
because there are issues that need to be dealt with for these models
(e.g. different types of parameters: fixed, random-effects parameters,
conditional modes/BLUPs of random effects, etc.) that are not especially
relevant to the broader universe of models that broom deals
with.
Mixed-model-specific issues
Terminology
- the upper-level parameters that describe the distribution of random
variables (variance, covariance, precision, standard deviation, or
correlation) are called random-effect parameters
(
ran_parsin theeffectsargument when tidying) - the values that describe the deviation of the observations in a
group level from the population-level effect (which could be posterior
means or medians, conditional modes, or BLUPs depending on the model)
are called random-effect values (
ran_valsin theeffectsargument when tidying) - the parameters that describe the population-level effects of
(categorical and continuous) predictor variables are called fixed
effects (
fixedin theeffectsargument when tidying) - the categorical variable (factor) that identifies which group or
cluster an observation belongs to is called a grouping variable
(
groupcolumn intidy()output) - the particular level of a factor that specifies which level of the
grouping variable an observation belongs to is called a group
level (
levelcolumn intidy()output) - the categorical or continuous predictor variables that control the
expected value (i.e., enter into the linear predictor for some part of
the model) are called terms (
termcolumn intidy()output); note that unlike in basebroom, the term column may have duplicated values, because the same term may enter multiple model components (e.g. zero-inflated and conditional models; models for more than one response; fixed effects and random effects)
Time-consuming computations
Some kinds of computations needed for mixed model summaries are
computationally expensive, e.g. likelihood profiles or parametric
bootstrapping. In this case broom.mixed may offer an option
for passing a pre-computed object to tidy(), eg. the
profile argument in the tidy.merMod
(lmer/glmer) method.
Related packages
There are many, many things one might want to do with a fitted model,
and broom.mixed can only do a few of them.
emmeansmultcompcarafex-
sjStats/sjPlots rockchalk
dotwhisker + broom.mixed
dotwhisker is a convenient platform for creating
dot-whisker plots - either directly from models or lists of models
(tidy() methods are automatically called to convert the
models to a tidy format), or from the (possibly post-processed) output
of a tidy() call. There are a couple of caveats and issues
to be aware of when using dotwhisker in conjunction with
broom.mixed, however.
- For fixed effects, the
groupvalue is set toNA: in the current CRAN version (0.5.0), an unfortunatena.omit()within thedwplotcode will eliminate all of the fixed effects unless you drop this column before passing the results todwplot(this has been fixed in the current GitHub version, which you can install withdevtools::install_github("fsolt/dotwhisker")). - In
broom.mixedoutput, it is fairly common for a single tidied model to have duplicated entries in thetermcolumn (e.g. effects that appear in both the conditional and the zero-inflated model, or intercept standard deviations for several different grouping variables).dotwhisker::dwplottakes this as evidence that it has been handed a tidied object containing the results from several different models, and asks for amodelcolumn that will distinguish the non-unique terms. There are (at least) two strategies you can take:-
dwplot(list(fitted_model))will plot all of the non-unique values together -
tidy(fitted_model) %>% tidyr::unite(term, group, term)will create a newtermcolumn that’s the combination of thegroupandtermcolumns (which will disambiguate random-effect terms from different grouping variables);unite(term, component, term)will disambiguate conditional and zero-inflation parameters. The code below shows a slightly more complicated (but prettier) approach. (Some sort ofdisambiguate_terms()function could be added in a future version of the package …)
-
library(dplyr)
library(tidyr)
require(rstan) ## workaround for r-devel problem
library(broom.mixed)
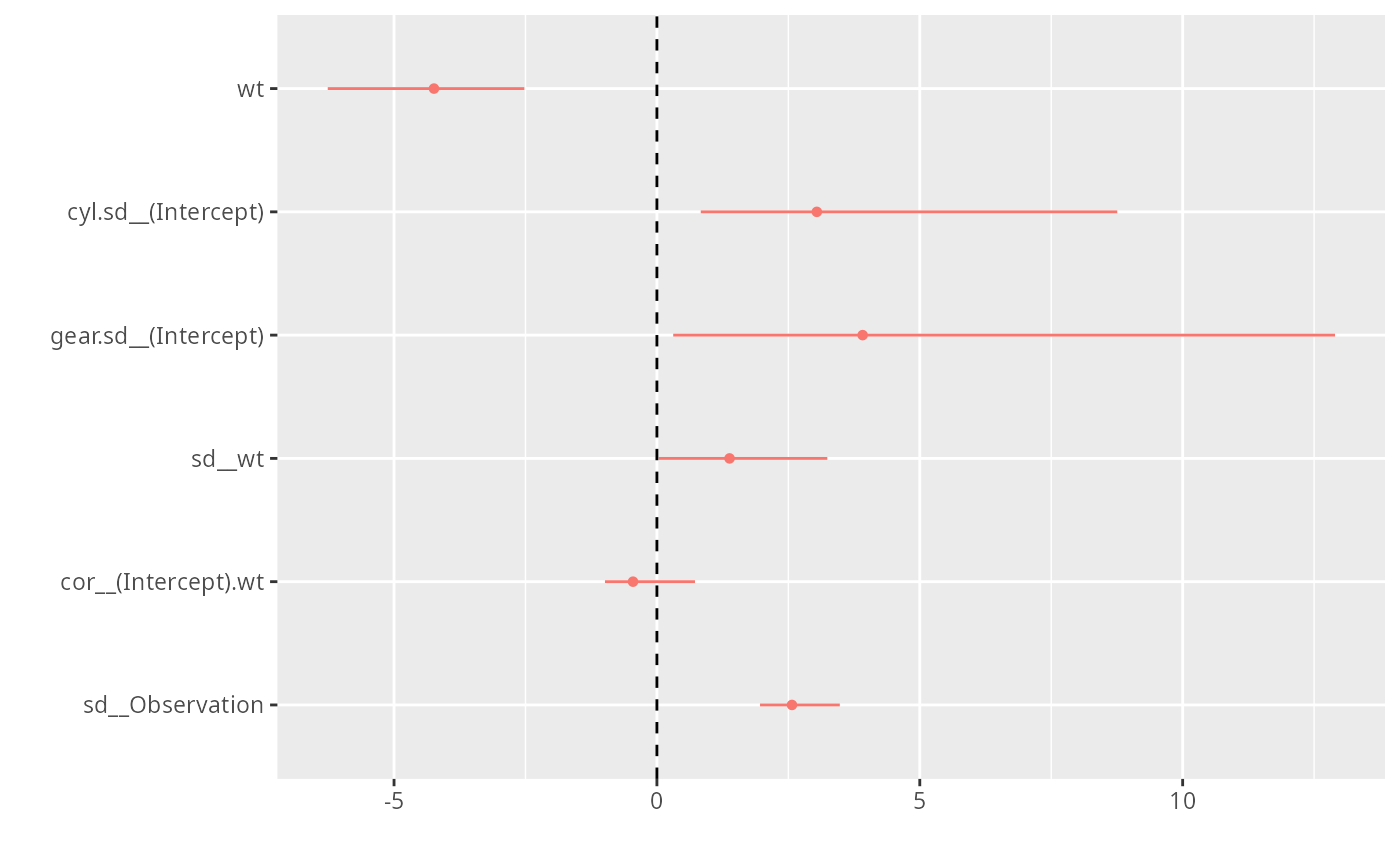
if (require("brms") && require("dotwhisker") && require("ggplot2")) {
L <- load(system.file("extdata", "brms_example.rda", package="broom.mixed"))
gg0 <- (tidy(brms_crossedRE)
## disambiguate
%>% mutate(term=ifelse(grepl("sd__(Int",term,fixed=TRUE),
paste(group,term,sep="."),
term))
%>% dwplot
)
gg0 + geom_vline(xintercept=0,lty=2)
}
Capabilities
Automatically retrieve table of available methods:
## # A tibble: 27 × 4
## class tidy glance augment
## <chr> <lgl> <lgl> <lgl>
## 1 allFit TRUE TRUE FALSE
## 2 brmsfit TRUE TRUE TRUE
## 3 clmm FALSE FALSE TRUE
## 4 gamlss TRUE TRUE FALSE
## 5 gamm4 TRUE TRUE TRUE
## 6 glmm TRUE FALSE FALSE
## 7 glmmadmb TRUE TRUE TRUE
## 8 glmmTMB TRUE TRUE TRUE
## 9 gls TRUE TRUE TRUE
## 10 lme TRUE TRUE TRUE
## # ℹ 17 more rowsManually compiled list of capabilities (possibly out of date):
| package | object | tidy | glance | augment | effects.fixed | effects.ran_vals | effects.ran_pars | effects.ran_coefs | confint…Ww.ald. | confint..profile. | confint..boot. | component.zi | component.disp | covstruct |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| lme4 | glmer | y | y | y | y | y | y | y | NA | y | NA | NA | NA | ? |
| lme4 | lmer | y | y | y | y | y | y | y | NA | y | NA | NA | NA | ? |
| nlme | lme | y | y | y | y | y | y | y | NA | n | NA | NA | ? | ? |
| nlme | gls | y | y | y | y | NA | NA | NA | NA | n | NA | NA | ? | ? |
| nlme | nlme | y | y | y | y | y | n | y | NA | n | NA | NA | ? | ? |
| glmmTMB | glmmTMB | y | y | y | y | y | y | n | NA | NA | y | y | ? | |
| glmmADMB | glmmadmb | y | y | y | y | y | y | n | NA | NA | y | ? | ? | |
| brms | brmsfit | y | y | y | y | y | y | n | NA | NA | y | ? | ? | |
| rstanarm | stanreg | y | y | y | y | y | y | n | NA | NA | NA | NA | ? | |
| MCMCglmm | MCMCglmm | y | y | y | y | y | y | n | NA | NA | ? | ? | ? | |
| TMB | TMB | y | n | n | n | n | n | n | NA | NA | NA | NA | ? | |
| INLA | n | n | n | n | n | n | n | NA | NA | ? | ? | ? |