Trajectory plot for QRR-VGLMs
trplot.qrrvglm.RdProduces a trajectory plot for
quadratic reduced-rank vector generalized linear models
(QRR-VGLMs).
It is only applicable for rank-1 models with argument
noRRR = ~ 1.
Usage
trplot.qrrvglm(object, which.species = NULL, add = FALSE,
show.plot = TRUE,
label.sites = FALSE, sitenames = rownames(object@y),
axes.equal = TRUE, cex = par()$cex,
col = 1:(nos * (nos - 1)/2), log = "",
lty = rep_len(par()$lty, nos * (nos - 1)/2),
lwd = rep_len(par()$lwd, nos * (nos - 1)/2),
tcol = rep_len(par()$col, nos * (nos - 1)/2),
xlab = NULL, ylab = NULL, main = "",
type = "b", asp = NULL, check.ok = TRUE, ...)Arguments
- object
Object of class
"qrrvglm", i.e., a CQO object.- which.species
Integer or character vector specifying the species to be plotted. If integer, these are the columns of the response matrix. If character, these must match exactly with the species' names. The default is to use all species.
- add
Logical. Add to an existing plot? If
FALSE(default), a new plot is made.- show.plot
Logical. Plot it?
- label.sites
Logical. If
TRUE, the points on the curves/trajectories are labelled with thesitenames.- sitenames
Character vector. The names of the sites.
- axes.equal
Logical. If
TRUE, the x- and y-axes will be on the same scale.- cex
Character expansion of the labelling of the site names. Used only if
label.sitesisTRUE. See thecexargument inpar.- col
Color of the lines. See the
colargument inpar. Here,nosis the number of species.- log
Character, specifying which (if any) of the x- and y-axes are to be on a logarithmic scale. See the
logargument inpar.- lty
Line type. See the
ltyargument ofpar.- lwd
Line width. See the
lwdargument ofpar.- tcol
Color of the text for the site names. See the
colargument inpar. Used only iflabel.sitesisTRUE.- asp
Aspect ratio. See the
aspargument inpar.- xlab
Character caption for the x-axis. By default, a suitable caption is found. See the
xlabargument inplotortitle.- ylab
Character caption for the y-axis. By default, a suitable caption is found. See the
xlabargument inplotortitle.- main
Character, giving the title of the plot. See the
mainargument inplotortitle.- type
Character, giving the type of plot. A common option is to use
type="l"for lines only. See thetypeargument ofplot.- check.ok
Logical. Whether a check is performed to see that
noRRR = ~ 1was used. It doesn't make sense to have a trace plot unless this is so.- ...
Arguments passed into the
plotfunction when setting up the entire plot. Useful arguments here includexlimandylim.
Details
A trajectory plot plots the fitted values of a `second' species
against a `first' species. The argument which.species must
therefore contain at least two species. By default, all of the
species that were fitted in object are plotted.
With more than a few species
the resulting plot will be very congested, and so it
is recommended
that only a few species be selected for plotting.
In the above, \(M\) is the number of species selected for plotting, so there will be \(M(M-1)/2\) curves/trajectories in total.
A trajectory plot will be fitted only
if noRRR = ~ 1 because
otherwise the trajectory will not be a smooth function
of the latent
variables.
Value
A list with the following components.
- species.names
A matrix of characters giving the `first' and `second' species. The number of different combinations of species is given by the number of rows. This is useful for creating a legend.
- sitenames
A character vector of site names, sorted by the latent variable (from low to high).
References
Yee, T. W. (2020). On constrained and unconstrained quadratic ordination. Manuscript in preparation.
Note
Plotting the axes on a log scale is often a good idea.
The use of xlim and ylim to control
the axis limits
is also a good idea, so as to limit the extent
of the curves at low
abundances or probabilities.
Setting label.sites = TRUE
is a good idea only if the number of
sites is small, otherwise there is too much clutter.
Examples
if (FALSE) set.seed(111) # Leads to the global solution
# hspider[,1:6] <- scale(hspider[,1:6]) # Stdze the environ vars
p1 <- cqo(cbind(Alopacce, Alopcune, Alopfabr, Arctlute,
Arctperi, Auloalbi, Pardlugu, Pardmont,
Pardnigr, Pardpull, Trocterr, Zoraspin) ~
WaterCon + BareSand + FallTwig + CoveMoss +
CoveHerb + ReflLux,
poissonff, data = hspider, trace = FALSE)
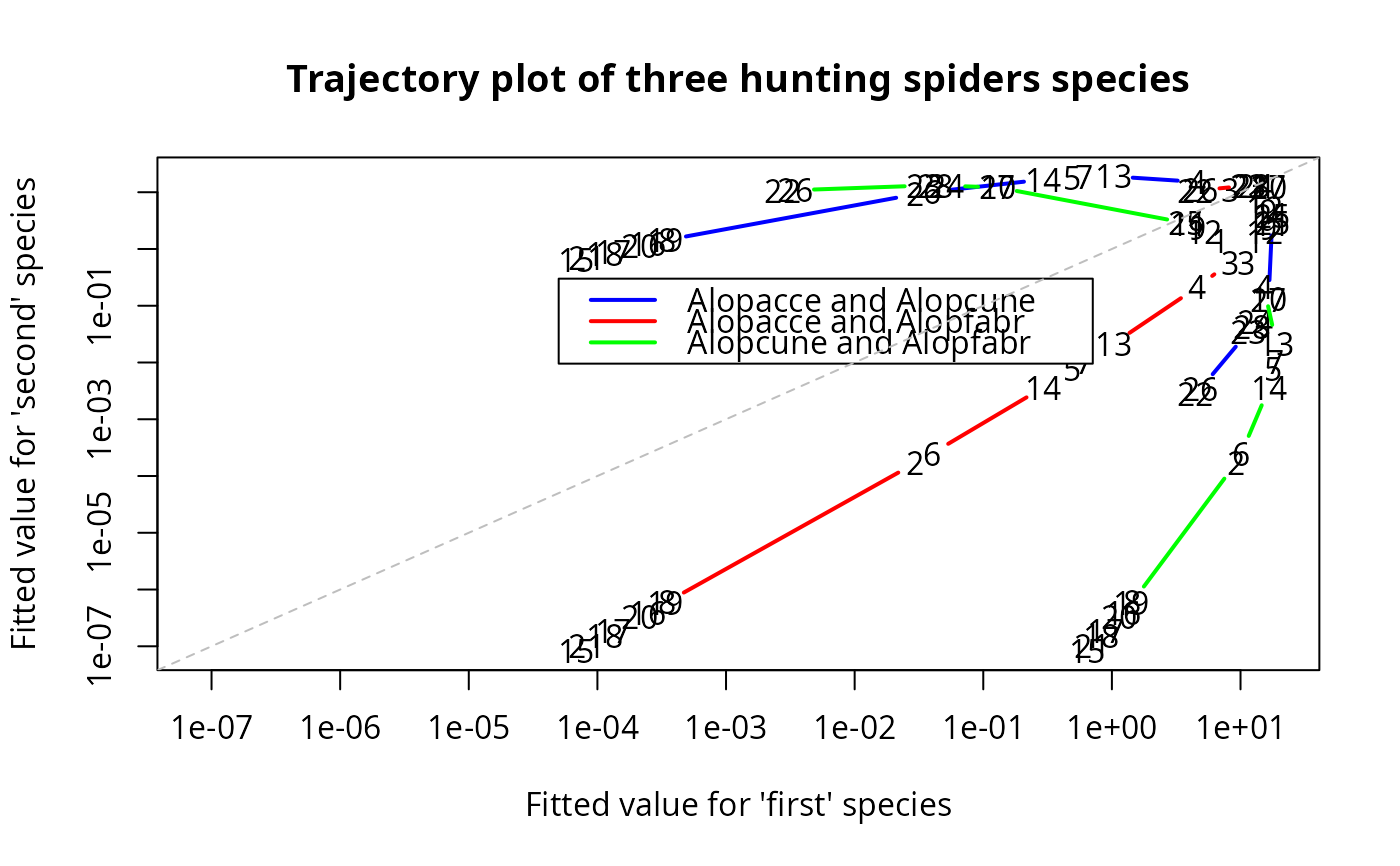
trplot(p1, which.species = 1:3, log = "xy", type = "b", lty = 1,
main = "Trajectory plot of three hunting spiders species",
col = c("blue","red","green"), lwd = 2, label = TRUE) -> ii
legend(0.00005, 0.3, lwd = 2, lty = 1,
col = c("blue", "red", "green"),
with(ii, paste(species.names[,1], species.names[,2],
sep = " and ")))
abline(a = 0, b = 1, lty = "dashed", col = "grey") # Ref. line
 # \dontrun{}
# \dontrun{}