Sample a Meth object with replacement. If how=="random", a random sample of the rows are
sampled, the existing values of meth, item and y are kept but new replicate numbers are
generated. If how=="linked", a random sample of the linked observations (i.e.
observations with identical item and repl values) are
sampled with replacement and replicate numbers are kept. If
how=="item", items are sampled with replacement, and their
observations are included the sampled numner of times.
Arguments
Value
A meth object
Examples
data(fat)
# Different ways of selecting columns and generating replicate numbers
Sub1 <- Meth(fat,meth=2,item=1,repl=3,y=4,print=TRUE)
#> The following variables from the dataframe
#> "fat" are used as the Meth variables:
#> meth: Obs
#> item: Id
#> repl: Rep
#> y: Sub
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
Sub2 <- Meth(fat,2,1,3,4,print=TRUE)
#> The following variables from the dataframe
#> "fat" are used as the Meth variables:
#> meth: Obs
#> item: Id
#> repl: Rep
#> y: Sub
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
Sub3 <- Meth(fat,meth="Obs",item="Id",repl="Rep",y="Sub",print=TRUE)
#> The following variables from the dataframe
#> "fat" are used as the Meth variables:
#> meth: Obs
#> item: Id
#> repl: Rep
#> y: Sub
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
summary( Sub3 )
#> #Replicates
#> Method 3 #Items #Obs: 258 Values: min med max
#> KL 43 43 129 0.39 1.7 4.2
#> SL 43 43 129 0.51 1.7 4.1
plot( Sub3 )
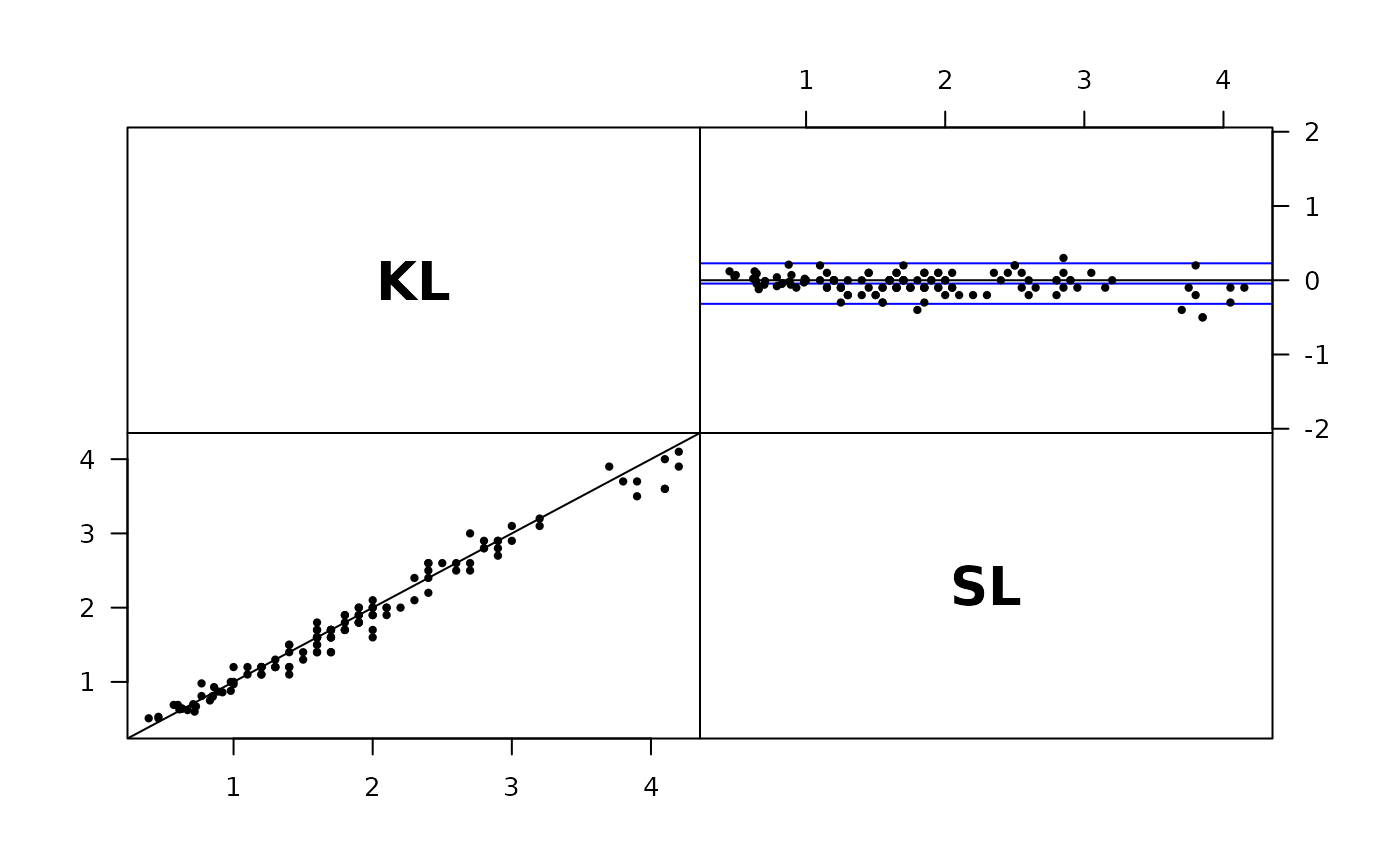 # Use observation in different columns as methods
data( CardOutput )
head( CardOutput )
#> Age Diag VO2 Svo2 Scvo2 TCO FCO Sex
#> 1 66 sepsis 207 63 62 4.9 5.3 M
#> 2 73 sepsis 300 75 78 7.4 10.2 M
#> 3 53 sepsis 289 69 70 8.1 8.7 F
#> 4 28 cardiogenic 150 70 71 3.1 3.7 F
#> 5 72 cardiogenic 350 66 68 5.7 7.6 M
#> 6 71 sepsis 150 70 72 3.1 3.7 M
sv <- Meth( CardOutput, y=c("Svo2","Scvo2") )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 #Items #Obs: 30 Values: min med max
#> Scvo2 15 15 15 61 71 84
#> Svo2 15 15 15 62 70 82
# Note that replicates are generated if a non-unique item-id is used
sv <- Meth( CardOutput, y=c("Svo2","Scvo2"), item="Age" )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> item: Age
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 2 3 #Items #Obs: 30 Values: min med max
#> Scvo2 8 2 1 11 15 61 71 84
#> Svo2 8 2 1 11 15 62 70 82
str( sv )
#> Classes ‘Meth’ and 'data.frame': 30 obs. of 9 variables:
#> $ meth: Factor w/ 2 levels "Scvo2","Svo2": 2 2 2 2 2 2 2 2 2 2 ...
#> $ item: Factor w/ 11 levels "24","28","53",..: 7 10 3 2 9 8 8 3 5 11 ...
#> $ repl: Factor w/ 3 levels "1","2","3": 1 1 1 1 1 1 2 2 1 1 ...
#> $ y : num 63 75 69 70 66 70 75 74 63 64 ...
#> $ Diag: Factor w/ 3 levels "sepsis","cardiogenic",..: 1 1 1 2 2 1 1 1 1 2 ...
#> $ VO2 : num 207 300 289 150 350 150 305 140 167 318 ...
#> $ TCO : num 4.9 7.4 8.1 3.1 5.7 3.1 6.3 3.1 4.1 6.1 ...
#> $ FCO : num 5.3 10.2 8.7 3.7 7.6 3.7 9.6 3.7 3.4 6.8 ...
#> $ Sex : Factor w/ 2 levels "F","M": 2 2 1 1 2 2 1 2 1 1 ...
# A summary is not created if the the first argument (data=) is not used:
sv <- Meth( y=CardOutput[,c("Svo2","Scvo2")], item=CardOutput$VO2 )
#>
#> NOTE: Replication numbers generated in the order of the data
summary(sv)
#> #Replicates
#> Method 1 2 #Items #Obs: 30 Values: min med max
#> Scvo2 13 1 14 15 61 71 84
#> Svo2 13 1 14 15 62 70 82
# Sample items
ssv <- sample.Meth( sv, how="item", N=8 )
# More than two methods
data( sbp )
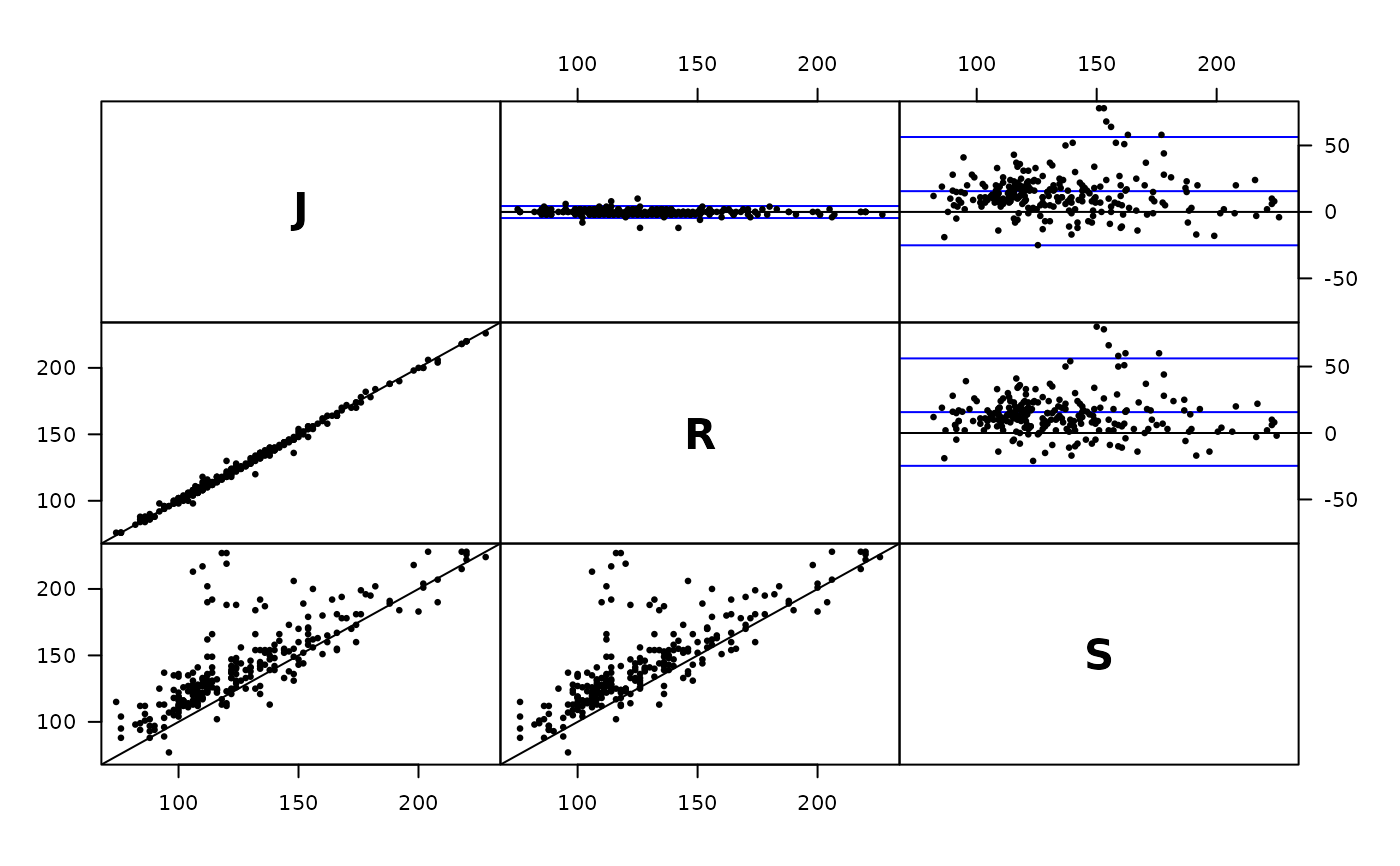
plot( Meth( sbp ) )
#> The following variables from the dataframe
#> "sbp" are used as the Meth variables:
#> meth: meth
#> item: item
#> repl: repl
#> y: y
#> #Replicates
#> Method 3 #Items #Obs: 765 Values: min med max
#> J 85 85 255 74 120 228
#> R 85 85 255 76 120 226
#> S 85 85 255 77 135 228
# Use observation in different columns as methods
data( CardOutput )
head( CardOutput )
#> Age Diag VO2 Svo2 Scvo2 TCO FCO Sex
#> 1 66 sepsis 207 63 62 4.9 5.3 M
#> 2 73 sepsis 300 75 78 7.4 10.2 M
#> 3 53 sepsis 289 69 70 8.1 8.7 F
#> 4 28 cardiogenic 150 70 71 3.1 3.7 F
#> 5 72 cardiogenic 350 66 68 5.7 7.6 M
#> 6 71 sepsis 150 70 72 3.1 3.7 M
sv <- Meth( CardOutput, y=c("Svo2","Scvo2") )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 #Items #Obs: 30 Values: min med max
#> Scvo2 15 15 15 61 71 84
#> Svo2 15 15 15 62 70 82
# Note that replicates are generated if a non-unique item-id is used
sv <- Meth( CardOutput, y=c("Svo2","Scvo2"), item="Age" )
#> The following variables from the dataframe
#> "CardOutput" are used as the Meth variables:
#> item: Age
#> y: Svo2 Scvo2
#> #Replicates
#> Method 1 2 3 #Items #Obs: 30 Values: min med max
#> Scvo2 8 2 1 11 15 61 71 84
#> Svo2 8 2 1 11 15 62 70 82
str( sv )
#> Classes ‘Meth’ and 'data.frame': 30 obs. of 9 variables:
#> $ meth: Factor w/ 2 levels "Scvo2","Svo2": 2 2 2 2 2 2 2 2 2 2 ...
#> $ item: Factor w/ 11 levels "24","28","53",..: 7 10 3 2 9 8 8 3 5 11 ...
#> $ repl: Factor w/ 3 levels "1","2","3": 1 1 1 1 1 1 2 2 1 1 ...
#> $ y : num 63 75 69 70 66 70 75 74 63 64 ...
#> $ Diag: Factor w/ 3 levels "sepsis","cardiogenic",..: 1 1 1 2 2 1 1 1 1 2 ...
#> $ VO2 : num 207 300 289 150 350 150 305 140 167 318 ...
#> $ TCO : num 4.9 7.4 8.1 3.1 5.7 3.1 6.3 3.1 4.1 6.1 ...
#> $ FCO : num 5.3 10.2 8.7 3.7 7.6 3.7 9.6 3.7 3.4 6.8 ...
#> $ Sex : Factor w/ 2 levels "F","M": 2 2 1 1 2 2 1 2 1 1 ...
# A summary is not created if the the first argument (data=) is not used:
sv <- Meth( y=CardOutput[,c("Svo2","Scvo2")], item=CardOutput$VO2 )
#>
#> NOTE: Replication numbers generated in the order of the data
summary(sv)
#> #Replicates
#> Method 1 2 #Items #Obs: 30 Values: min med max
#> Scvo2 13 1 14 15 61 71 84
#> Svo2 13 1 14 15 62 70 82
# Sample items
ssv <- sample.Meth( sv, how="item", N=8 )
# More than two methods
data( sbp )
plot( Meth( sbp ) )
#> The following variables from the dataframe
#> "sbp" are used as the Meth variables:
#> meth: meth
#> item: item
#> repl: repl
#> y: y
#> #Replicates
#> Method 3 #Items #Obs: 765 Values: min med max
#> J 85 85 255 74 120 228
#> R 85 85 255 76 120 226
#> S 85 85 255 77 135 228
 # Creating non-unique replicate numbers per (meth,item) creates a warning:
data( hba1c )
hb1 <- with( hba1c,
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 6 7 8 #Items #Obs: 835 Values: min med max
#> BR.V2 0 19 19 38 285 5.3 8.0 12.6
#> BR.VC 19 0 19 38 266 5.3 8.1 12.1
#> Tosoh 0 20 18 38 284 5.0 7.9 12.1
#>
#> WARNING: You have chosen a replicate variable which is not unique
#> within each (meth,item).
hb2 <- with( subset(hba1c,type=="Cap"),
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 3 4 #Items #Obs: 437 Values: min med max
#> BR.V2 0 38 38 152 5.3 8.0 12.6
#> BR.VC 19 19 38 133 5.3 8.2 12.1
#> Tosoh 0 38 38 152 5.0 7.8 11.8
# Creating non-unique replicate numbers per (meth,item) creates a warning:
data( hba1c )
hb1 <- with( hba1c,
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 6 7 8 #Items #Obs: 835 Values: min med max
#> BR.V2 0 19 19 38 285 5.3 8.0 12.6
#> BR.VC 19 0 19 38 266 5.3 8.1 12.1
#> Tosoh 0 20 18 38 284 5.0 7.9 12.1
#>
#> WARNING: You have chosen a replicate variable which is not unique
#> within each (meth,item).
hb2 <- with( subset(hba1c,type=="Cap"),
Meth( meth=dev, item=item, repl=d.ana-d.samp, y=y, print=TRUE ) )
#> #Replicates
#> Method 3 4 #Items #Obs: 437 Values: min med max
#> BR.V2 0 38 38 152 5.3 8.0 12.6
#> BR.VC 19 19 38 133 5.3 8.2 12.1
#> Tosoh 0 38 38 152 5.0 7.8 11.8