Re-state Restricted Cubic Spline Function
rcspline.restate.RdThis function re-states a restricted cubic spline function in
the un-linearly-restricted form. Coefficients for that form are
returned, along with an R functional representation of this function
and a LaTeX character representation of the function.
rcsplineFunction is a fast function that creates a function to
compute a restricted cubic spline function with given coefficients and
knots, without reformatting the function to be pretty (i.e., into
unrestricted form).
Arguments
- knots
vector of knots used in the regression fit
- coef
vector of coefficients from the fit. If the length of
coefis \(k-1\), where k is equal to thelength(knots), the first coefficient must be for the linear term and remaining \(k-2\) coefficients must be for the constructed terms (e.g., fromrcspline.eval). If the length ofcoefis k, an intercept is assumed to be in the first element (or a zero is prepended tocoefforrcsplineFunction).- type
The default is to represent the cubic spline function corresponding to the coefficients and knots. Set
type = "integral"to instead represent its anti-derivative.- x
a character string to use as the variable name in the LaTeX expression for the formula.
- lx
length of
xto count with respect tocolumns. Default is length of character string contained byx. You may want to setlxsmaller than this if it includes non-printable LaTeX commands.- norm
normalization that was used in deriving the original nonlinear terms used in the fit. See
rcspline.evalfor definitions.- columns
maximum number of symbols in the LaTeX expression to allow before inserting a newline (\\) command. Set to a very large number to keep text all on one line.
- before
text to place before each line of LaTeX output. Use "& &" for an equation array environment in LaTeX where you want to have a left-hand prefix e.g. "f(X) & = &" or using "\lefteqn".
- after
text to place at the end of each line of output.
- begin
text with which to start the first line of output. Useful when adding LaTeX output to part of an existing formula
- nbegin
number of columns of printable text in
begin- digits
number of significant digits to write for coefficients and knots
Value
rcspline.restate returns a vector of coefficients. The
coefficients are un-normalized and two coefficients are added that are
linearly dependent on the other coefficients and knots. The vector of
coefficients has four attributes. knots is a vector of knots,
latex is a vector of text strings with the LaTeX
representation of the formula. columns.used is the number of
columns used in the output string since the last newline command.
function is an R function, which is also return in character
string format as the text attribute. rcsplineFunction
returns an R function with arguments x (a user-supplied
numeric vector at which to evaluate the function), and some
automatically-supplied other arguments.
Author
Frank Harrell
Department of Biostatistics, Vanderbilt University
fh@fharrell.com
See also
rcspline.eval, ns, rcs,
latex, Function.transcan
Examples
set.seed(1)
x <- 1:100
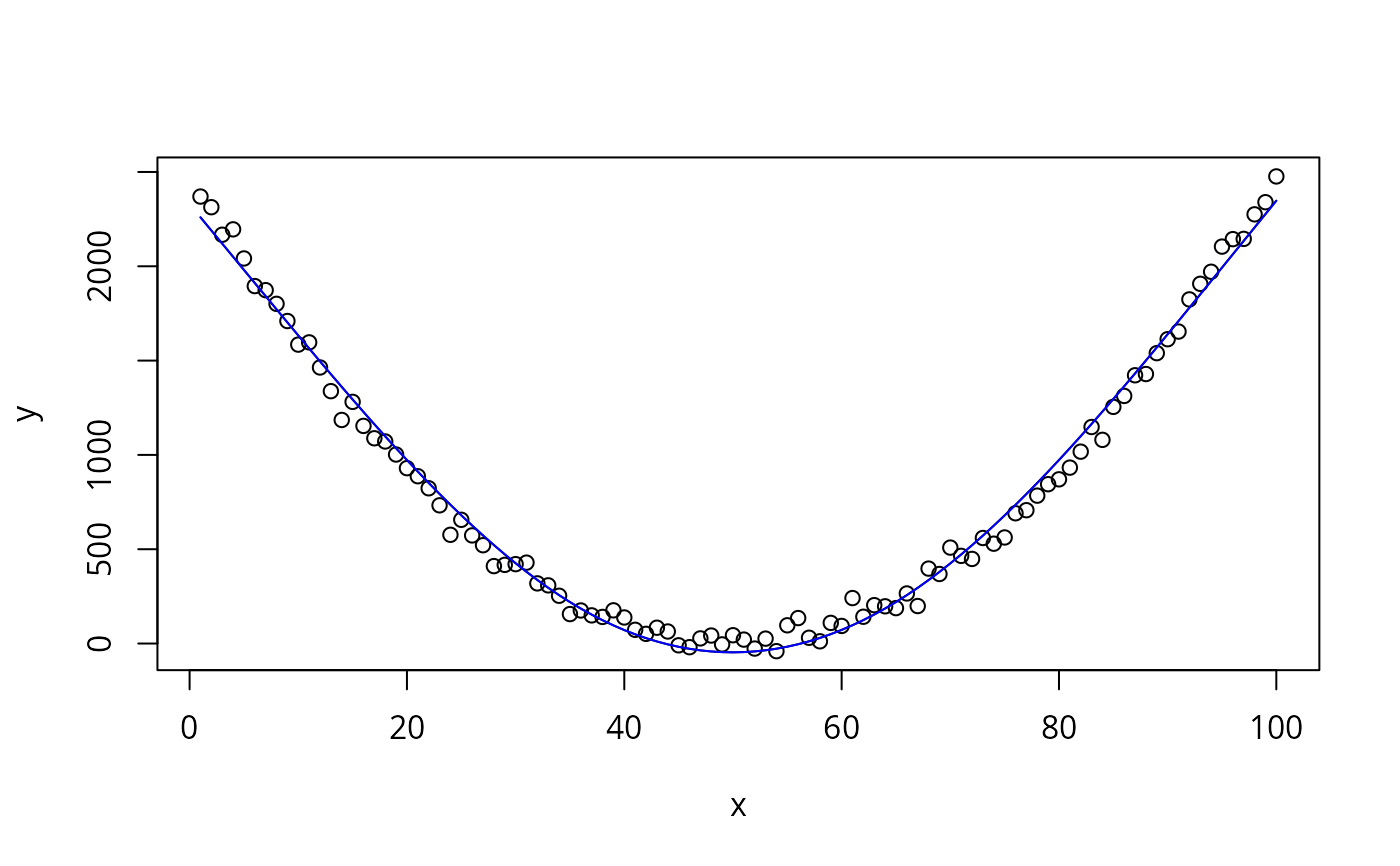
y <- (x - 50)^2 + rnorm(100, 0, 50)
plot(x, y)
xx <- rcspline.eval(x, inclx=TRUE, nk=4)
knots <- attr(xx, "knots")
coef <- lsfit(xx, y)$coef
options(digits=4)
# rcspline.restate must ignore intercept
w <- rcspline.restate(knots, coef[-1], x="{\\rm BP}")
# could also have used coef instead of coef[-1], to include intercept
cat(attr(w,"latex"), sep="\n")
#> & & -69.685564 {\rm BP}+0.013443742 ({\rm BP}-5.95)_{+}^{3} \
#> & & -0.013754656({\rm BP}-35.65)_{+}^{3}-0.012821915({\rm BP}-65.35)_{+}^{3} \
#> & & +0.013132828 ({\rm BP}-95.05)_{+}^{3} \
xtrans <- eval(attr(w, "function"))
# This is an S function of a single argument
lines(x, coef[1] + xtrans(x), type="l")
# Plots fitted transformation
xtrans <- rcsplineFunction(knots, coef)
xtrans
#> function (x = numeric(0), knots = c(5.95, 35.65, 65.35, 95.05
#> ), coef = c(Intercept = 2329.20175140574, x = -69.6855639459134,
#> 106.727315060936, -109.195599900753), kd = 19.9488777705554,
#> type = "ordinary")
#> {
#> k <- length(knots)
#> knotnk <- knots[k]
#> knotnk1 <- knots[k - 1]
#> knot1 <- knots[1]
#> if (length(coef) < k)
#> coef <- c(0, coef)
#> if (type == "ordinary") {
#> y <- coef[1] + coef[2] * x
#> for (j in 1:(k - 2)) y <- y + coef[j + 2] * (pmax((x -
#> knots[j])/kd, 0)^3 + ((knotnk1 - knots[j]) * pmax((x -
#> knotnk)/kd, 0)^3 - (knotnk - knots[j]) * (pmax((x -
#> knotnk1)/kd, 0)^3))/(knotnk - knotnk1))
#> return(y)
#> }
#> y <- coef[1] * x + 0.5 * coef[2] * x * x
#> for (j in 1:(k - 2)) y <- y + 0.25 * coef[j + 2] * kd * (pmax((x -
#> knots[j])/kd, 0)^4 + ((knotnk1 - knots[j]) * pmax((x -
#> knotnk)/kd, 0)^4 - (knotnk - knots[j]) * (pmax((x - knotnk1)/kd,
#> 0)^4))/(knotnk - knotnk1))
#> y
#> }
#> <environment: 0x5e2f8b148688>
lines(x, xtrans(x), col='blue')
 #x <- blood.pressure
xx.simple <- cbind(x, pmax(x-knots[1],0)^3, pmax(x-knots[2],0)^3,
pmax(x-knots[3],0)^3, pmax(x-knots[4],0)^3)
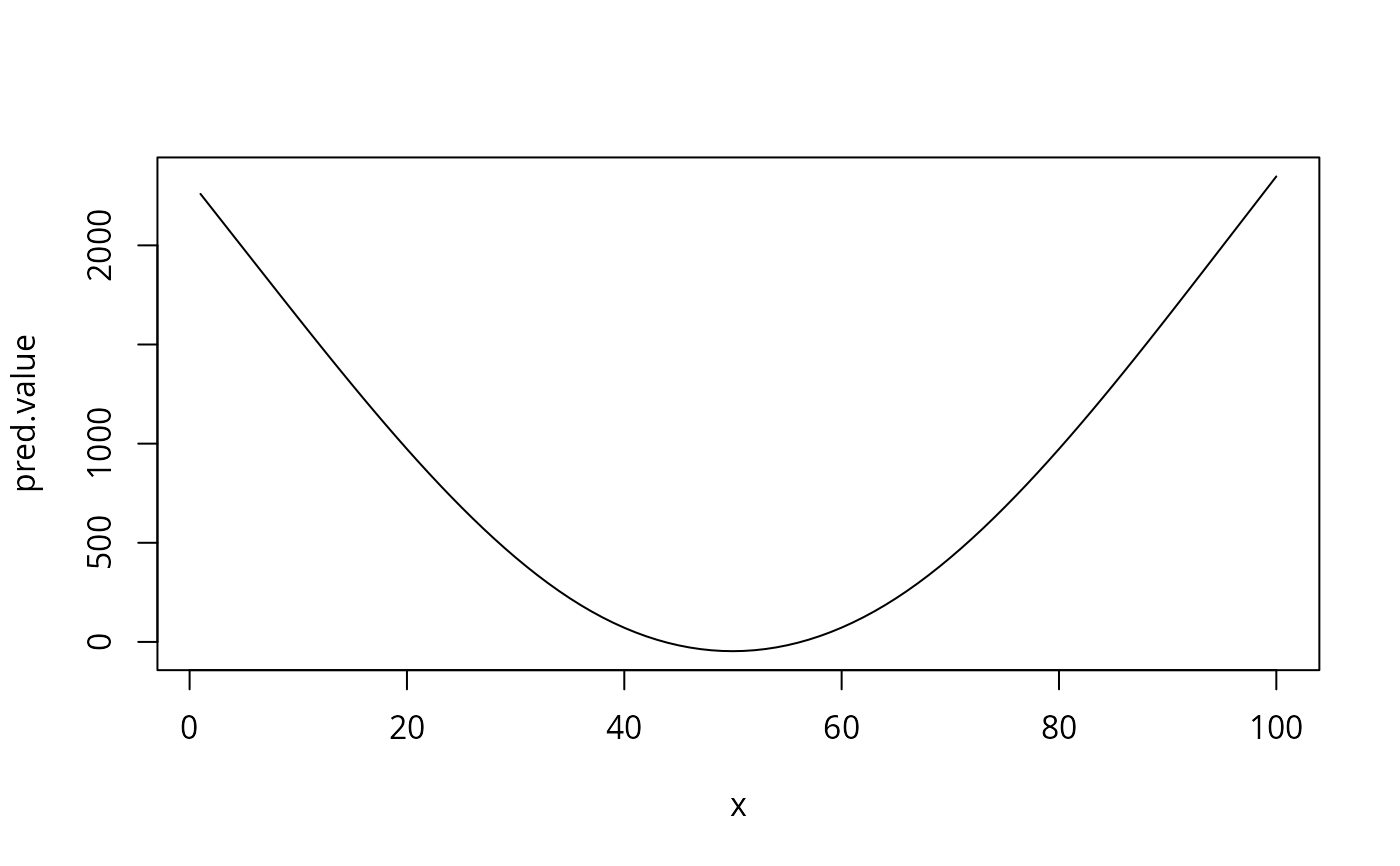
pred.value <- coef[1] + xx.simple %*% w
plot(x, pred.value, type='l') # same as above
#x <- blood.pressure
xx.simple <- cbind(x, pmax(x-knots[1],0)^3, pmax(x-knots[2],0)^3,
pmax(x-knots[3],0)^3, pmax(x-knots[4],0)^3)
pred.value <- coef[1] + xx.simple %*% w
plot(x, pred.value, type='l') # same as above