fit_coxreg_univar(variables, data, at = list(), control = control_coxreg())
fit_coxreg_multivar(variables, data, control = control_coxreg())Arguments
- variables
(named
list)
the names of the variables found indata, passed as a named list and corresponding to thetime,event,arm,strata, andcovariatesterms. Ifarmis missing fromvariables, then only Cox model(s) including thecovariateswill be fitted and the corresponding effect estimates will be tabulated later.- data
(
data.frame)
the dataset containing the variables to fit the models.- at
(
listofnumeric)
when the candidate covariate is anumeric, useatto specify the value of the covariate at which the effect should be estimated.- control
(
list)
a list of parameters as returned by the helper functioncontrol_coxreg().
Value
fit_coxreg_univar()returns acoxreg.univarclass object which is a namedlistwith 5 elements:mod: Cox regression models fitted bysurvival::coxph().data: The original data frame input.control: The original control input.vars: The variables used in the model.at: Value of the covariate at which the effect should be estimated.
fit_coxreg_multivar()returns acoxreg.multivarclass object which is a named list with 4 elements:mod: Cox regression model fitted bysurvival::coxph().data: The original data frame input.control: The original control input.vars: The variables used in the model.
Functions
fit_coxreg_univar(): Fit a series of univariate Cox regression models given the inputs.fit_coxreg_multivar(): Fit a multivariate Cox regression model.
Note
When using fit_coxreg_univar there should be two study arms.
See also
h_cox_regression for relevant helper functions, cox_regression.
Examples
library(survival)
set.seed(1, kind = "Mersenne-Twister")
# Testing dataset [survival::bladder].
dta_bladder <- with(
data = bladder[bladder$enum < 5, ],
data.frame(
time = stop,
status = event,
armcd = as.factor(rx),
covar1 = as.factor(enum),
covar2 = factor(
sample(as.factor(enum)),
levels = 1:4, labels = c("F", "F", "M", "M")
)
)
)
labels <- c("armcd" = "ARM", "covar1" = "A Covariate Label", "covar2" = "Sex (F/M)")
formatters::var_labels(dta_bladder)[names(labels)] <- labels
dta_bladder$age <- sample(20:60, size = nrow(dta_bladder), replace = TRUE)
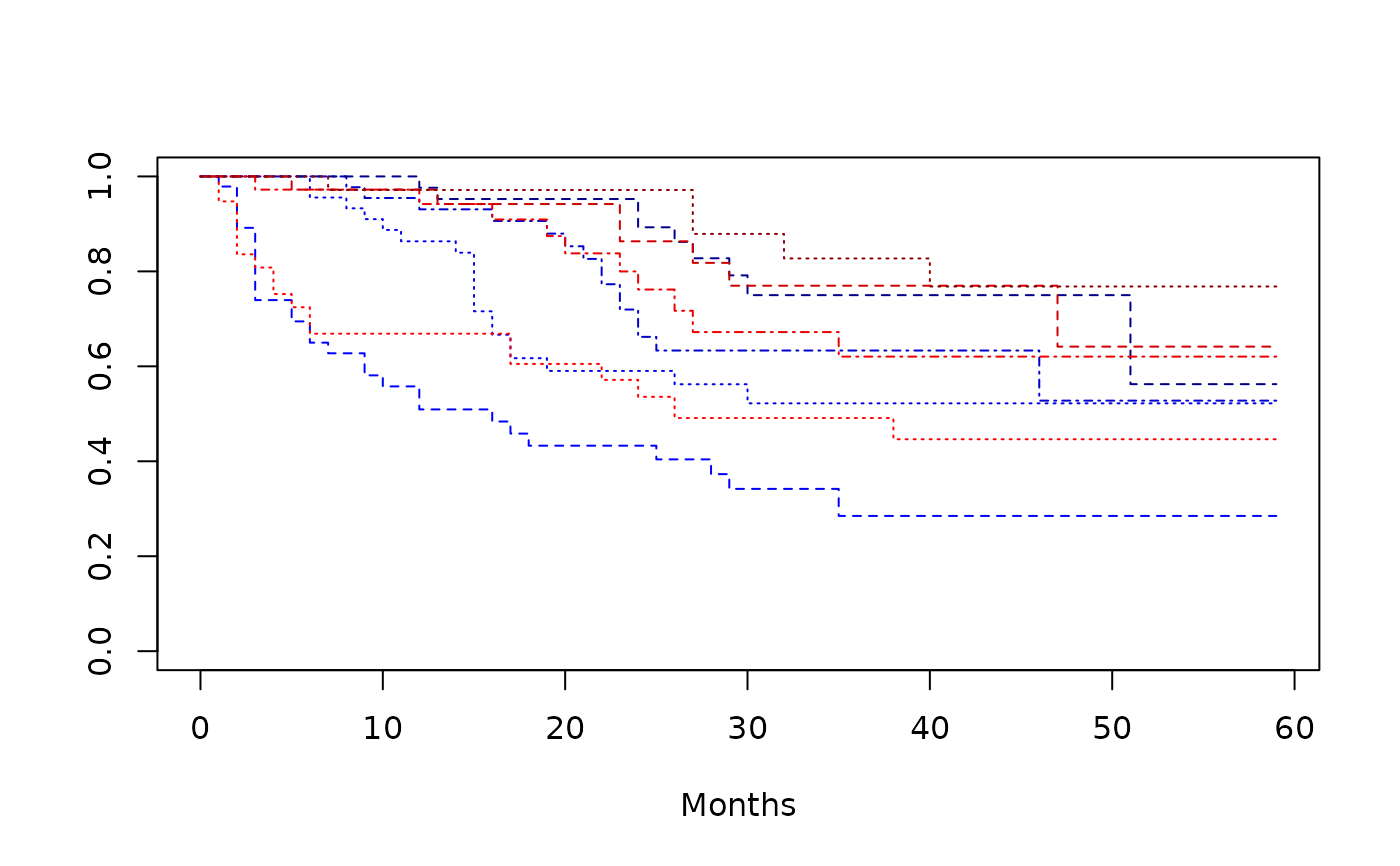
plot(
survfit(Surv(time, status) ~ armcd + covar1, data = dta_bladder),
lty = 2:4,
xlab = "Months",
col = c("blue1", "blue2", "blue3", "blue4", "red1", "red2", "red3", "red4")
)
 # fit_coxreg_univar
## Cox regression: arm + 1 covariate.
mod1 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status", arm = "armcd",
covariates = "covar1"
),
data = dta_bladder,
control = control_coxreg(conf_level = 0.91)
)
## Cox regression: arm + 1 covariate + interaction, 2 candidate covariates.
mod2 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status", arm = "armcd",
covariates = c("covar1", "covar2")
),
data = dta_bladder,
control = control_coxreg(conf_level = 0.91, interaction = TRUE)
)
## Cox regression: arm + 1 covariate, stratified analysis.
mod3 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status", arm = "armcd", strata = "covar2",
covariates = c("covar1")
),
data = dta_bladder,
control = control_coxreg(conf_level = 0.91)
)
## Cox regression: no arm, only covariates.
mod4 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status",
covariates = c("covar1", "covar2")
),
data = dta_bladder
)
# fit_coxreg_multivar
## Cox regression: multivariate Cox regression.
multivar_model <- fit_coxreg_multivar(
variables = list(
time = "time", event = "status", arm = "armcd",
covariates = c("covar1", "covar2")
),
data = dta_bladder
)
# Example without treatment arm.
multivar_covs_model <- fit_coxreg_multivar(
variables = list(
time = "time", event = "status",
covariates = c("covar1", "covar2")
),
data = dta_bladder
)
# fit_coxreg_univar
## Cox regression: arm + 1 covariate.
mod1 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status", arm = "armcd",
covariates = "covar1"
),
data = dta_bladder,
control = control_coxreg(conf_level = 0.91)
)
## Cox regression: arm + 1 covariate + interaction, 2 candidate covariates.
mod2 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status", arm = "armcd",
covariates = c("covar1", "covar2")
),
data = dta_bladder,
control = control_coxreg(conf_level = 0.91, interaction = TRUE)
)
## Cox regression: arm + 1 covariate, stratified analysis.
mod3 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status", arm = "armcd", strata = "covar2",
covariates = c("covar1")
),
data = dta_bladder,
control = control_coxreg(conf_level = 0.91)
)
## Cox regression: no arm, only covariates.
mod4 <- fit_coxreg_univar(
variables = list(
time = "time", event = "status",
covariates = c("covar1", "covar2")
),
data = dta_bladder
)
# fit_coxreg_multivar
## Cox regression: multivariate Cox regression.
multivar_model <- fit_coxreg_multivar(
variables = list(
time = "time", event = "status", arm = "armcd",
covariates = c("covar1", "covar2")
),
data = dta_bladder
)
# Example without treatment arm.
multivar_covs_model <- fit_coxreg_multivar(
variables = list(
time = "time", event = "status",
covariates = c("covar1", "covar2")
),
data = dta_bladder
)