
Smoothing splines using a pspline basis
pspline.RdSpecifies a penalised spline basis for the predictor. This is done by fitting a comparatively small set of splines and penalising the integrated second derivative. Traditional smoothing splines use one basis per observation, but several authors have pointed out that the final results of the fit are indistinguishable for any number of basis functions greater than about 2-3 times the degrees of freedom. Eilers and Marx point out that if the basis functions are evenly spaced, this leads to significant computational simplification, they refer to the result as a p-spline.
Usage
pspline(x, df=4, theta, nterm=2.5 * df, degree=3, eps=0.1, method,
Boundary.knots=range(x), intercept=FALSE, penalty=TRUE, combine, ...)
psplineinverse(x)Arguments
- x
for psline: a covariate vector. The function does not apply to factor variables. For psplineinverse x will be the result of a pspline call.
- df
the desired degrees of freedom. One of the arguments
dfortheta' must be given, but not both. Ifdf=0, then the AIC = (loglik -df) is used to choose an "optimal" degrees of freedom. If AIC is chosen, then an optional argument `caic=T' can be used to specify the corrected AIC of Hurvich et. al.- theta
roughness penalty for the fit. It is a monotone function of the degrees of freedom, with theta=1 corresponding to a linear fit and theta=0 to an unconstrained fit of nterm degrees of freedom.
- nterm
number of splines in the basis
- degree
degree of splines
- eps
accuracy for
df- method
the method for choosing the tuning parameter
theta. If theta is given, then 'fixed' is assumed. If the degrees of freedom is given, then 'df' is assumed. If method='aic' then the degrees of freedom is chosen automatically using Akaike's information criterion.- ...
optional arguments to the control function
- Boundary.knots
the spline is linear beyond the boundary knots. These default to the range of the data.
- intercept
if TRUE, the basis functions include the intercept.
- penalty
if FALSE a large number of attributes having to do with penalized fits are excluded. This is useful to create a pspline basis matrix for other uses.
- combine
an optional vector of increasing integers. If two adjacent values of
combineare equal, then the corresponding coefficients of the fit are forced to be equal. This is useful for monotone fits, see the vignette for more details.
Value
Object of class pspline, coxph.penalty containing the spline basis,
with the appropriate attributes to be
recognized as a penalized term by the coxph or survreg functions.
For psplineinverse the original x vector is reconstructed.
References
Eilers, Paul H. and Marx, Brian D. (1996). Flexible smoothing with B-splines and penalties. Statistical Science, 11, 89-121.
Hurvich, C.M. and Simonoff, J.S. and Tsai, Chih-Ling (1998). Smoothing parameter selection in nonparametric regression using an improved Akaike information criterion, JRSSB, volume 60, 271–293.
Examples
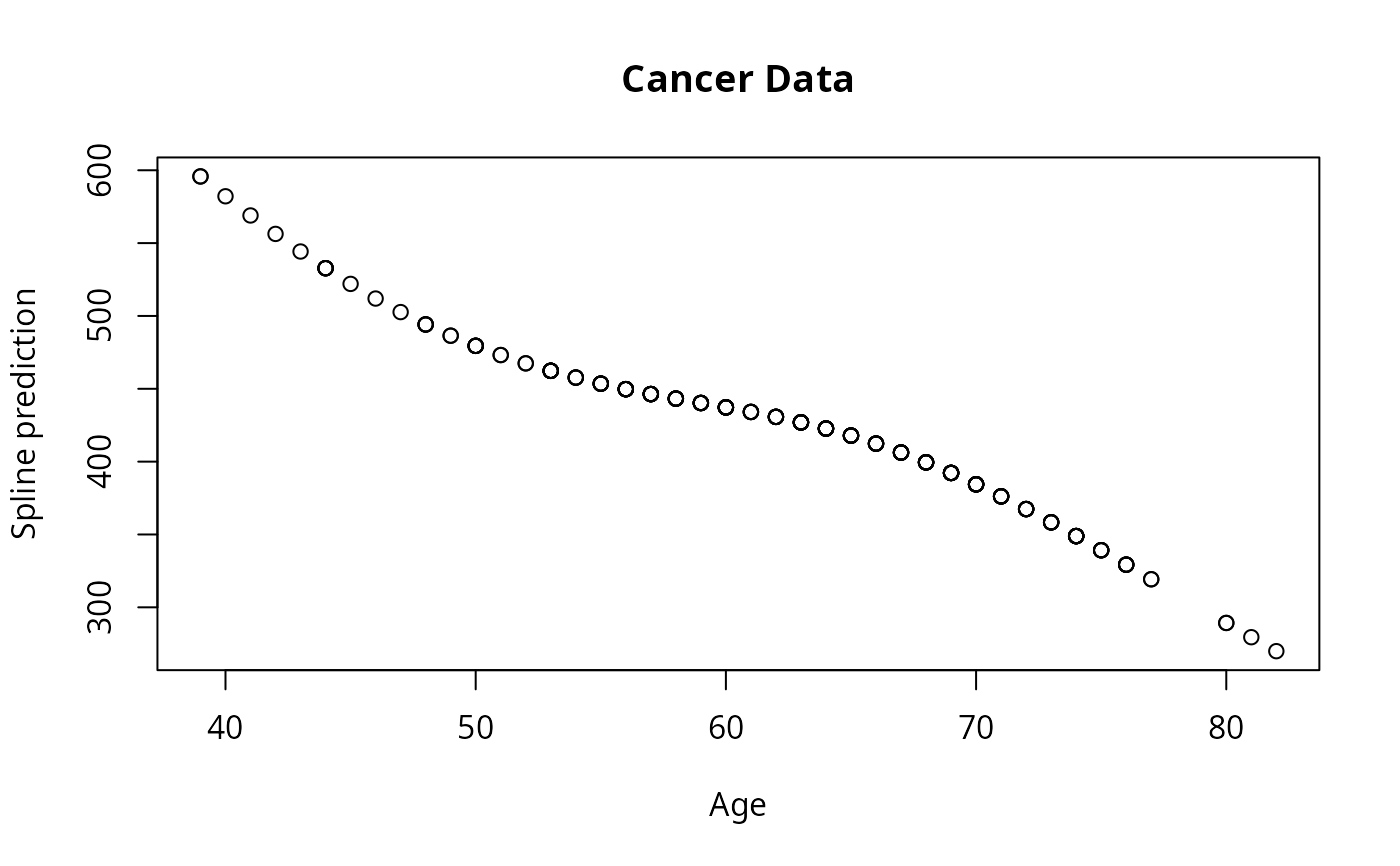
lfit6 <- survreg(Surv(time, status)~pspline(age, df=2), lung)
plot(lung$age, predict(lfit6), xlab='Age', ylab="Spline prediction")
title("Cancer Data")
 fit0 <- coxph(Surv(time, status) ~ ph.ecog + age, lung)
fit1 <- coxph(Surv(time, status) ~ ph.ecog + pspline(age,3), lung)
fit3 <- coxph(Surv(time, status) ~ ph.ecog + pspline(age,8), lung)
fit0
#> Call:
#> coxph(formula = Surv(time, status) ~ ph.ecog + age, data = lung)
#>
#> coef exp(coef) se(coef) z p
#> ph.ecog 0.443485 1.558128 0.115831 3.829 0.000129
#> age 0.011281 1.011345 0.009319 1.211 0.226082
#>
#> Likelihood ratio test=19.06 on 2 df, p=7.279e-05
#> n= 227, number of events= 164
#> (1 observation deleted due to missingness)
fit1
#> Call:
#> coxph(formula = Surv(time, status) ~ ph.ecog + pspline(age, 3),
#> data = lung)
#>
#> coef se(coef) se2 Chisq DF p
#> ph.ecog 0.44802 0.11707 0.11678 14.64453 1.00 0.00013
#> pspline(age, 3), linear 0.01126 0.00928 0.00928 1.47231 1.00 0.22498
#> pspline(age, 3), nonlin 2.07924 2.08 0.37143
#>
#> Iterations: 4 outer, 12 Newton-Raphson
#> Theta= 0.861
#> Degrees of freedom for terms= 1.0 3.1
#> Likelihood ratio test=21.9 on 4.08 df, p=2e-04
#> n= 227, number of events= 164
#> (1 observation deleted due to missingness)
fit3
#> Call:
#> coxph(formula = Surv(time, status) ~ ph.ecog + pspline(age, 8),
#> data = lung)
#>
#> coef se(coef) se2 Chisq DF p
#> ph.ecog 0.47640 0.12024 0.11925 15.69732 1.00 7.4e-05
#> pspline(age, 8), linear 0.01172 0.00923 0.00923 1.61161 1.00 0.20
#> pspline(age, 8), nonlin 6.93188 6.99 0.43
#>
#> Iterations: 5 outer, 15 Newton-Raphson
#> Theta= 0.691
#> Degrees of freedom for terms= 1 8
#> Likelihood ratio test=27.6 on 8.97 df, p=0.001
#> n= 227, number of events= 164
#> (1 observation deleted due to missingness)
fit0 <- coxph(Surv(time, status) ~ ph.ecog + age, lung)
fit1 <- coxph(Surv(time, status) ~ ph.ecog + pspline(age,3), lung)
fit3 <- coxph(Surv(time, status) ~ ph.ecog + pspline(age,8), lung)
fit0
#> Call:
#> coxph(formula = Surv(time, status) ~ ph.ecog + age, data = lung)
#>
#> coef exp(coef) se(coef) z p
#> ph.ecog 0.443485 1.558128 0.115831 3.829 0.000129
#> age 0.011281 1.011345 0.009319 1.211 0.226082
#>
#> Likelihood ratio test=19.06 on 2 df, p=7.279e-05
#> n= 227, number of events= 164
#> (1 observation deleted due to missingness)
fit1
#> Call:
#> coxph(formula = Surv(time, status) ~ ph.ecog + pspline(age, 3),
#> data = lung)
#>
#> coef se(coef) se2 Chisq DF p
#> ph.ecog 0.44802 0.11707 0.11678 14.64453 1.00 0.00013
#> pspline(age, 3), linear 0.01126 0.00928 0.00928 1.47231 1.00 0.22498
#> pspline(age, 3), nonlin 2.07924 2.08 0.37143
#>
#> Iterations: 4 outer, 12 Newton-Raphson
#> Theta= 0.861
#> Degrees of freedom for terms= 1.0 3.1
#> Likelihood ratio test=21.9 on 4.08 df, p=2e-04
#> n= 227, number of events= 164
#> (1 observation deleted due to missingness)
fit3
#> Call:
#> coxph(formula = Surv(time, status) ~ ph.ecog + pspline(age, 8),
#> data = lung)
#>
#> coef se(coef) se2 Chisq DF p
#> ph.ecog 0.47640 0.12024 0.11925 15.69732 1.00 7.4e-05
#> pspline(age, 8), linear 0.01172 0.00923 0.00923 1.61161 1.00 0.20
#> pspline(age, 8), nonlin 6.93188 6.99 0.43
#>
#> Iterations: 5 outer, 15 Newton-Raphson
#> Theta= 0.691
#> Degrees of freedom for terms= 1 8
#> Likelihood ratio test=27.6 on 8.97 df, p=0.001
#> n= 227, number of events= 164
#> (1 observation deleted due to missingness)