Plot method for segmented objects
plot.segmented.RdTakes a fitted segmented object returned by segmented() and plots (or adds)
the fitted broken-line relationship for the selected segmented term.
Usage
# S3 method for class 'segmented'
plot(x, term, add=FALSE, res=FALSE, conf.level=0, interc=TRUE, link=TRUE,
res.col=grey(.15, alpha = .4), rev.sgn=FALSE, const=NULL, shade=FALSE, rug=!add,
dens.rug=FALSE, dens.col = grey(0.8), transf=I, isV=FALSE, is=FALSE, var.diff=FALSE,
p.df="p", .vcov=NULL, .coef=NULL, prev.trend=FALSE, smoos=NULL, hide.zeros=FALSE,
leg="topleft", psi.lines=FALSE, ...)Arguments
- x
a fitted
segmentedobject.- term
Numerical or character to indicate the segmented variable having the piece-wise relationship to be plotted. If there is a single segmented variable in the fitted model
x,termcan be omitted. If vector, multiple segmented lines will be drawn on the same plot.- add
when
TRUEthe fitted lines are added to the current device.- res
when
TRUEthe fitted lines are plotted along with corresponding partial residuals. See Details.- conf.level
If greater than zero, it means the confidence level at which the pointwise confidence itervals have to be plotted.
- interc
If
TRUEthe computed segmented components include the model intercept (if it exists).- link
when
TRUE(default), the fitted lines are plotted on the link scale, otherwise they are tranformed on the response scale before plotting. Ignored for linear segmented fits.- res.col
when
res=TRUEit means the color of the points representing the partial residuals.- rev.sgn
when
TRUEit is assumed that currenttermis `minus' the actual segmented variable, therefore the sign is reversed before plotting. This is useful when a null-constraint has been set on the last slope.- const
constant to add to each fitted segmented relationship (on the scale of the linear predictor) before plotting. If
const=NULLand the fit includes a segmented interaction term (obtained viaseg(..,by)in the formula), the group-specific intercept is included.- shade
if
TRUEandconf.level>0it produces shaded regions (in grey color) for the pointwise confidence intervals embracing the fitted segmented line.- rug
when
TRUEthe covariate values are displayed as a rug plot at the foot of the plot. Default is to!add.- dens.rug
when
TRUEthen smooth covariate distribution is plotted on the x-axis.- dens.col
if
dens.rug=TRUE, it means the colour to be used to plot the density.
- transf
A possible function to convert the fitted values before plotting. It is only effective if
res=FALSE. Ifres=TRUEany transformation is ignored.- isV
logical value (to be passed to
broken.line). Ignored ifconf.level=0- is
logical value (to be passed to
broken.line) indicating if the covariance matrix based on the induced smoothing should be used. Ignored ifconf.level=0- var.diff
logical value to be passed to
summary.segmentedto compute dthe standard errors of fitted values (ifconf.level>0).- p.df
degrees of freedom when
var.diff=TRUE, seesummary.segmented- .vcov
The full covariance matrix of estimates to be used when
conf.level>0. If unspecified (i.e.NULL), the covariance matrix is computed internally by the functionvcov.segmented.- .coef
The regression parameter estimates. If unspecified (i.e.
NULL), it is computed internally bycoef().- prev.trend
logical. If
TRUEdashed lines corresponding to the `previous' trends (i.e. the trends if the breakpoints would not have occurred) are also drawn.- smoos
logical, indicating if the residuals (provided that
res=TRUE) will be drawn using a smoothed scatterplot. IfNULL(default) the smoothed scatterplot will be employed when the number of observation is larger than 10000.- hide.zeros
logical, indicating if the residuals (provided that
res=TRUE) corresponding to the covariate zero values should be deleted. Useful when the fit includes an interaction term in the formula, such asseg(.., by=..), and the zeros in covariates indicate units in other groups.- leg
If the plot refers to segmented relationships in groups, i.e.
termhas been specified as a vector, a legend is placed at the specifiedlegposition. PutNAnot to draw the legend.- psi.lines
if
TRUEvertical lines corresponding to the estimated breakpoints are also drawn. Ignored iftermis not a vector.- ...
other graphics parameters to pass to plotting commands: `col', `lwd' and `lty' (that can be vectors and are recycled if necessary, see the example below) for the fitted piecewise lines; `ylab', `xlab', `main', `sub', `cex.axis', `cex.lab', `xlim' and `ylim' when a new plot is produced (i.e. when
add=FALSE); `pch' and `cex' for the partial residuals (whenres=TRUE,res.colis for the color);col.shadefor the shaded regions (provided thatshade=TRUEandconf.level>0).
Details
Produces (or adds to the current device) the fitted segmented relationship between the
response and the selected term. If the fitted model includes just a single `segmented' variable,
term may be omitted.
The partial residuals are computed as `fitted + residuals', where `fitted' are the fitted values of the
segmented relationship relevant to the covariate specified in term.
Notice that for GLMs the residuals are the response residuals if link=FALSE and
the working residuals if link=TRUE.
Note
For models with offset, partial residuals on the response scale are not defined. Thus plot.segmented does not work
when link=FALSE, res=TRUE, and the fitted model includes an offset.
When term is a vector and multiple segmented relationships are being drawn on the same plot, col and res.col can be vectors. Also pch, cex, lty, and lwd can be vectors, if specified.
See also
segmented to fit the model, lines.segmented to add the estimated breakpoints on the current plot.
points.segmented to add the joinpoints of the segmented relationship.
predict.segmented to compute standard errors and confidence intervals for predictions from a
"segmented" fit.
Examples
set.seed(1234)
z<-runif(100)
y<-rpois(100,exp(2+1.8*pmax(z-.6,0)))
o<-glm(y~z,family=poisson)
o.seg<-segmented(o) #single segmented covariate and one breakpoint: 'seg.Z' and 'psi' can be omitted
par(mfrow=c(1,2))
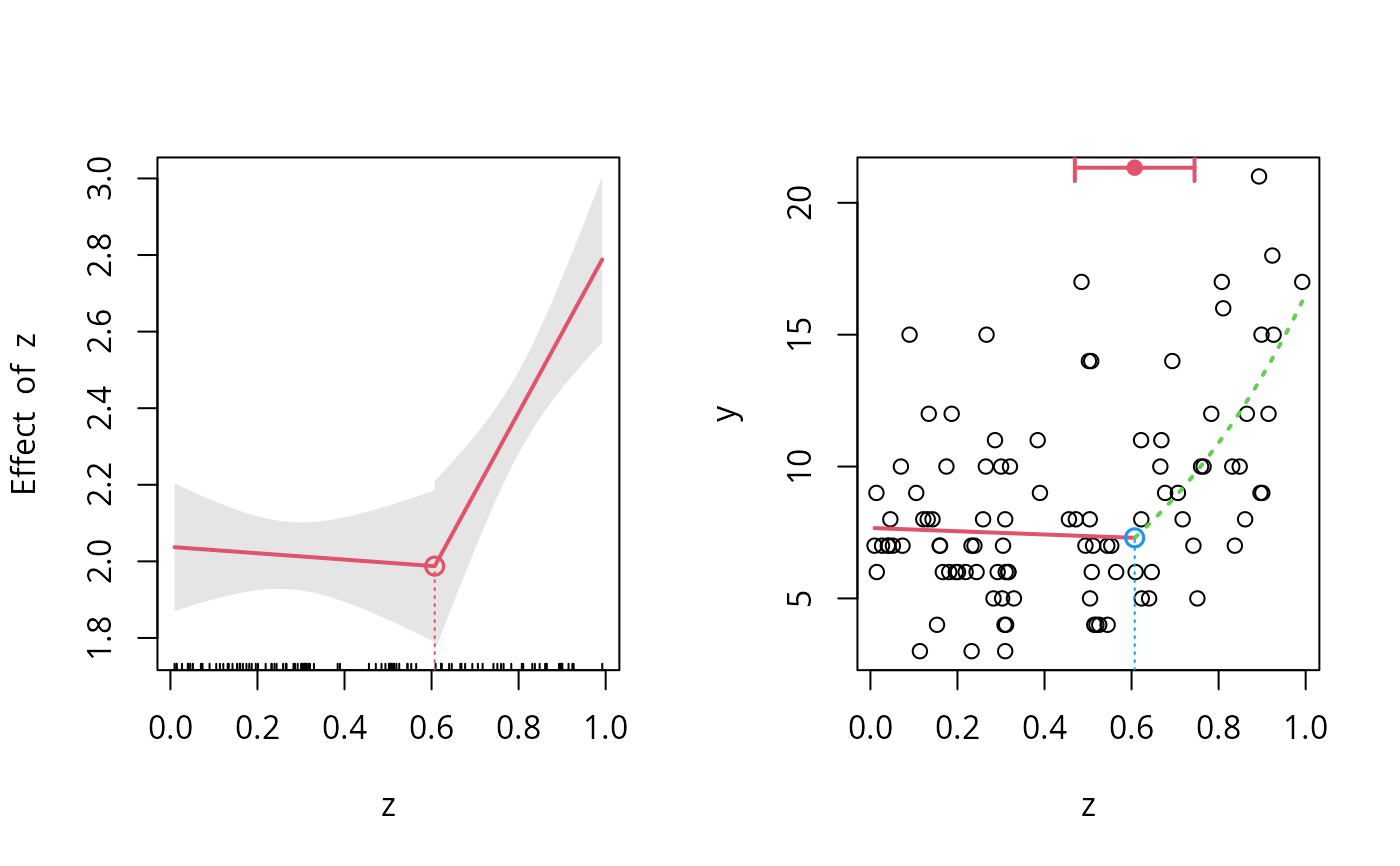
plot(o.seg, conf.level=0.95, shade=TRUE)
points(o.seg, link=TRUE, col=2)
## new plot
plot(z,y)
## add the fitted lines using different colors and styles..
plot(o.seg,add=TRUE,link=FALSE,lwd=2,col=2:3, lty=c(1,3))
lines(o.seg,col=2,pch=19,bottom=FALSE,lwd=2) #for the CI for the breakpoint
points(o.seg,col=4, link=FALSE)
 ## using the options 'is', 'isV', 'shade' and 'col.shade'.
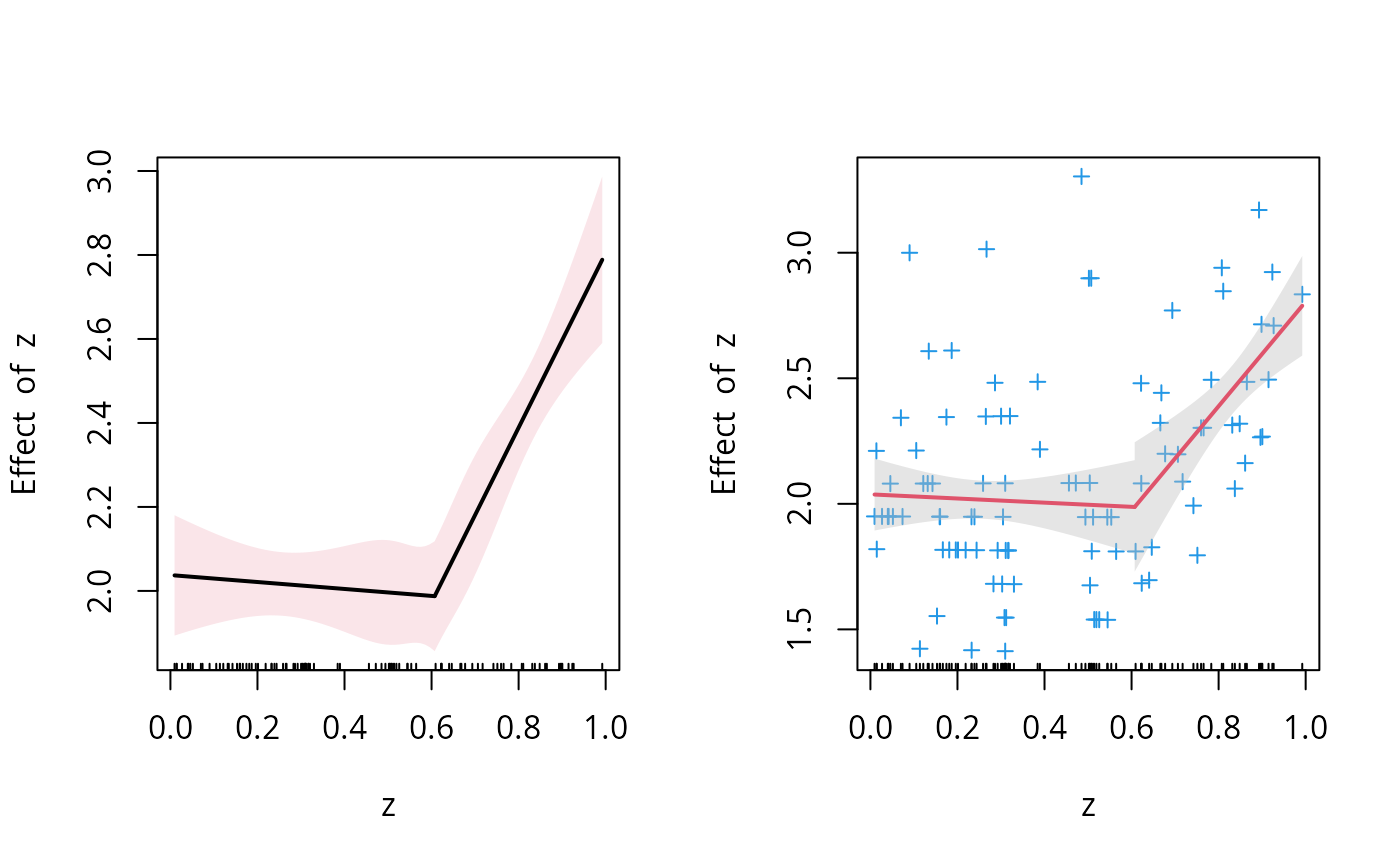
par(mfrow=c(1,2))
plot(o.seg, conf.level=.9, is=TRUE, isV=TRUE, col=1, shade = TRUE, col.shade=2)
plot(o.seg, conf.level=.9, is=TRUE, isV=FALSE, col=2, shade = TRUE, res=TRUE, res.col=4, pch=3)
## using the options 'is', 'isV', 'shade' and 'col.shade'.
par(mfrow=c(1,2))
plot(o.seg, conf.level=.9, is=TRUE, isV=TRUE, col=1, shade = TRUE, col.shade=2)
plot(o.seg, conf.level=.9, is=TRUE, isV=FALSE, col=2, shade = TRUE, res=TRUE, res.col=4, pch=3)