Cate-Nelson models for bivariate data with a fixed critical Y value
cateNelsonFixedY.RdProduces critical-x values for bivariate data according to a Cate-Nelson analysis for a given critical Y value.
cateNelsonFixedY(
x,
y,
cly = 0.95,
plotit = TRUE,
hollow = TRUE,
xlab = "X",
ylab = "Y",
trend = "positive",
clx = 1,
outlength = 20,
sortstat = "error"
)Arguments
- x
A vector of values for the x variable.
- y
A vector of values for the y variable.
- cly
= Critical Y value.
- plotit
If
TRUE, produces plots of the output.- hollow
If
TRUE, uses hollow circles on the plot to indicate data not fitting the model.- xlab
The label for the x-axis.
- ylab
The label for the y-axis.
- trend
"postive"if the trend of y vs. x is generally positive."negative"if negative.- clx
Indicates which of the listed critical x values should be chosen as the critical x value for the plot.
- outlength
Indicates the number of potential critical x values to display in the output.
- sortstat
The statistic to sort by. Any of
"error"(the default),"phi","fisher", or"pearson".
Value
A data frame of statistics from the analysis: critical level for x, critical value for y, the number of observations in each of the quadrants (I, II, III, IV), the number of observations that conform with the model, the number of observations that do not conform to the model, the proportion of observations that conform with the model, the proportion of observations that do not conform to the model, a p-value for the Fisher exact test for the data divided into the groups indicated by the model, phi for the data divided into the groups indicated by the model, and Pearson's chi-square for the data divided into the groups indicated by the model.
Details
Cate-Nelson analysis divides bivariate data into two groups.
For data with a positive trend, one group has a
large x value associated with a large y value, and
the other group has a small x value associated with a small
y value. For a negative trend, a small x is
associated with a large y, and so on.
The analysis is useful for bivariate data which don't conform well to linear, curvilinear, or plateau models.
References
See also
Examples
data(Nurseries)
cateNelsonFixedY(x = Nurseries$Size,
y = Nurseries$Proportion,
cly = 0.70,
plotit = TRUE,
hollow = TRUE,
xlab = "Nursery size in hectares",
ylab = "Proportion of good practices adopted",
trend = "positive",
clx = 1,
outlength = 15)
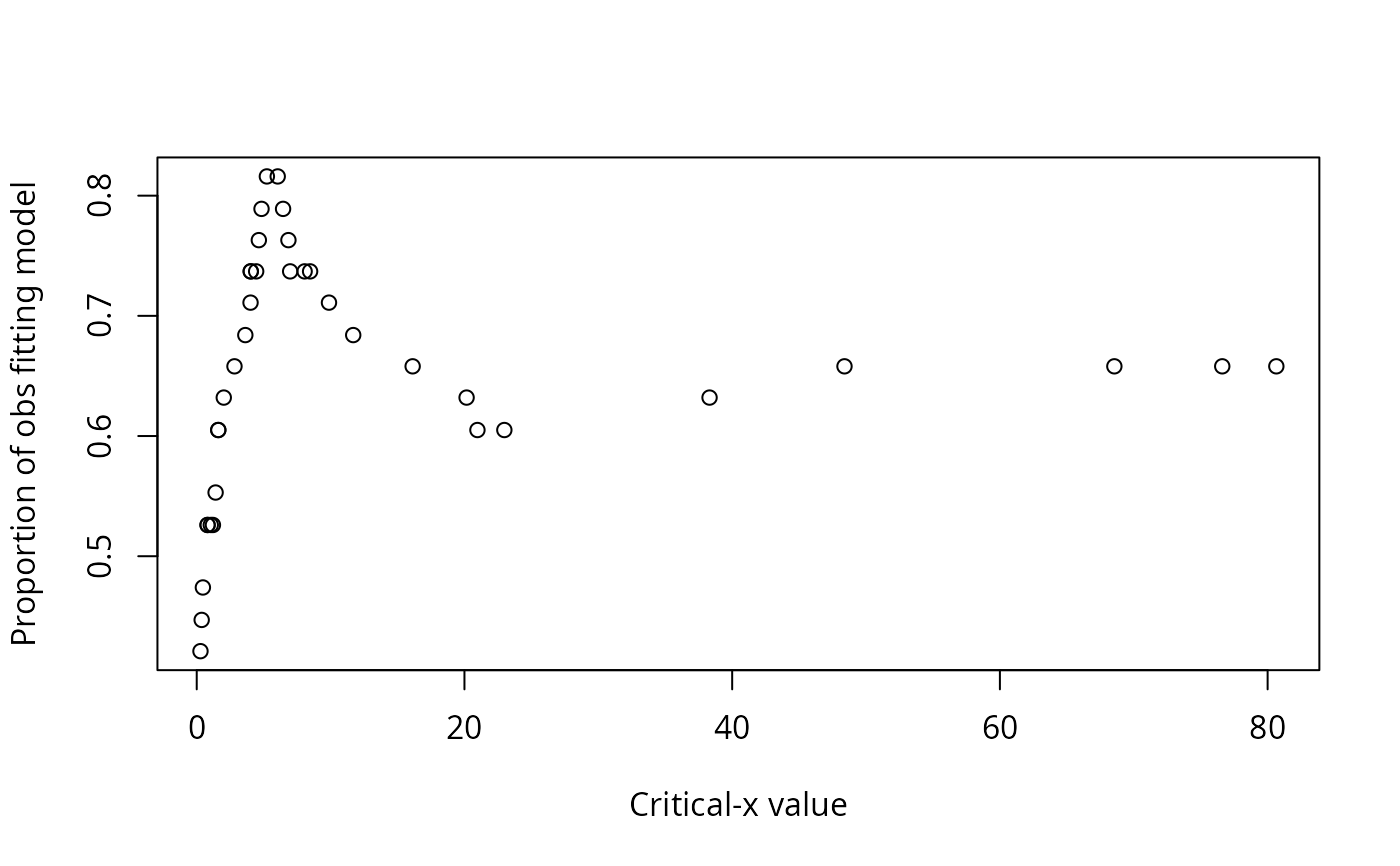
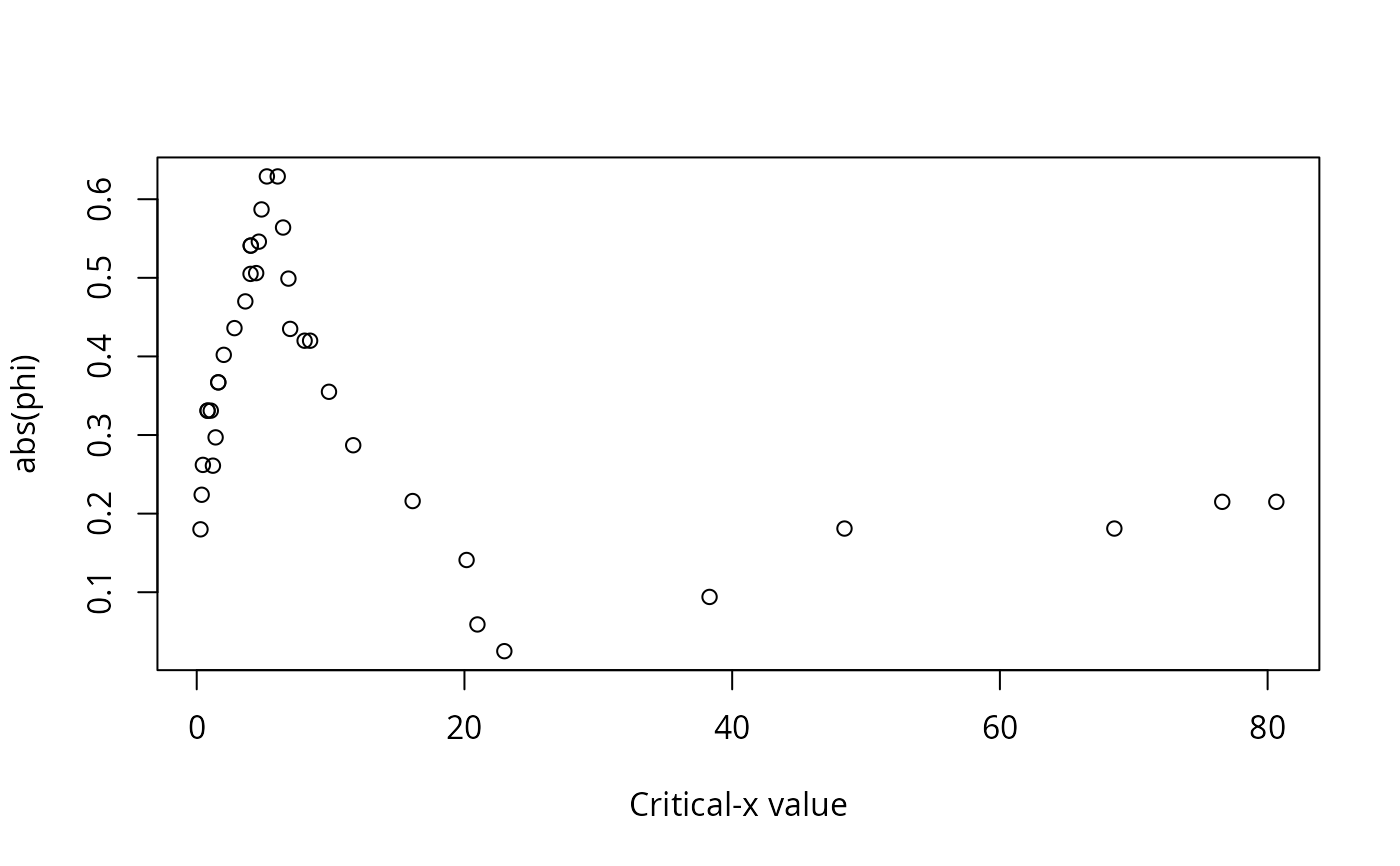
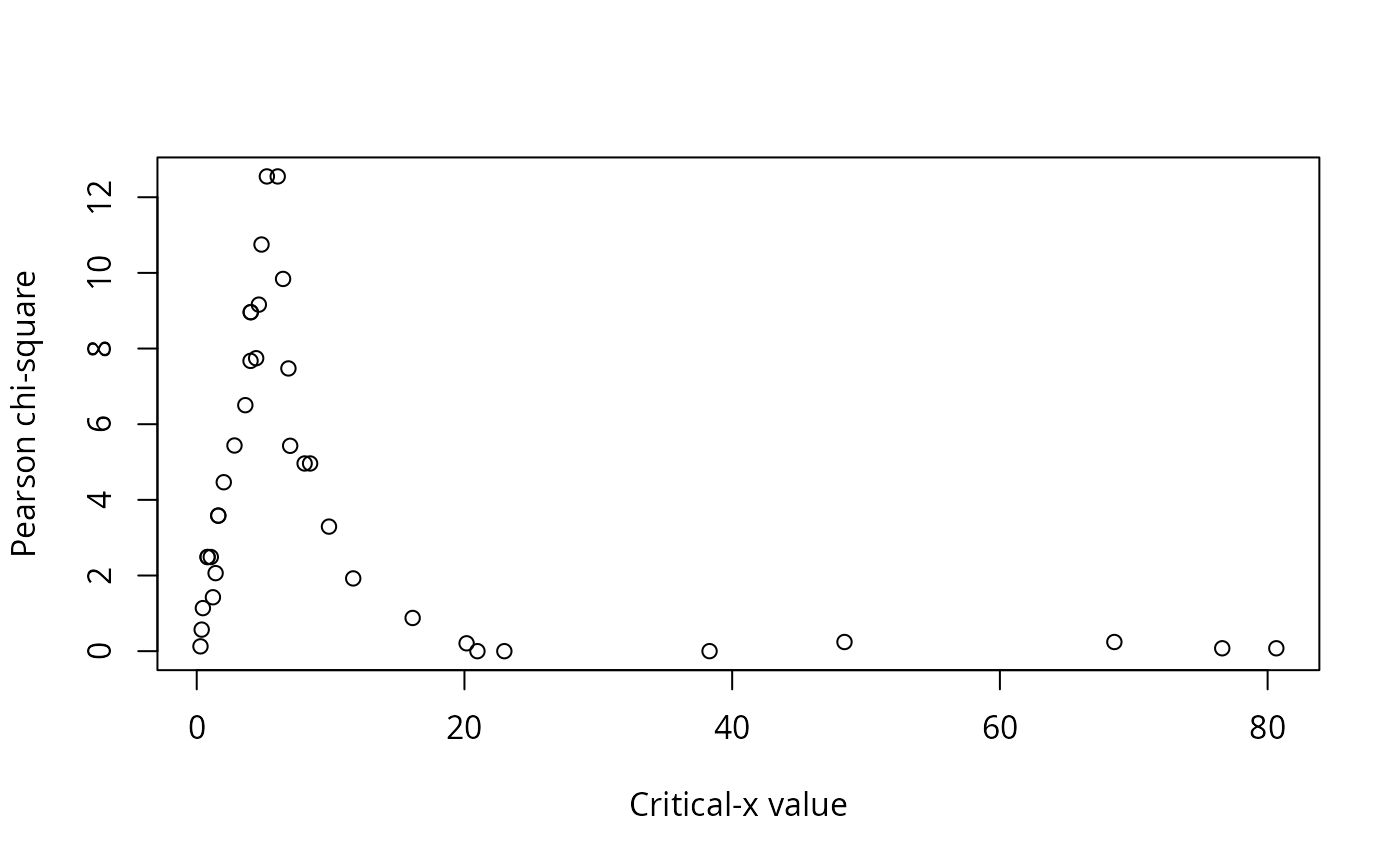
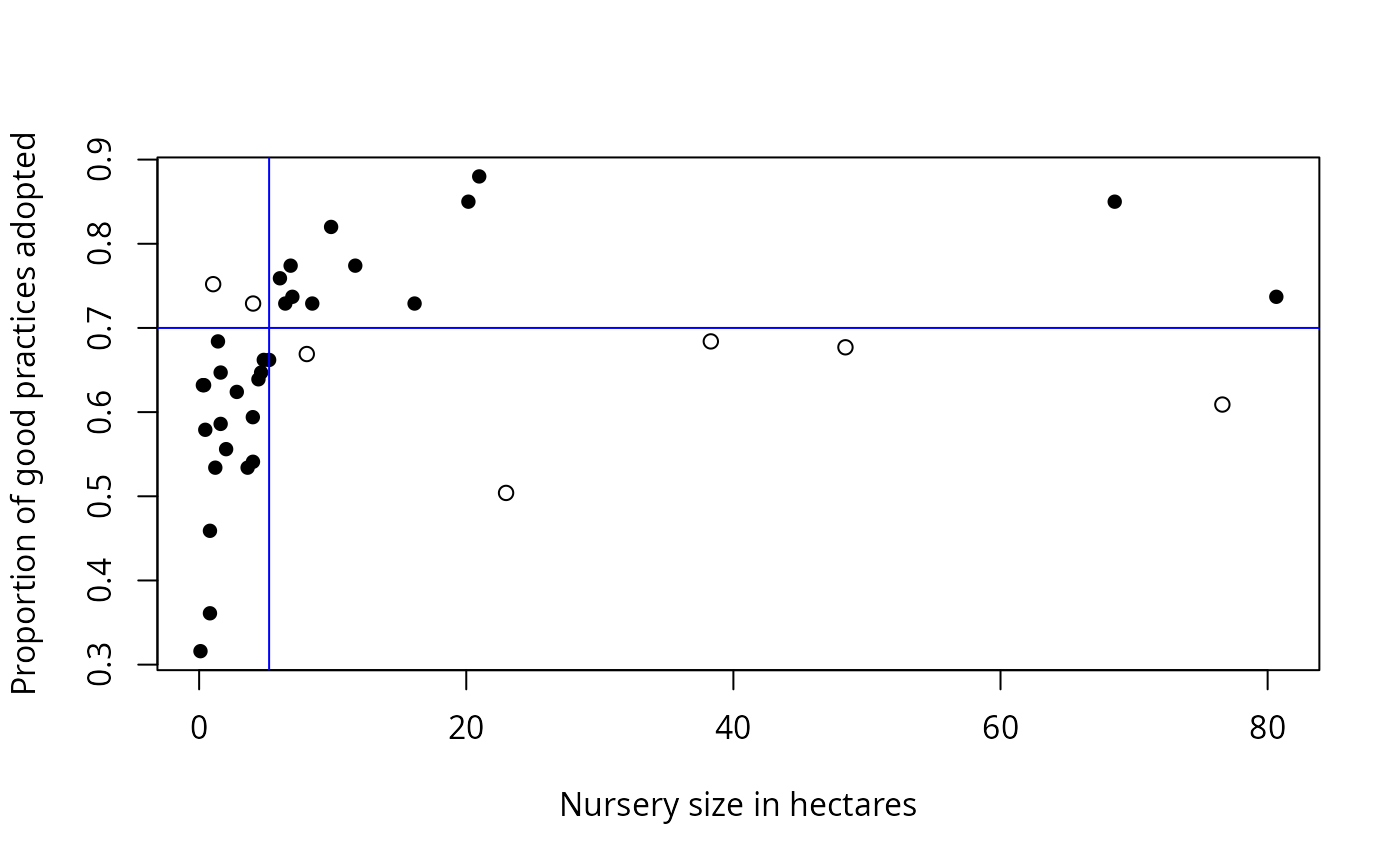 #> Critx Crity Q1 Q2 Q3 Q4 Model Error N pQ1 pQ2 pQ3 pQ4 pModel pError
#> 1 5.24 0.7 2 12 5 19 31 7 38 0.053 0.316 0.132 0.500 0.816 0.184
#> 2 6.05 0.7 2 12 5 19 31 7 38 0.053 0.316 0.132 0.500 0.816 0.184
#> 3 4.84 0.7 2 12 6 18 30 8 38 0.053 0.316 0.158 0.474 0.789 0.211
#> 4 6.45 0.7 3 11 5 19 30 8 38 0.079 0.289 0.132 0.500 0.789 0.211
#> 5 4.64 0.7 2 12 7 17 29 9 38 0.053 0.316 0.184 0.447 0.763 0.237
#> 6 6.85 0.7 4 10 5 19 29 9 38 0.105 0.263 0.132 0.500 0.763 0.237
#> 7 4.03 0.7 1 13 9 15 28 10 38 0.026 0.342 0.237 0.395 0.737 0.263
#> 8 4.04 0.7 1 13 9 15 28 10 38 0.026 0.342 0.237 0.395 0.737 0.263
#> 9 4.44 0.7 2 12 8 16 28 10 38 0.053 0.316 0.211 0.421 0.737 0.263
#> 10 6.98 0.7 5 9 5 19 28 10 38 0.132 0.237 0.132 0.500 0.737 0.263
#> 11 8.06 0.7 6 8 4 20 28 10 38 0.158 0.211 0.105 0.526 0.737 0.263
#> 12 8.47 0.7 6 8 4 20 28 10 38 0.158 0.211 0.105 0.526 0.737 0.263
#> 13 4.02 0.7 1 13 10 14 27 11 38 0.026 0.342 0.263 0.368 0.711 0.289
#> 14 9.88 0.7 7 7 4 20 27 11 38 0.184 0.184 0.105 0.526 0.711 0.289
#> 15 3.63 0.7 1 13 11 13 26 12 38 0.026 0.342 0.289 0.342 0.684 0.316
#> Fisher.p Pearson.chisq Pearson.p phi
#> 1 0.0001517 12.550 0.0003972 -0.629
#> 2 0.0001517 12.550 0.0003972 -0.629
#> 3 0.0005311 10.750 0.0010420 -0.587
#> 4 0.0007735 9.840 0.0017080 -0.564
#> 5 0.0018910 9.161 0.0024730 -0.546
#> 6 0.0048710 7.474 0.0062580 -0.499
#> 7 0.0015970 8.961 0.0027590 -0.541
#> 8 0.0015970 8.961 0.0027590 -0.541
#> 9 0.0025240 7.744 0.0053900 -0.506
#> 10 0.0138000 5.429 0.0198100 -0.435
#> 11 0.0143900 4.962 0.0259100 -0.420
#> 12 0.0143900 4.962 0.0259100 -0.420
#> 13 0.0021210 7.674 0.0056030 -0.505
#> 14 0.0607300 3.293 0.0695600 -0.355
#> 15 0.0049960 6.503 0.0107700 -0.470
#> Critx Crity Q1 Q2 Q3 Q4 Model Error N pQ1 pQ2 pQ3 pQ4 pModel pError
#> 1 5.24 0.7 2 12 5 19 31 7 38 0.053 0.316 0.132 0.500 0.816 0.184
#> 2 6.05 0.7 2 12 5 19 31 7 38 0.053 0.316 0.132 0.500 0.816 0.184
#> 3 4.84 0.7 2 12 6 18 30 8 38 0.053 0.316 0.158 0.474 0.789 0.211
#> 4 6.45 0.7 3 11 5 19 30 8 38 0.079 0.289 0.132 0.500 0.789 0.211
#> 5 4.64 0.7 2 12 7 17 29 9 38 0.053 0.316 0.184 0.447 0.763 0.237
#> 6 6.85 0.7 4 10 5 19 29 9 38 0.105 0.263 0.132 0.500 0.763 0.237
#> 7 4.03 0.7 1 13 9 15 28 10 38 0.026 0.342 0.237 0.395 0.737 0.263
#> 8 4.04 0.7 1 13 9 15 28 10 38 0.026 0.342 0.237 0.395 0.737 0.263
#> 9 4.44 0.7 2 12 8 16 28 10 38 0.053 0.316 0.211 0.421 0.737 0.263
#> 10 6.98 0.7 5 9 5 19 28 10 38 0.132 0.237 0.132 0.500 0.737 0.263
#> 11 8.06 0.7 6 8 4 20 28 10 38 0.158 0.211 0.105 0.526 0.737 0.263
#> 12 8.47 0.7 6 8 4 20 28 10 38 0.158 0.211 0.105 0.526 0.737 0.263
#> 13 4.02 0.7 1 13 10 14 27 11 38 0.026 0.342 0.263 0.368 0.711 0.289
#> 14 9.88 0.7 7 7 4 20 27 11 38 0.184 0.184 0.105 0.526 0.711 0.289
#> 15 3.63 0.7 1 13 11 13 26 12 38 0.026 0.342 0.289 0.342 0.684 0.316
#> Fisher.p Pearson.chisq Pearson.p phi
#> 1 0.0001517 12.550 0.0003972 -0.629
#> 2 0.0001517 12.550 0.0003972 -0.629
#> 3 0.0005311 10.750 0.0010420 -0.587
#> 4 0.0007735 9.840 0.0017080 -0.564
#> 5 0.0018910 9.161 0.0024730 -0.546
#> 6 0.0048710 7.474 0.0062580 -0.499
#> 7 0.0015970 8.961 0.0027590 -0.541
#> 8 0.0015970 8.961 0.0027590 -0.541
#> 9 0.0025240 7.744 0.0053900 -0.506
#> 10 0.0138000 5.429 0.0198100 -0.435
#> 11 0.0143900 4.962 0.0259100 -0.420
#> 12 0.0143900 4.962 0.0259100 -0.420
#> 13 0.0021210 7.674 0.0056030 -0.505
#> 14 0.0607300 3.293 0.0695600 -0.355
#> 15 0.0049960 6.503 0.0107700 -0.470