Model comparison for regression, mediation, cluster and factor analysis
anova.psych.RdWhen doing regressions from the data or from a correlation matrix using setCor or doing a mediation analysis using link{mediate}, it is useful to compare alternative models. Since these are both regression models, the appropriate test is an Analysis of Variance. Similar tests, using Chi Square may be done for factor analytic models.
# S3 method for class 'psych'
anova(object,...)Arguments
Details
setCor returns the SE.residual and degrees of freedom.
These are converted to SSR and then an analysis of variance is used to compare two (or more) models.
For omega or fa the change in the ML chisquare statistic as a function of change in df is reported.
Value
An ANOVA table comparing the models.
Note
The code has been adapted from the anova.lm function in stats and the anova.sem by John Fox.
Examples
if(require("psychTools")) {
m1 <- lmCor(reaction ~ import, data = Tal_Or,std=FALSE)
m2 <- lmCor(reaction ~ import+pmi, data = Tal_Or,std=FALSE)
m3 <- lmCor(reaction ~ import+pmi + cond, data = Tal_Or,std=FALSE)
anova(m1,m2,m3)
}
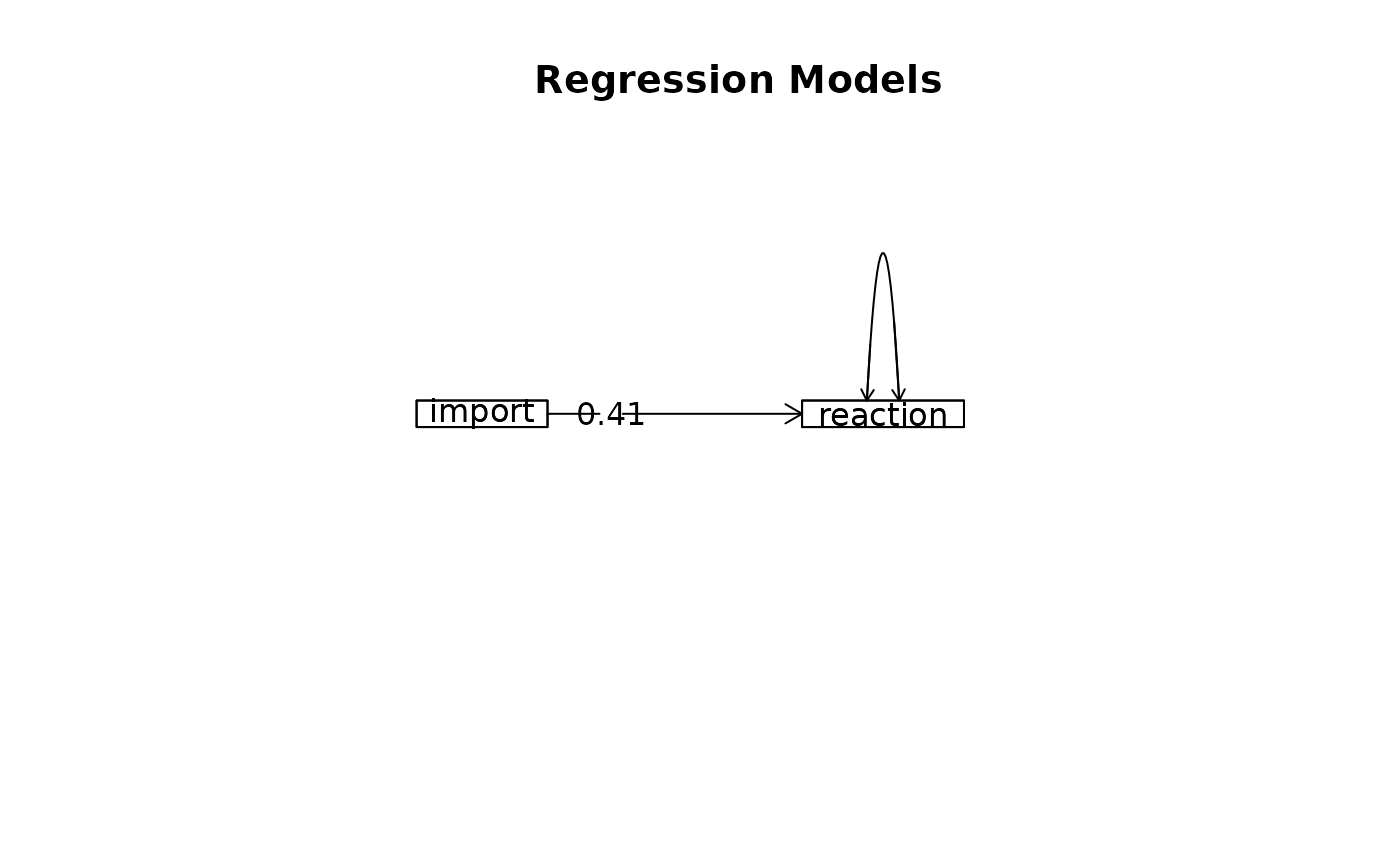
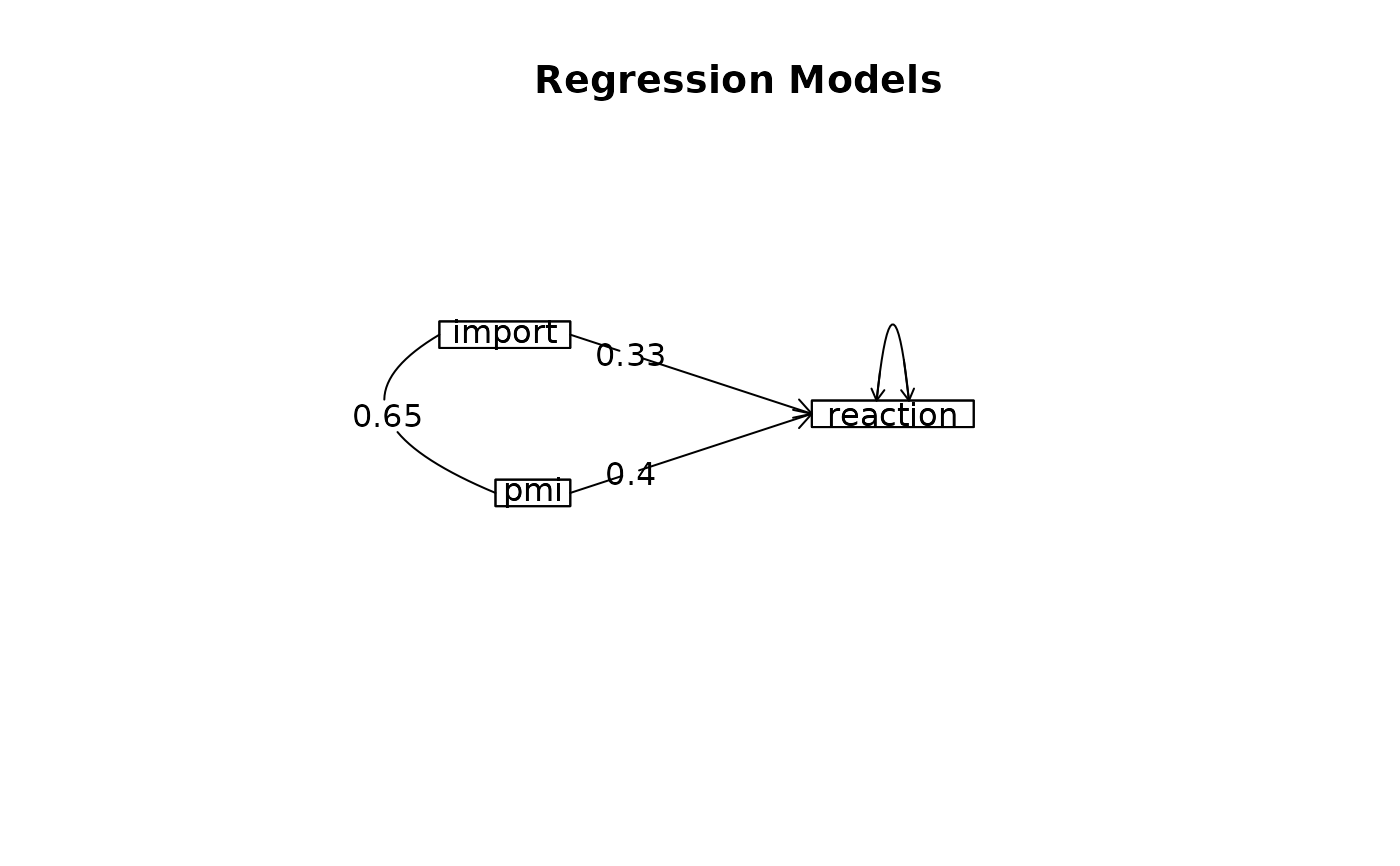
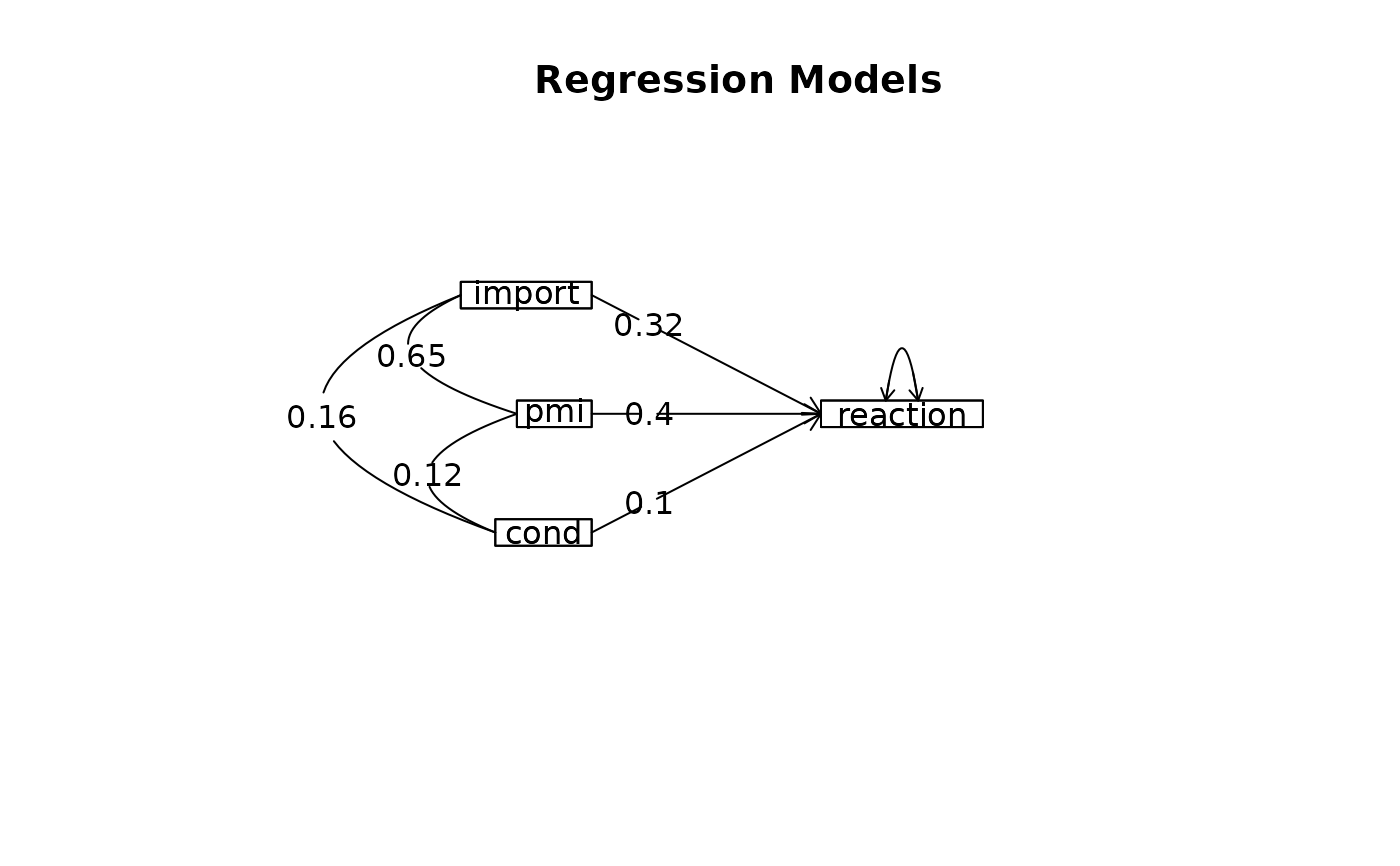 #> Model 1 = lmCor(y = reaction ~ import, data = Tal_Or, std = FALSE)
#> Model 2 = lmCor(y = reaction ~ import + pmi, data = Tal_Or, std = FALSE)
#> Model 3 = lmCor(y = reaction ~ import + pmi + cond, data = Tal_Or, std = FALSE)
#> $reaction
#> Res Df Res SS Diff df Diff SS F Pr(F > )
#> 1 121 229.8774 NA NA NA NA
#> 2 120 198.1888 1 31.6885717 19.0569054 2.705402e-05
#> 3 119 197.8779 1 0.3109252 0.1869845 6.662209e-01
#>
#Several interesting test cases are taken from analyses of the Spengler data set
#Although the sample sizes are actually very large in the first wave, I use the
#sample sizes from the last wave
#This data set is actually in psychTools but is copied here until we can update psychTools
#We set the n.iter to be 50 instead of the default value of 5,000
if(require("psychTools")) {
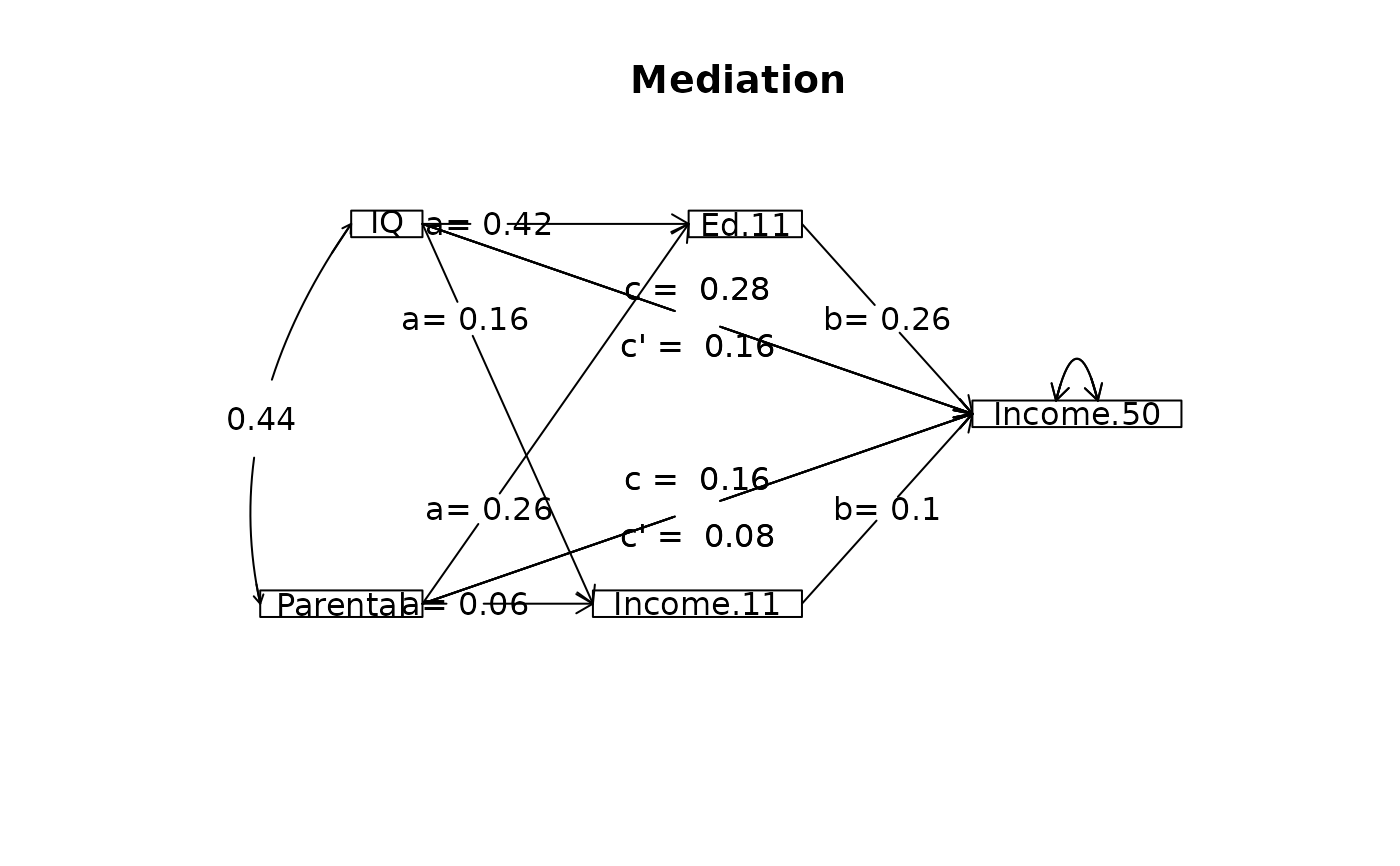
mod1 <- mediate(Income.50 ~ IQ + Parental+ (Ed.11) ,data=Spengler,
n.obs = 1952, n.iter=50)
mod2 <- mediate(Income.50 ~ IQ + Parental+ (Ed.11) + (Income.11)
,data=Spengler,n.obs = 1952, n.iter=50)
#Now, compare these models
anova(mod1,mod2)
}
#> The replication data matrices were simulated based upon the specified number of subjects and the observed correlation matrix.
#> Model 1 = lmCor(y = reaction ~ import, data = Tal_Or, std = FALSE)
#> Model 2 = lmCor(y = reaction ~ import + pmi, data = Tal_Or, std = FALSE)
#> Model 3 = lmCor(y = reaction ~ import + pmi + cond, data = Tal_Or, std = FALSE)
#> $reaction
#> Res Df Res SS Diff df Diff SS F Pr(F > )
#> 1 121 229.8774 NA NA NA NA
#> 2 120 198.1888 1 31.6885717 19.0569054 2.705402e-05
#> 3 119 197.8779 1 0.3109252 0.1869845 6.662209e-01
#>
#Several interesting test cases are taken from analyses of the Spengler data set
#Although the sample sizes are actually very large in the first wave, I use the
#sample sizes from the last wave
#This data set is actually in psychTools but is copied here until we can update psychTools
#We set the n.iter to be 50 instead of the default value of 5,000
if(require("psychTools")) {
mod1 <- mediate(Income.50 ~ IQ + Parental+ (Ed.11) ,data=Spengler,
n.obs = 1952, n.iter=50)
mod2 <- mediate(Income.50 ~ IQ + Parental+ (Ed.11) + (Income.11)
,data=Spengler,n.obs = 1952, n.iter=50)
#Now, compare these models
anova(mod1,mod2)
}
#> The replication data matrices were simulated based upon the specified number of subjects and the observed correlation matrix.
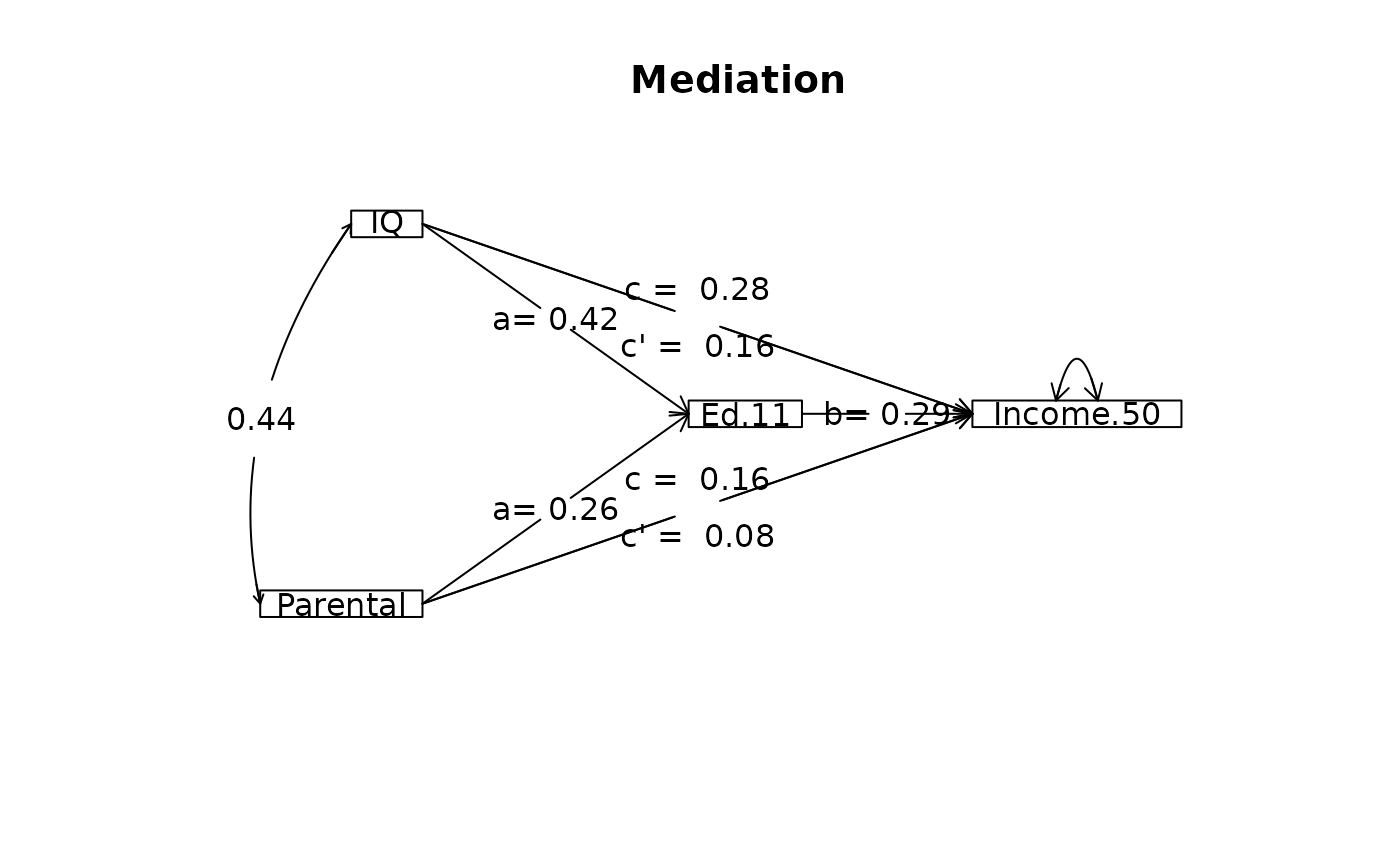 #> The replication data matrices were simulated based upon the specified number of subjects and the observed correlation matrix.
#> The replication data matrices were simulated based upon the specified number of subjects and the observed correlation matrix.
 #> Model 1 = mediate(y = Income.50 ~ IQ + Parental + (Ed.11), data = Spengler,
#> n.obs = 1952, n.iter = 50)
#> Model 2 = mediate(y = Income.50 ~ IQ + Parental + (Ed.11) + (Income.11),
#> data = Spengler, n.obs = 1952, n.iter = 50)
#> Res Df Res SS Diff df Diff SS F Pr(F > )
#> 1 1948 1565.401 NA NA NA NA
#> 2 1947 1547.155 1 18.2464 22.96198 1.777401e-06
f3 <- fa(Thurstone,3,n.obs=213) #we need to specifiy the n.obs for the test to work
f2 <- fa(Thurstone,2, n.obs=213)
anova(f2,f3)
#> Model 1 = fa(r = Thurstone, nfactors = 2, n.obs = 213)
#> Model 2 = fa(r = Thurstone, nfactors = 3, n.obs = 213)
#> df d.df chiSq d.chiSq PR test empirical d.empirical test.echi BIC d.BIC
#> 1 19 NA 86.62 NA NA NA 66.73 NA NA -15.25 NA
#> 2 12 7 2.98 83.64 0 11.95 0.52 66.2 9.46 -61.36 -83.64
#> RMSEA rms
#> 1 0.13 0.07
#> 2 0.00 0.01
#> Model 1 = mediate(y = Income.50 ~ IQ + Parental + (Ed.11), data = Spengler,
#> n.obs = 1952, n.iter = 50)
#> Model 2 = mediate(y = Income.50 ~ IQ + Parental + (Ed.11) + (Income.11),
#> data = Spengler, n.obs = 1952, n.iter = 50)
#> Res Df Res SS Diff df Diff SS F Pr(F > )
#> 1 1948 1565.401 NA NA NA NA
#> 2 1947 1547.155 1 18.2464 22.96198 1.777401e-06
f3 <- fa(Thurstone,3,n.obs=213) #we need to specifiy the n.obs for the test to work
f2 <- fa(Thurstone,2, n.obs=213)
anova(f2,f3)
#> Model 1 = fa(r = Thurstone, nfactors = 2, n.obs = 213)
#> Model 2 = fa(r = Thurstone, nfactors = 3, n.obs = 213)
#> df d.df chiSq d.chiSq PR test empirical d.empirical test.echi BIC d.BIC
#> 1 19 NA 86.62 NA NA NA 66.73 NA NA -15.25 NA
#> 2 12 7 2.98 83.64 0 11.95 0.52 66.2 9.46 -61.36 -83.64
#> RMSEA rms
#> 1 0.13 0.07
#> 2 0.00 0.01