Confidence intervals and profile likelihoods for parameters in cumulative link models
confintOld.RdComputes confidence intervals from the profiled likelihood for one or more parameters in a fitted cumulative link model, or plots the profile likelihood function.
Usage
# S3 method for class 'clm2'
confint(object, parm, level = 0.95, whichL = seq_len(p),
whichS = seq_len(k), lambda = TRUE, trace = 0, ...)
# S3 method for class 'profile.clm2'
confint(object, parm = seq_along(Pnames), level = 0.95, ...)
# S3 method for class 'clm2'
profile(fitted, whichL = seq_len(p), whichS = seq_len(k),
lambda = TRUE, alpha = 0.01, maxSteps = 50, delta = LrootMax/10,
trace = 0, stepWarn = 8, ...)
# S3 method for class 'profile.clm2'
plot(x, parm = seq_along(Pnames), level = c(0.95, 0.99),
Log = FALSE, relative = TRUE, fig = TRUE, n = 1e3, ..., ylim = NULL)Arguments
- object
a fitted
clm2object or aprofile.clm2object.- fitted
a fitted
clm2object.- x
a
profile.clm2object.- parm
not used in
confint.clm2.For
confint.profile.clm2: a specification of which parameters are to be given confidence intervals, either a vector of numbers or a vector of names. If missing, all parameters are considered.For
plot.profile.clm2: a specification of which parameters the profile likelihood are to be plotted for, either a vector of numbers or a vector of names. If missing, all parameters are considered.- level
the confidence level required.
- whichL
a specification of which location parameters are to be given confidence intervals, either a vector of numbers or a vector of names. If missing, all location parameters are considered.
- whichS
a specification of which scale parameters are to be given confidence intervals, either a vector of numbers or a vector of names. If missing, all scale parameters are considered.
- lambda
logical. Should profile or confidence intervals be computed for the link function parameter? Only used when one of the flexible link functions are used; see the
link-argument inclm2.- trace
logical. Should profiling be traced?
- alpha
Determines the range of profiling. By default the likelihood is profiled in the 99% confidence interval region as determined by the profile likelihood.
- maxSteps
the maximum number of profiling steps in each direction (up and down) for each parameter.
- delta
the length of profiling steps. To some extent this parameter determines the degree of accuracy of the profile likelihood in that smaller values, i.e. smaller steps gives a higher accuracy. Note however that a spline interpolation is used when constructing confidence intervals so fairly long steps can provide high accuracy.
- stepWarn
a warning is issued if the no. steps in each direction (up or down) for a parameter is less than
stepWarn(defaults to 8 steps) because this indicates an unreliable profile.- Log
should the profile likelihood be plotted on the log-scale?
- relative
should the relative or the absolute likelihood be plotted?
- fig
should the profile likelihood be plotted?
- n
the no. points used in the spline interpolation of the profile likelihood.
- ylim
overrules default y-limits on the plot of the profile likelihood.
- ...
additional argument(s) for methods including
range(for the hidden functionprofileLambda) that sets the range of values oflambdaat which the likelihood should be profiled for this parameter.
Value
confint:
A matrix (or vector) with columns giving lower and upper confidence
limits for each parameter. These will be labelled as (1-level)/2 and
1 - (1-level)/2 in % (by default 2.5% and 97.5%).
The parameter names are preceded with "loc." or "sca."
to indicate whether the confidence interval applies to a location or a
scale parameter.
plot.profile.clm2 invisibly returns the profile object.
Details
These confint methods call
the appropriate profile method, then finds the
confidence intervals by interpolation of the profile traces.
If the profile object is already available, this should be used as the
main argument rather than the fitted model object itself.
In plot.profile.clm2: at least one of Log and
relative arguments have to be TRUE.
Examples
options(contrasts = c("contr.treatment", "contr.poly"))
## More manageable data set:
(tab26 <- with(soup, table("Product" = PROD, "Response" = SURENESS)))
#> Response
#> Product 1 2 3 4 5 6
#> Ref 132 161 65 41 121 219
#> Test 96 99 50 57 156 650
dimnames(tab26)[[2]] <- c("Sure", "Not Sure", "Guess", "Guess", "Not Sure", "Sure")
dat26 <- expand.grid(sureness = as.factor(1:6), prod = c("Ref", "Test"))
dat26$wghts <- c(t(tab26))
m1 <- clm2(sureness ~ prod, scale = ~prod, data = dat26,
weights = wghts, link = "logistic")
## profile
pr1 <- profile(m1)
#> Error in eval(expr, p): object 'dat26' not found
par(mfrow = c(2, 2))
plot(pr1)
#> Error: object 'pr1' not found
m9 <- update(m1, link = "log-gamma")
pr9 <- profile(m9, whichL = numeric(0), whichS = numeric(0))
#> Error in eval(expr, p): object 'dat26' not found
par(mfrow = c(1, 1))
plot(pr9)
#> Error: object 'pr9' not found
plot(pr9, Log=TRUE, relative = TRUE)
#> Error: object 'pr9' not found
plot(pr9, Log=TRUE, relative = TRUE, ylim = c(-4, 0))
#> Error: object 'pr9' not found
plot(pr9, Log=TRUE, relative = FALSE)
#> Error: object 'pr9' not found
## confint
confint(pr9)
#> Error: object 'pr9' not found
confint(pr1)
#> Error: object 'pr1' not found
## Extend example from polr in package MASS:
## Fit model from polr example:
if(require(MASS)) {
fm1 <- clm2(Sat ~ Infl + Type + Cont, scale = ~ Cont, weights = Freq,
data = housing)
pr1 <- profile(fm1)
confint(pr1)
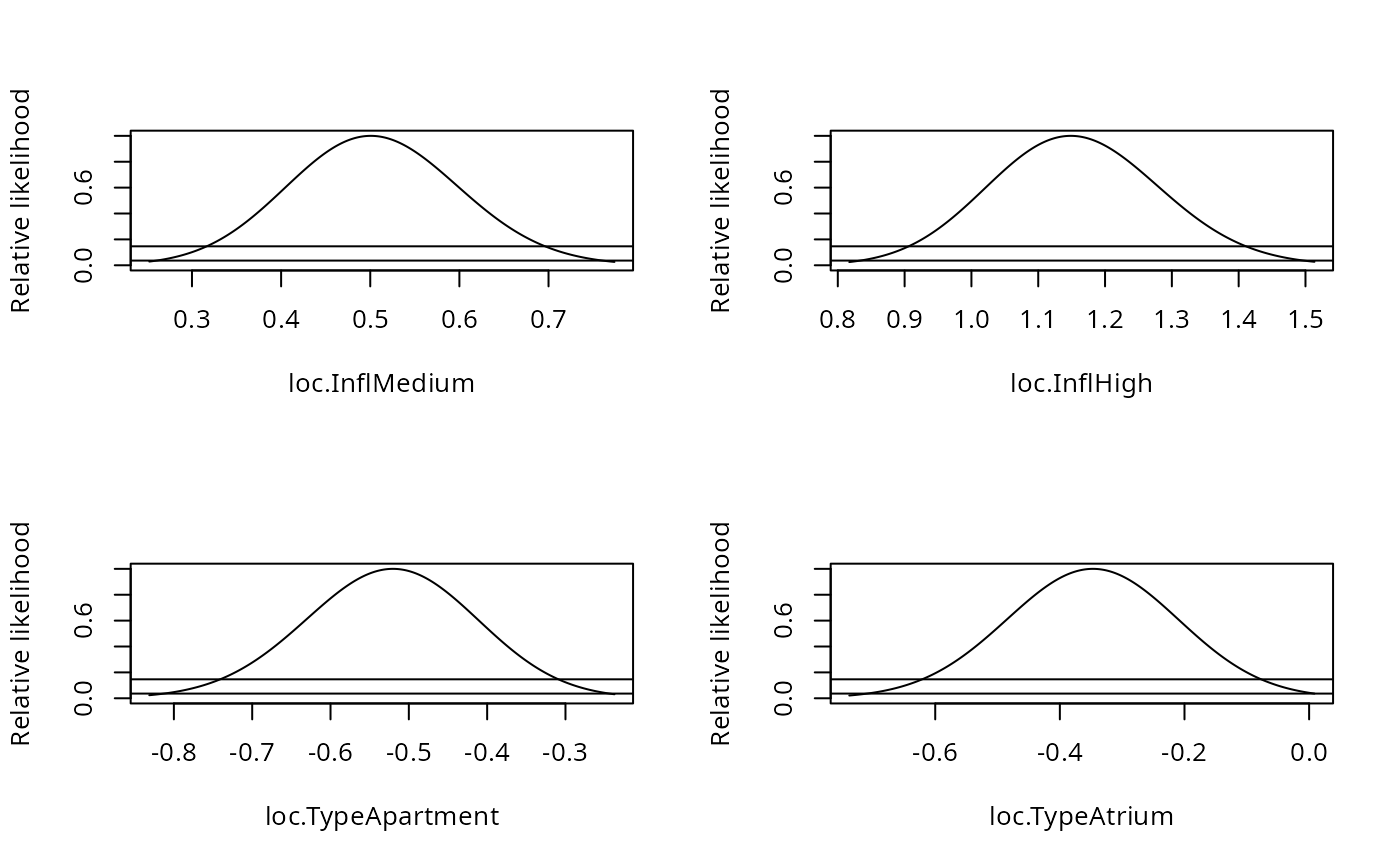
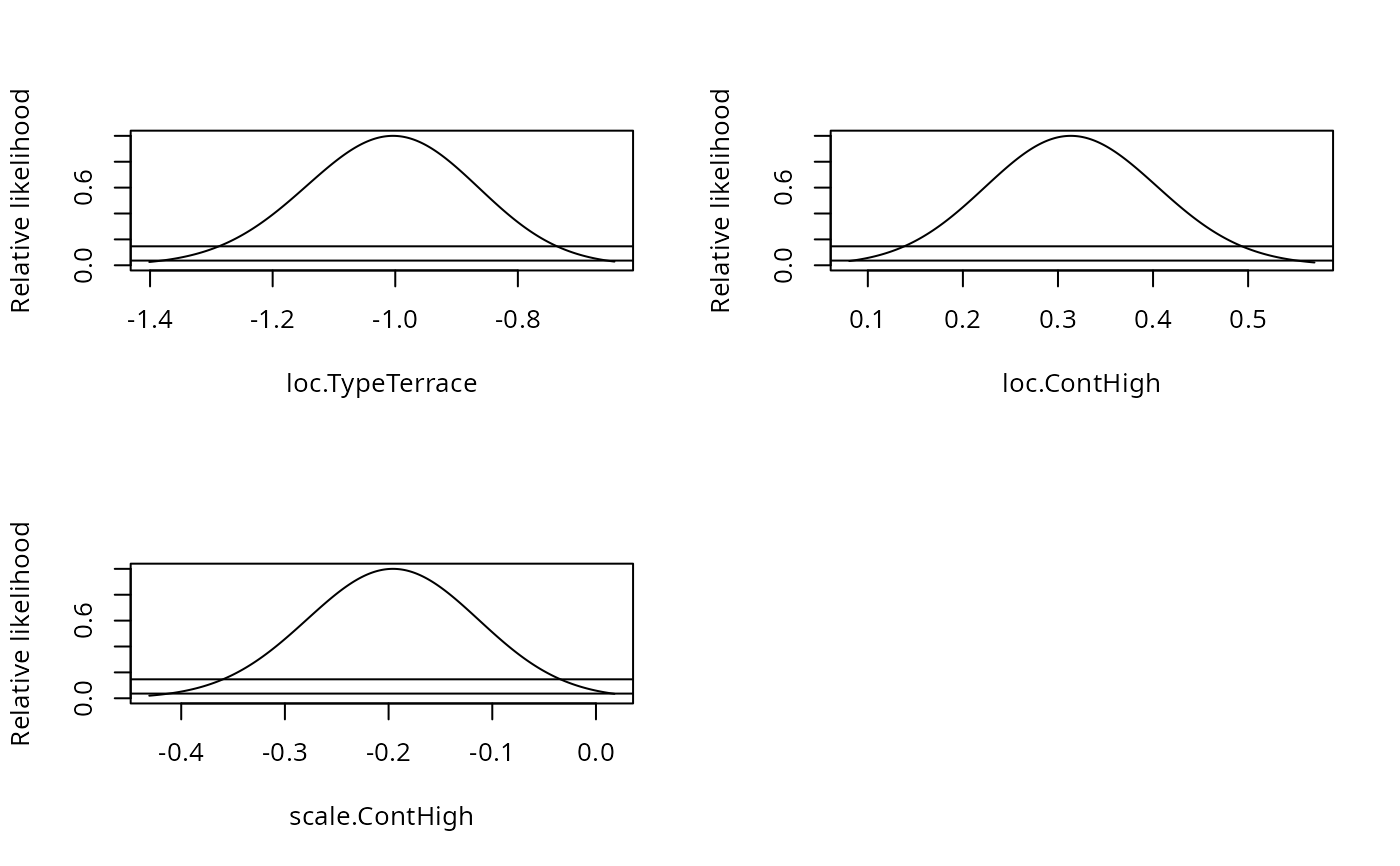
par(mfrow=c(2,2))
plot(pr1)
}