Methods for General Linear Hypotheses
methods.RdSimultaneous tests and confidence intervals for general linear hypotheses.
Usage
# S3 method for class 'glht'
summary(object, test = adjusted(), ...)
# S3 method for class 'glht'
confint(object, parm, level = 0.95, calpha = adjusted_calpha(),
...)
# S3 method for class 'glht'
coef(object, rhs = FALSE, ...)
# S3 method for class 'glht'
vcov(object, ...)
# S3 method for class 'confint.glht'
plot(x, xlim, xlab, ylim, ...)
# S3 method for class 'glht'
plot(x, ...)
univariate()
adjusted(type = c("single-step", "Shaffer", "Westfall", "free",
p.adjust.methods), ...)
Ftest()
Chisqtest()
adjusted_calpha(...)
univariate_calpha(...)Arguments
- object
an object of class
glht.- test
a function for computing p values.
- parm
additional parameters, currently ignored.
- level
the confidence level required.
- calpha
either a function computing the critical value or the critical value itself.
- rhs
logical, indicating whether the linear function \(K \hat{\theta}\) or the right hand side \(m\) (
rhs = TRUE) of the linear hypothesis should be returned.- type
the multiplicity adjustment (
adjusted) to be applied. See below andp.adjust.- x
an object of class
glhtorconfint.glht.- xlim
the
xlimits(x1, x2)of the plot.- ylim
the y limits of the plot.
- xlab
a label for the
xaxis.- ...
additional arguments, such as
maxpts,absepsorrelepstopmvnorminadjustedorqmvnorminconfint. Note that additional arguments specified tosummary,confint,coefandvcovmethods are currently ignored.
Details
The methods for general linear hypotheses as described by objects returned
by glht can be used to actually test the global
null hypothesis, each of the partial hypotheses and for
simultaneous confidence intervals for the linear function \(K \theta\).
The coef and vcov methods compute the linear
function \(K \hat{\theta}\) and its covariance, respectively.
The test argument to summary takes a function specifying
the type of test to be applied. Classical Chisq (Wald test) or F statistics
for testing the global hypothesis \(H_0\) are implemented in functions
Chisqtest and Ftest. Several approaches to multiplicity adjusted p
values for each of the linear hypotheses are implemented
in function adjusted. The type
argument to adjusted specifies the method to be applied:
"single-step" implements adjusted p values based on the joint
normal or t distribution of the linear function, and
"Shaffer" and "Westfall" implement logically constraint
multiplicity adjustments
multcomp::shaffer1986,multcomp::westfall1997.
"free" implements multiple testing procedures under free
combinations multcomp::westfall1999.
In addition, all adjustment methods
implemented in p.adjust are available as well.
Simultaneous confidence intervals for linear functions can be computed
using method confint. Univariate confidence intervals
can be computed by specifying calpha = univariate_calpha()
to confint. The critical value can directly be specified as a scalar
to calpha as well. Note that plot(a) for some object a of class
glht is equivalent to plot(confint(a)).
All simultaneous inference procedures implemented here control
the family-wise error rate (FWER). Multivariate
normal and t distributions, the latter one only for models of
class lm, are evaluated using the procedures
implemented in package mvtnorm. Note that the default
procedure is stochastic. Reproducible p-values and confidence
intervals require appropriate settings of seeds.
A more detailed description of the underlying methodology is available from hothorn2008b and Bretz+Hothorn+Westfall_2010.
Value
summary computes (adjusted) p values for general linear hypotheses,
confint computes (adjusted) confidence intervals.
coef returns estimates of the linear function \(K \theta\)
and vcov its covariance.
Examples
### set up a two-way ANOVA
amod <- aov(breaks ~ wool + tension, data = warpbreaks)
### set up all-pair comparisons for factor `tension'
wht <- glht(amod, linfct = mcp(tension = "Tukey"))
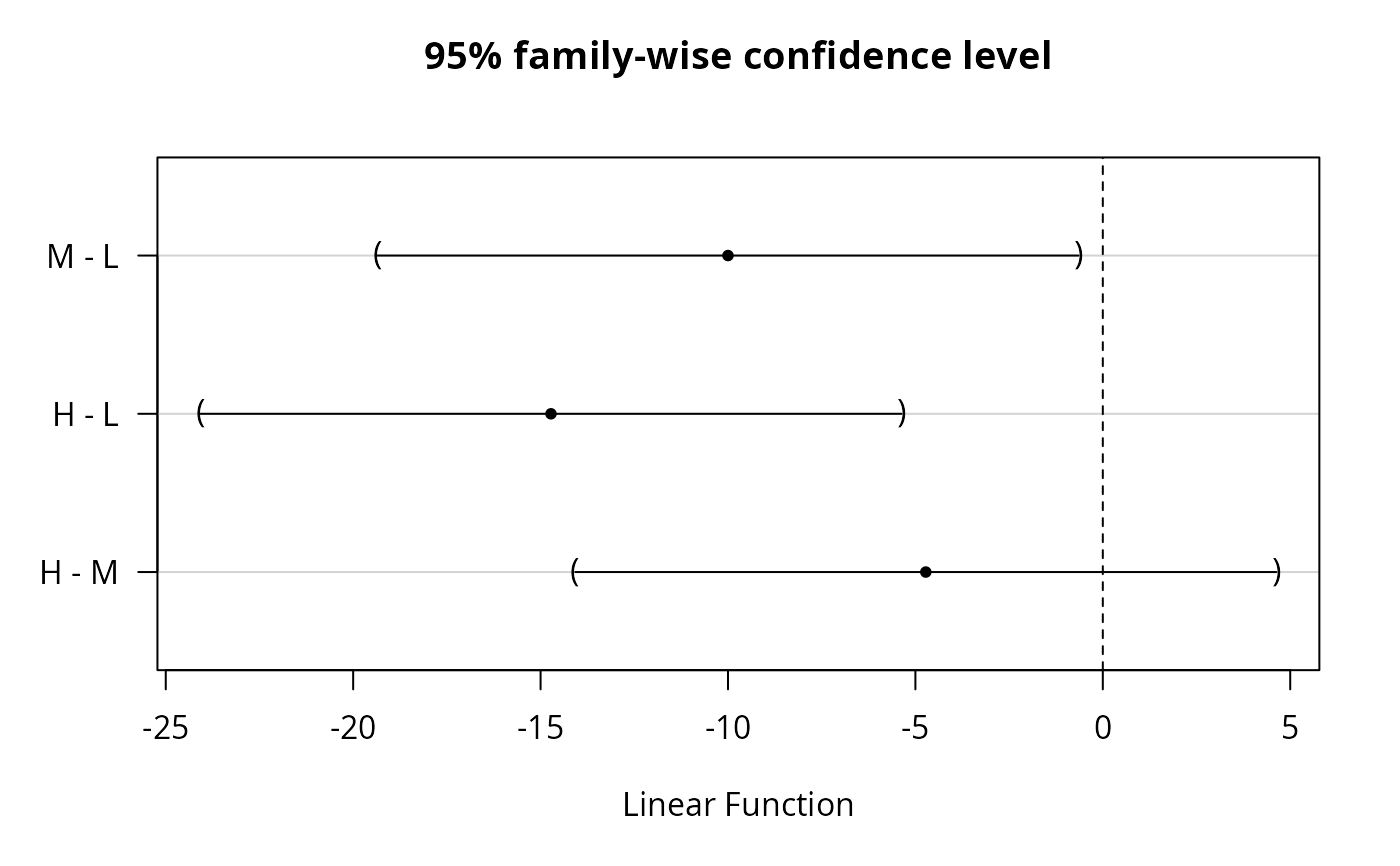
### 95% simultaneous confidence intervals
plot(print(confint(wht)))
#>
#> Simultaneous Confidence Intervals
#>
#> Multiple Comparisons of Means: Tukey Contrasts
#>
#>
#> Fit: aov(formula = breaks ~ wool + tension, data = warpbreaks)
#>
#> Quantile = 2.4149
#> 95% family-wise confidence level
#>
#>
#> Linear Hypotheses:
#> Estimate lwr upr
#> M - L == 0 -10.0000 -19.3515 -0.6485
#> H - L == 0 -14.7222 -24.0737 -5.3707
#> H - M == 0 -4.7222 -14.0737 4.6293
#>
 ### the same (for balanced designs only)
TukeyHSD(amod, "tension")
#> Tukey multiple comparisons of means
#> 95% family-wise confidence level
#>
#> Fit: aov(formula = breaks ~ wool + tension, data = warpbreaks)
#>
#> $tension
#> diff lwr upr p adj
#> M-L -10.000000 -19.35342 -0.6465793 0.0336262
#> H-L -14.722222 -24.07564 -5.3688015 0.0011218
#> H-M -4.722222 -14.07564 4.6311985 0.4474210
#>
### corresponding adjusted p values
summary(wht)
#>
#> Simultaneous Tests for General Linear Hypotheses
#>
#> Multiple Comparisons of Means: Tukey Contrasts
#>
#>
#> Fit: aov(formula = breaks ~ wool + tension, data = warpbreaks)
#>
#> Linear Hypotheses:
#> Estimate Std. Error t value Pr(>|t|)
#> M - L == 0 -10.000 3.872 -2.582 0.03356 *
#> H - L == 0 -14.722 3.872 -3.802 0.00114 **
#> H - M == 0 -4.722 3.872 -1.219 0.44742
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#> (Adjusted p values reported -- single-step method)
#>
### all means for levels of `tension'
amod <- aov(breaks ~ tension, data = warpbreaks)
glht(amod, linfct = matrix(c(1, 0, 0,
1, 1, 0,
1, 0, 1), byrow = TRUE, ncol = 3))
#>
#> General Linear Hypotheses
#>
#> Linear Hypotheses:
#> Estimate
#> 1 == 0 36.39
#> 2 == 0 26.39
#> 3 == 0 21.67
#>
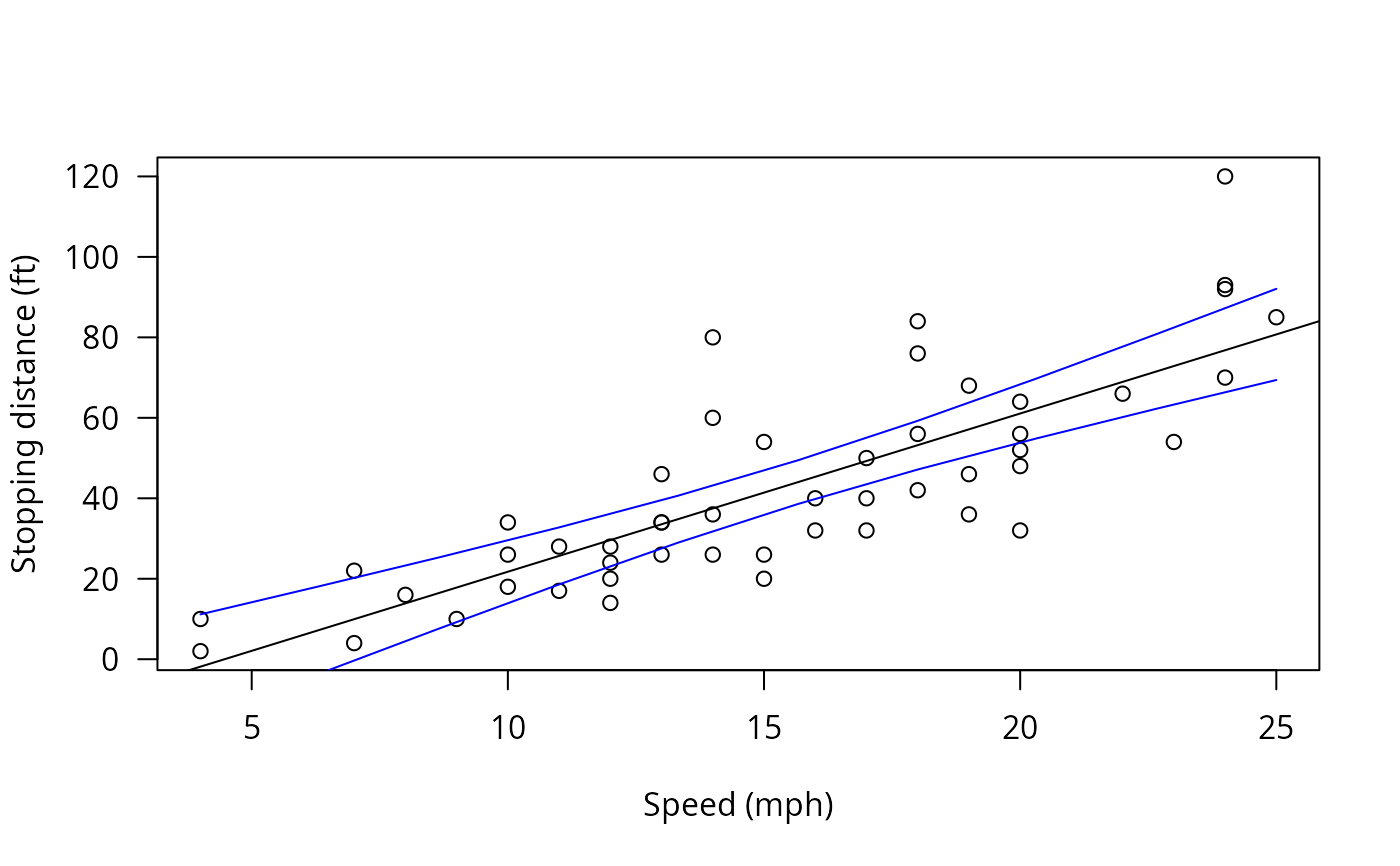
### confidence bands for a simple linear model, `cars' data
plot(cars, xlab = "Speed (mph)", ylab = "Stopping distance (ft)",
las = 1)
### fit linear model and add regression line to plot
lmod <- lm(dist ~ speed, data = cars)
abline(lmod)
### a grid of speeds
speeds <- seq(from = min(cars$speed), to = max(cars$speed),
length.out = 10)
### linear hypotheses: 10 selected points on the regression line != 0
K <- cbind(1, speeds)
### set up linear hypotheses
cht <- glht(lmod, linfct = K)
### confidence intervals, i.e., confidence bands, and add them plot
cci <- confint(cht)
lines(speeds, cci$confint[,"lwr"], col = "blue")
lines(speeds, cci$confint[,"upr"], col = "blue")
### the same (for balanced designs only)
TukeyHSD(amod, "tension")
#> Tukey multiple comparisons of means
#> 95% family-wise confidence level
#>
#> Fit: aov(formula = breaks ~ wool + tension, data = warpbreaks)
#>
#> $tension
#> diff lwr upr p adj
#> M-L -10.000000 -19.35342 -0.6465793 0.0336262
#> H-L -14.722222 -24.07564 -5.3688015 0.0011218
#> H-M -4.722222 -14.07564 4.6311985 0.4474210
#>
### corresponding adjusted p values
summary(wht)
#>
#> Simultaneous Tests for General Linear Hypotheses
#>
#> Multiple Comparisons of Means: Tukey Contrasts
#>
#>
#> Fit: aov(formula = breaks ~ wool + tension, data = warpbreaks)
#>
#> Linear Hypotheses:
#> Estimate Std. Error t value Pr(>|t|)
#> M - L == 0 -10.000 3.872 -2.582 0.03356 *
#> H - L == 0 -14.722 3.872 -3.802 0.00114 **
#> H - M == 0 -4.722 3.872 -1.219 0.44742
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#> (Adjusted p values reported -- single-step method)
#>
### all means for levels of `tension'
amod <- aov(breaks ~ tension, data = warpbreaks)
glht(amod, linfct = matrix(c(1, 0, 0,
1, 1, 0,
1, 0, 1), byrow = TRUE, ncol = 3))
#>
#> General Linear Hypotheses
#>
#> Linear Hypotheses:
#> Estimate
#> 1 == 0 36.39
#> 2 == 0 26.39
#> 3 == 0 21.67
#>
### confidence bands for a simple linear model, `cars' data
plot(cars, xlab = "Speed (mph)", ylab = "Stopping distance (ft)",
las = 1)
### fit linear model and add regression line to plot
lmod <- lm(dist ~ speed, data = cars)
abline(lmod)
### a grid of speeds
speeds <- seq(from = min(cars$speed), to = max(cars$speed),
length.out = 10)
### linear hypotheses: 10 selected points on the regression line != 0
K <- cbind(1, speeds)
### set up linear hypotheses
cht <- glht(lmod, linfct = K)
### confidence intervals, i.e., confidence bands, and add them plot
cci <- confint(cht)
lines(speeds, cci$confint[,"lwr"], col = "blue")
lines(speeds, cci$confint[,"upr"], col = "blue")
 ### simultaneous p values for parameters in a Cox model
if (require("survival") && require("MASS")) {
data("leuk", package = "MASS")
leuk.cox <- coxph(Surv(time) ~ ag + log(wbc), data = leuk)
### set up linear hypotheses
lht <- glht(leuk.cox, linfct = diag(length(coef(leuk.cox))))
### adjusted p values
print(summary(lht))
}
#>
#> Simultaneous Tests for General Linear Hypotheses
#>
#> Fit: coxph(formula = Surv(time) ~ ag + log(wbc), data = leuk)
#>
#> Linear Hypotheses:
#> Estimate Std. Error z value Pr(>|z|)
#> 1 == 0 -1.0691 0.4293 -2.490 0.0253 *
#> 2 == 0 0.3677 0.1360 2.703 0.0137 *
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#> (Adjusted p values reported -- single-step method)
#>
### simultaneous p values for parameters in a Cox model
if (require("survival") && require("MASS")) {
data("leuk", package = "MASS")
leuk.cox <- coxph(Surv(time) ~ ag + log(wbc), data = leuk)
### set up linear hypotheses
lht <- glht(leuk.cox, linfct = diag(length(coef(leuk.cox))))
### adjusted p values
print(summary(lht))
}
#>
#> Simultaneous Tests for General Linear Hypotheses
#>
#> Fit: coxph(formula = Surv(time) ~ ag + log(wbc), data = leuk)
#>
#> Linear Hypotheses:
#> Estimate Std. Error z value Pr(>|z|)
#> 1 == 0 -1.0691 0.4293 -2.490 0.0253 *
#> 2 == 0 0.3677 0.1360 2.703 0.0137 *
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#> (Adjusted p values reported -- single-step method)
#>