plot Method for logistf Likelihood Profiles
Source: R/plot.logistf.profile.r
plot.logistf.profile.RdProvides the plot method for objects created by profile.logistf or CLIP.profile
# S3 method for class 'logistf.profile'
plot(
x,
type = "profile",
max1 = TRUE,
colmain = "black",
colimp = "gray",
plotmain = T,
ylim = NULL,
...
)Arguments
- x
A
profile.logistfobject- type
Type of plot: one of c("profile", "cdf", "density")
- max1
If
type="density", normalizes density to maximum 1- colmain
Color for main profile line
- colimp
color for completed-data profile lines (for
logistf.profileobjects that also carry theCLIP.profileclass attribute)- plotmain
if
FALSE, suppresses the main profile line (forlogistf.profileobjects that also carry theCLIP.profileclass attribute)- ylim
Limits for the y-axis
- ...
Further arguments to be passed to
plot().
Value
The function is called for its side effects
Details
The plot method provides three types of plots (profile, CDF, and density representation of a profile likelihood). For objects generated by CLIP.profile, it also allows to show the completed-data profiles along with the pooled profile.
References
Heinze G, Ploner M, Beyea J (2013). Confidence intervals after multiple imputation: combining profile likelihood information from logistic regressions. Statistics in Medicine, to appear.
Examples
data(sex2)
fit<-logistf(case ~ age+oc+vic+vicl+vis+dia, data=sex2)
plot(profile(fit,variable="dia"))
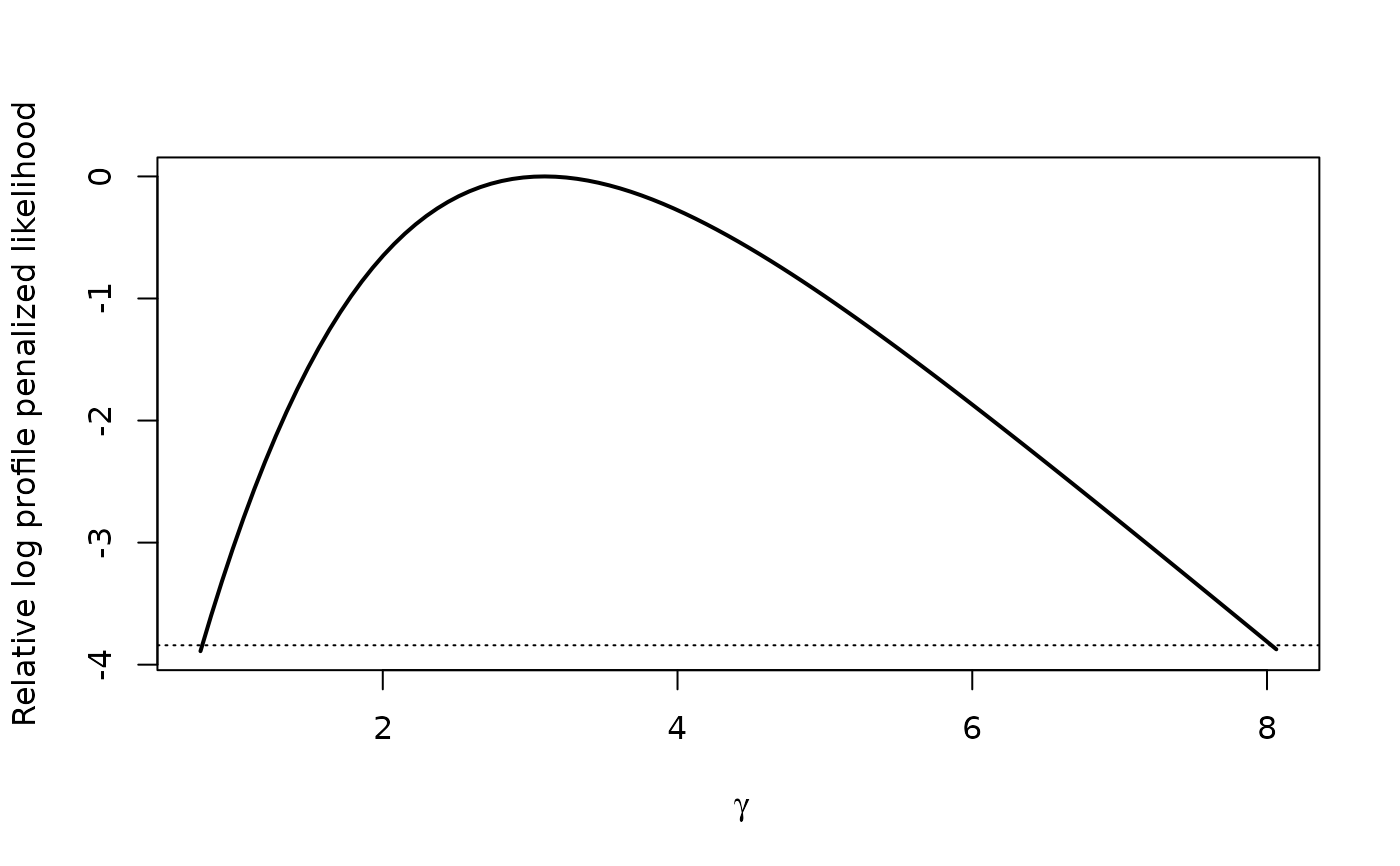 plot(profile(fit,variable="dia"), "cdf")
plot(profile(fit,variable="dia"), "cdf")
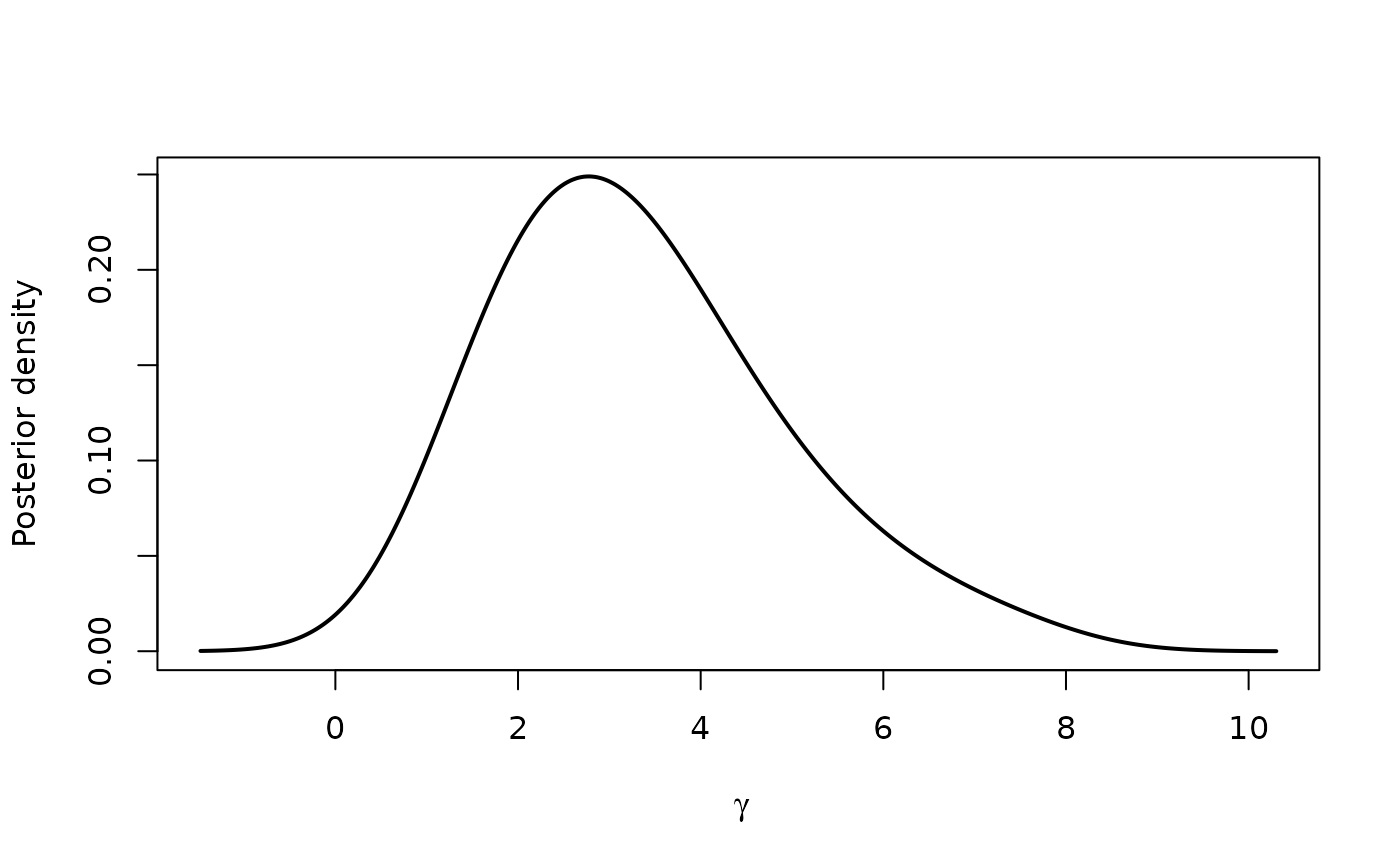
 plot(profile(fit,variable="dia"), "density")
#> Warning: Selecting bandwidth *not* using 'weights'
plot(profile(fit,variable="dia"), "density")
#> Warning: Selecting bandwidth *not* using 'weights'
 #generate data set with NAs
freq=c(5,2,2,7,5,4)
y<-c(rep(1,freq[1]+freq[2]), rep(0,freq[3]+freq[4]), rep(1,freq[5]), rep(0,freq[6]))
x<-c(rep(1,freq[1]), rep(0,freq[2]), rep(1,freq[3]), rep(0,freq[4]), rep(NA,freq[5]),
rep(NA,freq[6]))
toy<-data.frame(x=x,y=y)
# impute data set 5 times
set.seed(169)
toymi<-list(0)
for(i in 1:5){
toymi[[i]]<-toy
y1<-toymi[[i]]$y==1 & is.na(toymi[[i]]$x)
y0<-toymi[[i]]$y==0 & is.na(toymi[[i]]$x)
xnew1<-rbinom(sum(y1),1,freq[1]/(freq[1]+freq[2]))
xnew0<-rbinom(sum(y0),1,freq[3]/(freq[3]+freq[4]))
toymi[[i]]$x[y1==TRUE]<-xnew1
toymi[[i]]$x[y0==TRUE]<-xnew0
}
# logistf analyses of each imputed data set
fit.list<-lapply(1:5, function(X) logistf(data=toymi[[X]], y~x, pl=TRUE))
# CLIP profile
xprof<-CLIP.profile(obj=fit.list, variable="x", data=toymi, keep=TRUE)
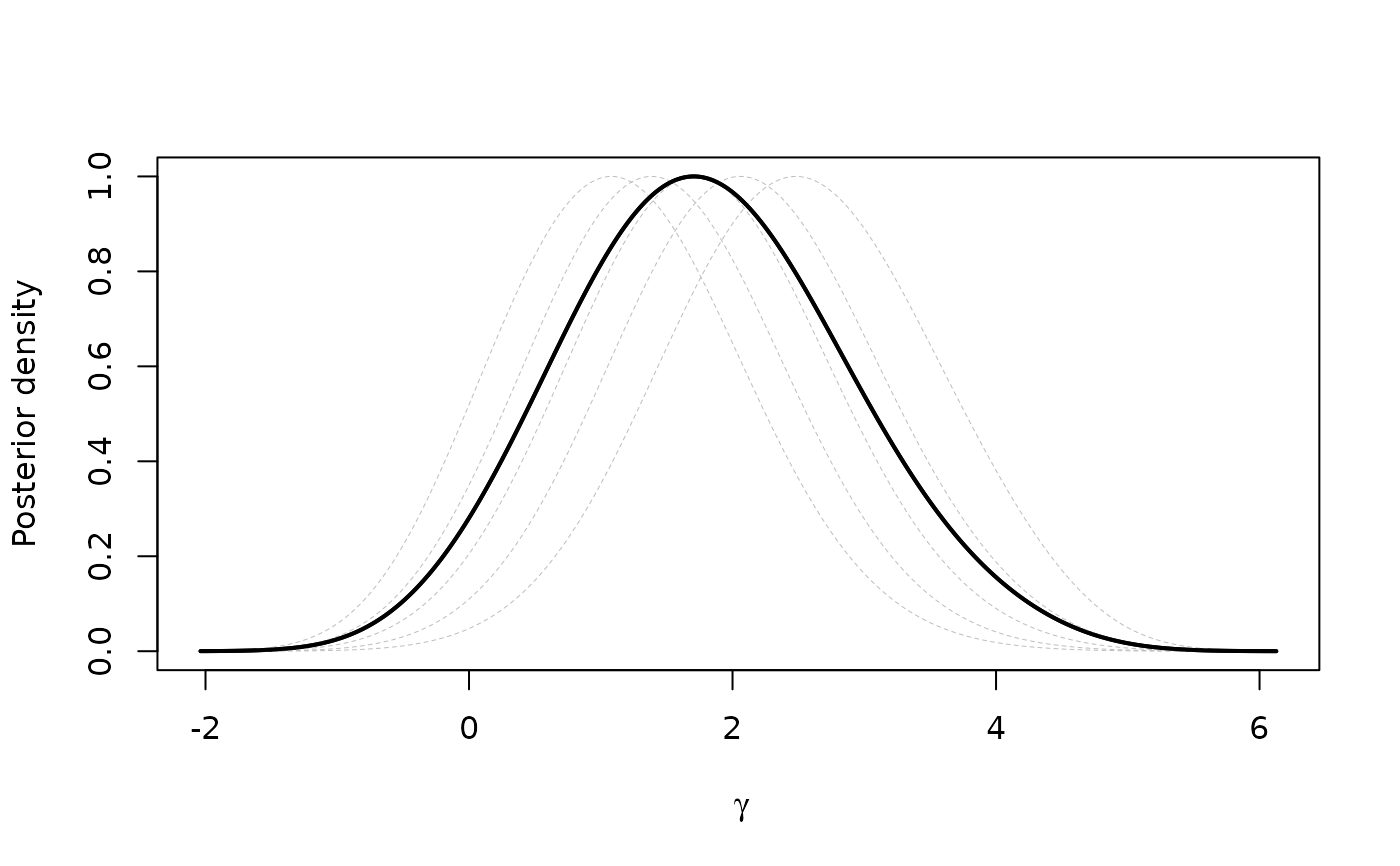
plot(xprof)
#generate data set with NAs
freq=c(5,2,2,7,5,4)
y<-c(rep(1,freq[1]+freq[2]), rep(0,freq[3]+freq[4]), rep(1,freq[5]), rep(0,freq[6]))
x<-c(rep(1,freq[1]), rep(0,freq[2]), rep(1,freq[3]), rep(0,freq[4]), rep(NA,freq[5]),
rep(NA,freq[6]))
toy<-data.frame(x=x,y=y)
# impute data set 5 times
set.seed(169)
toymi<-list(0)
for(i in 1:5){
toymi[[i]]<-toy
y1<-toymi[[i]]$y==1 & is.na(toymi[[i]]$x)
y0<-toymi[[i]]$y==0 & is.na(toymi[[i]]$x)
xnew1<-rbinom(sum(y1),1,freq[1]/(freq[1]+freq[2]))
xnew0<-rbinom(sum(y0),1,freq[3]/(freq[3]+freq[4]))
toymi[[i]]$x[y1==TRUE]<-xnew1
toymi[[i]]$x[y0==TRUE]<-xnew0
}
# logistf analyses of each imputed data set
fit.list<-lapply(1:5, function(X) logistf(data=toymi[[X]], y~x, pl=TRUE))
# CLIP profile
xprof<-CLIP.profile(obj=fit.list, variable="x", data=toymi, keep=TRUE)
plot(xprof)
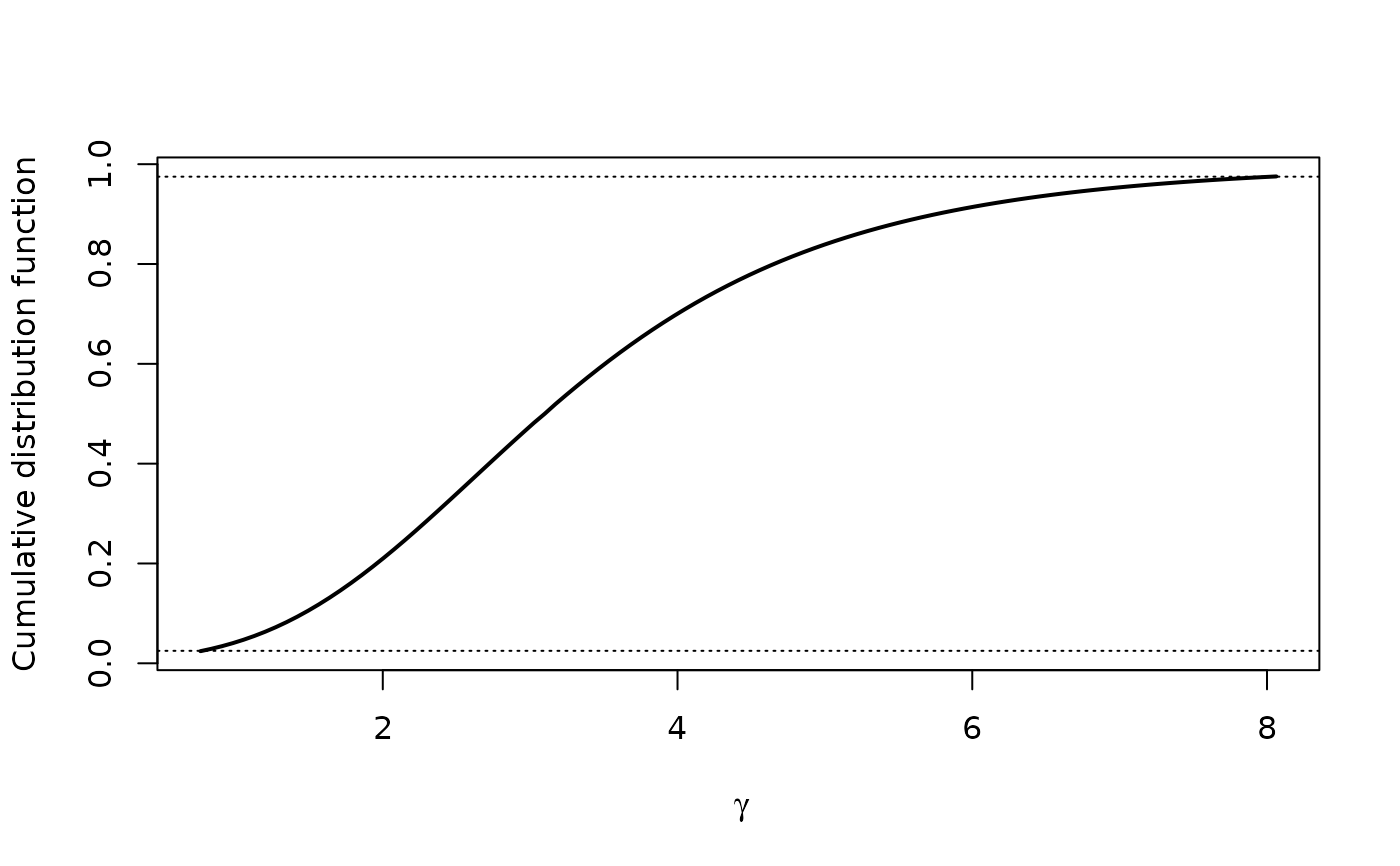
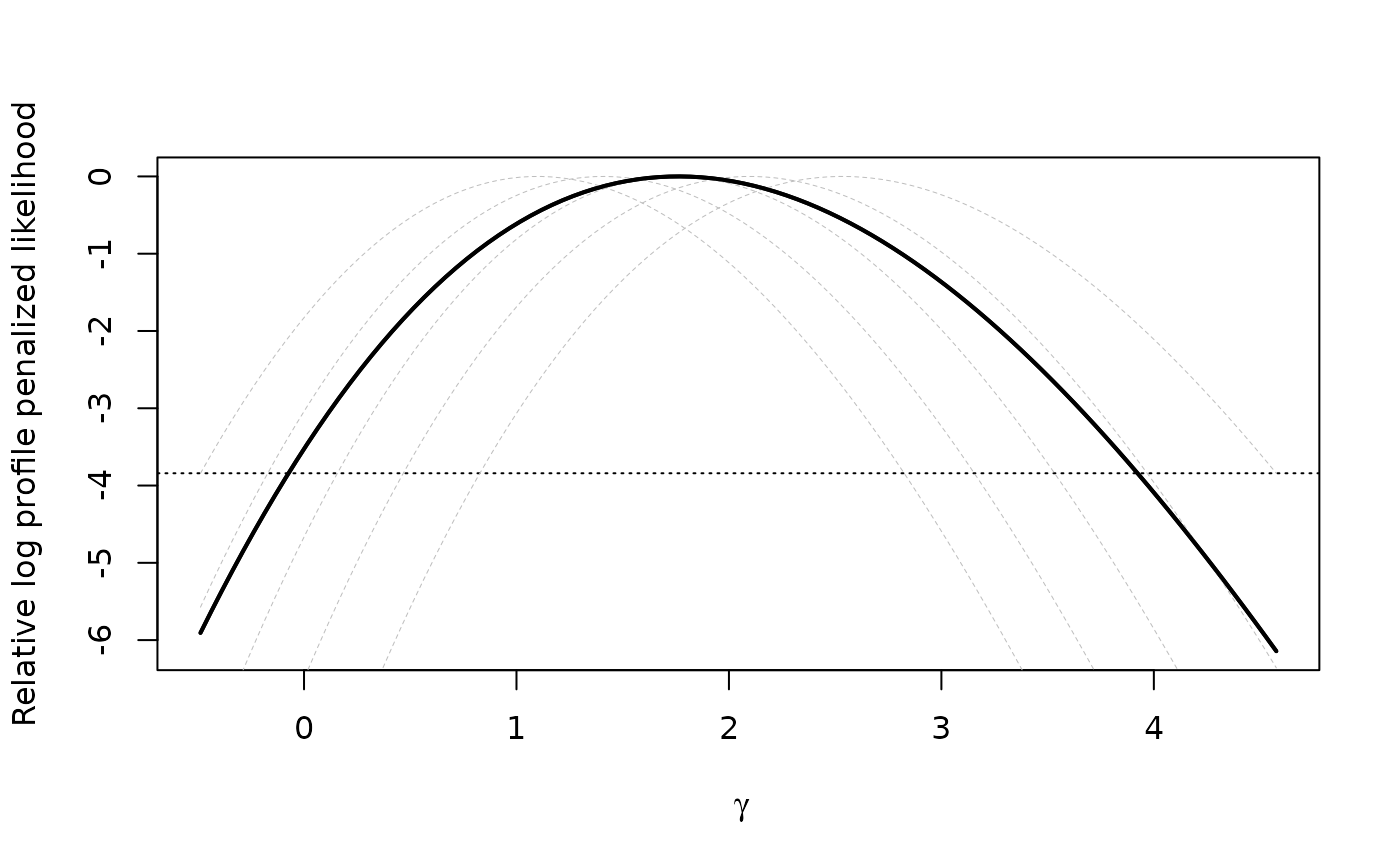 #plot as CDF
plot(xprof, "cdf")
#plot as CDF
plot(xprof, "cdf")
 #plot as density
plot(xprof, "density")
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#plot as density
plot(xprof, "density")
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'
#> Warning: Selecting bandwidth *not* using 'weights'