Model Fit for Survival Data
sbrier.RdModel fit for survival data: the integrated Brier score for censored observations.
sbrier(obj, pred, btime= range(obj[,1]))Arguments
Details
There is no obvious criterion of model fit for censored data. The Brier score for censoring as well as it's integrated version were suggested by Graf et al (1999).
The integrated Brier score is always computed over a subset of the
interval given by the range of the time slot of the survival object obj.
Value
The (integrated) Brier score with attribute time is returned.
See also
More measures for the validation of predicted surival probabilities
are implemented in package pec.
References
Erika Graf, Claudia Schmoor, Willi Sauerbrei and Martin Schumacher (1999), Assessment and comparison of prognostic classification schemes for survival data. Statistics in Medicine 18(17-18), 2529–2545.
Examples
library("survival")
data("DLBCL", package = "ipred")
smod <- Surv(DLBCL$time, DLBCL$cens)
KM <- survfit(smod ~ 1)
# integrated Brier score up to max(DLBCL$time)
sbrier(smod, KM)
#> [,1]
#> [1,] 0.2237226
#> attr(,"names")
#> [1] "integrated Brier score"
#> attr(,"time")
#> [1] 1.3 129.9
# integrated Brier score up to time=50
sbrier(smod, KM, btime=c(0, 50))
#> Warning: btime[1] is smaller than min(time)
#> [,1]
#> [1,] 0.2174081
#> attr(,"names")
#> [1] "integrated Brier score"
#> attr(,"time")
#> [1] 1.3 39.6
# Brier score for time=50
sbrier(smod, KM, btime=50)
#> Brier score
#> 0.249375
#> attr(,"time")
#> [1] 50
# a "real" model: one single survival tree with Intern. Prognostic Index
# and mean gene expression in the first cluster as predictors
mod <- bagging(Surv(time, cens) ~ MGEc.1 + IPI, data=DLBCL, nbagg=1)
# this is a list of survfit objects (==KM-curves), one for each observation
# in DLBCL
pred <- predict(mod, newdata=DLBCL)
# integrated Brier score up to max(time)
sbrier(smod, pred)
#> [,1]
#> [1,] 0.1442559
#> attr(,"names")
#> [1] "integrated Brier score"
#> attr(,"time")
#> [1] 1.3 129.9
# Brier score at time=50
sbrier(smod, pred, btime=50)
#> Brier score
#> 0.1774478
#> attr(,"time")
#> [1] 50
# artificial examples and illustrations
cleans <- function(x) { attr(x, "time") <- NULL; names(x) <- NULL; x }
n <- 100
time <- rpois(n, 20)
cens <- rep(1, n)
# checks, Graf et al. page 2536, no censoring at all!
# no information: \pi(t) = 0.5
a <- sbrier(Surv(time, cens), rep(0.5, n), time[50])
stopifnot(all.equal(cleans(a),0.25))
# some information: \pi(t) = S(t)
n <- 100
time <- 1:100
mod <- survfit(Surv(time, cens) ~ 1)
a <- sbrier(Surv(time, cens), rep(list(mod), n))
mymin <- mod$surv * (1 - mod$surv)
cleans(a)
#> [,1]
#> [1,] 0.1682833
sum(mymin)/diff(range(time))
#> [1] 0.1683333
# independent of ordering
rand <- sample(1:100)
b <- sbrier(Surv(time, cens)[rand], rep(list(mod), n)[rand])
stopifnot(all.equal(cleans(a), cleans(b)))
# 2 groups at different risk
time <- c(1:10, 21:30)
strata <- c(rep(1, 10), rep(2, 10))
cens <- rep(1, length(time))
# no information about the groups
a <- sbrier(Surv(time, cens), survfit(Surv(time, cens) ~ 1))
b <- sbrier(Surv(time, cens), rep(list(survfit(Surv(time, cens) ~1)), 20))
stopifnot(all.equal(a, b))
# risk groups known
mod <- survfit(Surv(time, cens) ~ strata)
b <- sbrier(Surv(time, cens), c(rep(list(mod[1]), 10), rep(list(mod[2]), 10)))
stopifnot(a > b)
### GBSG2 data
data("GBSG2", package = "TH.data")
thsum <- function(x) {
ret <- c(median(x), quantile(x, 0.25), quantile(x,0.75))
names(ret)[1] <- "Median"
ret
}
t(apply(GBSG2[,c("age", "tsize", "pnodes",
"progrec", "estrec")], 2, thsum))
#> Median 25% 75%
#> age 53.0 46 61.00
#> tsize 25.0 20 35.00
#> pnodes 3.0 1 7.00
#> progrec 32.5 7 131.75
#> estrec 36.0 8 114.00
table(GBSG2$menostat)
#>
#> Pre Post
#> 290 396
table(GBSG2$tgrade)
#>
#> I II III
#> 81 444 161
table(GBSG2$horTh)
#>
#> no yes
#> 440 246
# pooled Kaplan-Meier
mod <- survfit(Surv(time, cens) ~ 1, data=GBSG2)
# integrated Brier score
sbrier(Surv(GBSG2$time, GBSG2$cens), mod)
#> [,1]
#> [1,] 0.1939366
#> attr(,"names")
#> [1] "integrated Brier score"
#> attr(,"time")
#> [1] 8 2659
# Brier score at 5 years
sbrier(Surv(GBSG2$time, GBSG2$cens), mod, btime=1825)
#> Brier score
#> 0.2499984
#> attr(,"time")
#> [1] 1825
# Nottingham prognostic index
GBSG2 <- GBSG2[order(GBSG2$time),]
NPI <- 0.2*GBSG2$tsize/10 + 1 + as.integer(GBSG2$tgrade)
NPI[NPI < 3.4] <- 1
NPI[NPI >= 3.4 & NPI <=5.4] <- 2
NPI[NPI > 5.4] <- 3
mod <- survfit(Surv(time, cens) ~ NPI, data=GBSG2)
plot(mod)
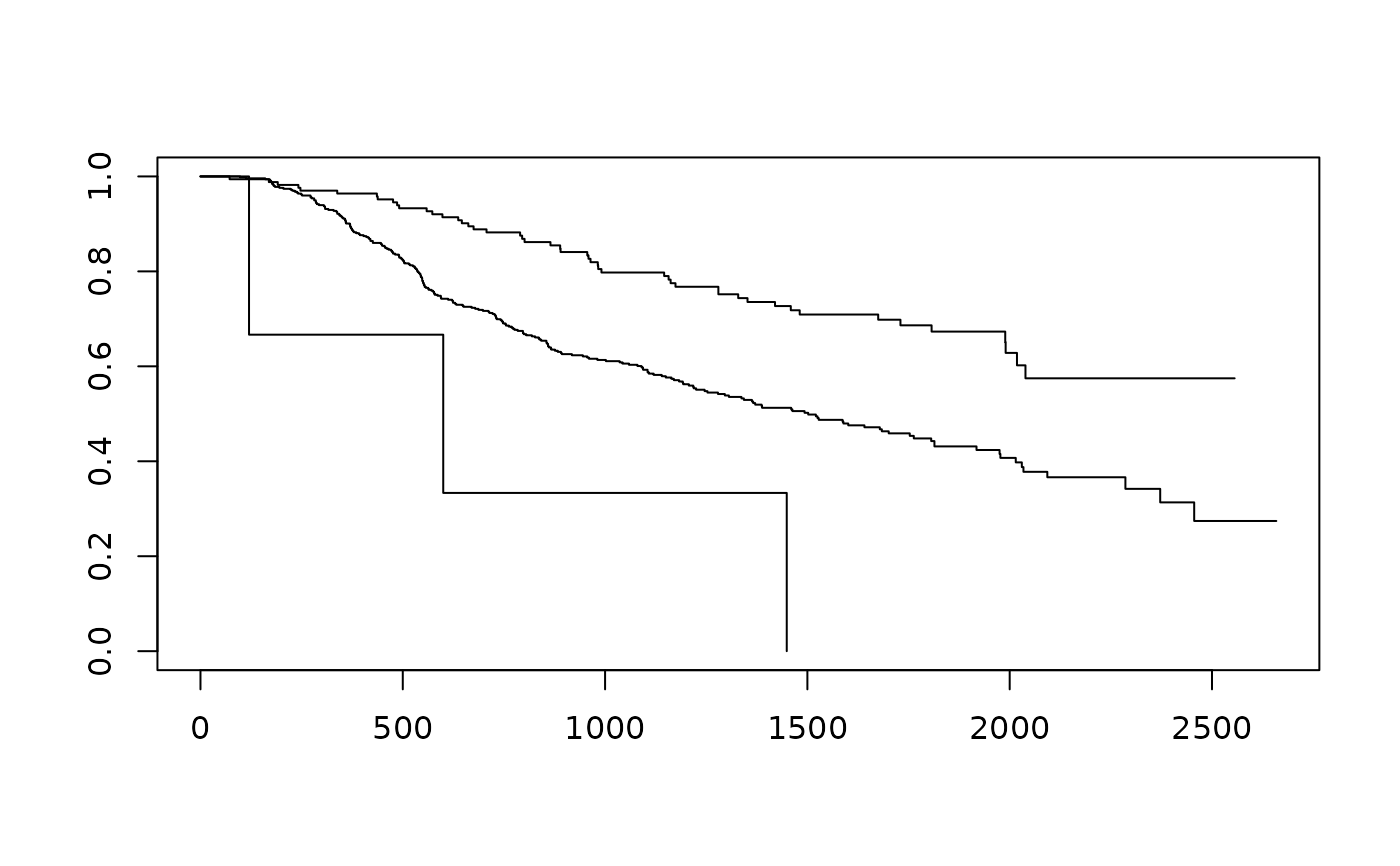 pred <- c()
survs <- c()
for (i in sort(unique(NPI)))
survs <- c(survs, getsurv(mod[i], 1825))
for (i in 1:nrow(GBSG2))
pred <- c(pred, survs[NPI[i]])
# Brier score of NPI at t=5 years
sbrier(Surv(GBSG2$time, GBSG2$cens), pred, btime=1825)
#> Brier score
#> 0.233823
#> attr(,"time")
#> [1] 1825
pred <- c()
survs <- c()
for (i in sort(unique(NPI)))
survs <- c(survs, getsurv(mod[i], 1825))
for (i in 1:nrow(GBSG2))
pred <- c(pred, survs[NPI[i]])
# Brier score of NPI at t=5 years
sbrier(Surv(GBSG2$time, GBSG2$cens), pred, btime=1825)
#> Brier score
#> 0.233823
#> attr(,"time")
#> [1] 1825