Fitting Generalized Additive Models
gam.Rdgam is used to fit generalized additive models, specified by giving a
symbolic description of the additive predictor and a description of the
error distribution. gam uses the backfitting algorithm to
combine different smoothing or fitting methods. The methods currently
supported are local regression and smoothing splines.
Usage
gam(
formula,
family = gaussian,
data,
weights,
subset,
na.action,
start = NULL,
etastart,
mustart,
control = gam.control(...),
model = TRUE,
method = "glm.fit",
x = FALSE,
y = TRUE,
...
)
gam.fit(
x,
y,
smooth.frame,
weights = rep(1, nobs),
start = NULL,
etastart = NULL,
mustart = NULL,
offset = rep(0, nobs),
family = gaussian(),
control = gam.control()
)Arguments
- formula
a formula expression as for other regression models, of the form
response ~ predictors. See the documentation oflmandformulafor details. Built-in nonparametric smoothing terms are indicated bysfor smoothing splines orloforloesssmooth terms. See the documentation forsandlofor their arguments. Additional smoothers can be added by creating the appropriate interface functions. Interactions with nonparametric smooth terms are not fully supported, but will not produce errors; they will simply produce the usual parametric interaction.- family
a description of the error distribution and link function to be used in the model. This can be a character string naming a family function, a family function or the result of a call to a family function. (See
familyfor details of family functions.)- data
an optional data frame containing the variables in the model. If not found in
data, the variables are taken fromenvironment(formula), typically the environment from whichgamis called.- weights
an optional vector of weights to be used in the fitting process.
- subset
an optional vector specifying a subset of observations to be used in the fitting process.
- na.action
a function which indicates what should happen when the data contain
NAs. The default is set by thena.actionsetting ofoptions, and isna.failif that is unset. The “factory-fresh” default isna.omit. A special methodna.gam.replaceallows for mean-imputation of missing values (assumes missing at random), and works gracefully withgam- start
starting values for the parameters in the additive predictor.
- etastart
starting values for the additive predictor.
- mustart
starting values for the vector of means.
- control
a list of parameters for controlling the fitting process. See the documentation for
gam.controlfor details. These can also be set as arguments togam()itself.- model
a logical value indicating whether model frame should be included as a component of the returned value. Needed if
gamis called and predicted from inside a user function. Default isTRUE.- method
the method to be used in fitting the parametric part of the model. The default method
"glm.fit"uses iteratively reweighted least squares (IWLS). The only current alternative is"model.frame"which returns the model frame and does no fitting.- x, y
For
gam: logical values indicating whether the response vector and model matrix used in the fitting process should be returned as components of the returned value.For
gam.fit:xis a model matrix of dimensionn * p, andyis a vector of observations of lengthn.- ...
further arguments passed to or from other methods.
- smooth.frame
for
gam.fitonly. This is essentially a subset of the model frame corresponding to the smooth terms, and has the ingredients needed for smoothing each variable in the backfitting algorithm. The elements of this frame are produced by the formula functionsloands.- offset
this can be used to specify an a priori known component to be included in the additive predictor during fitting.
Value
gam returns an object of class Gam, which inherits
from both glm and lm.
Gam objects can be examined by print, summary, plot,
and anova. Components can be extracted using extractor functions
predict, fitted, residuals, deviance,
formula, and family. Can be modified using update. It
has all the components of a glm object, with a few more. This also
means it can be queried, summarized etc by methods for glm and
lm objects. Other generic functions that have methods for Gam
objects are step and preplot.
The following components must be included in a legitimate `Gam' object. The
residuals, fitted values, coefficients and effects should be extracted by
the generic functions of the same name, rather than by the "$"
operator. The family function returns the entire family object used
in the fitting, and deviance can be used to extract the deviance of
the fit.
- coefficients
the coefficients of the parametric part of the
additive.predictors, which multiply the columns of the model matrix. The names of the coefficients are the names of the single-degree-of-freedom effects (the columns of the model matrix). If the model is overdetermined there will be missing values in the coefficients corresponding to inestimable coefficients.- additive.predictors
the additive fit, given by the product of the model matrix and the coefficients, plus the columns of the
$smoothcomponent.- fitted.values
the fitted mean values, obtained by transforming the component
additive.predictorsusing the inverse link function.- smooth, nl.df, nl.chisq, var
these four characterize the nonparametric aspect of the fit.
smoothis a matrix of smooth terms, with a column corresponding to each smooth term in the model; if no smooth terms are in theGammodel, all these components will be missing. Each column corresponds to the strictly nonparametric part of the term, while the parametric part is obtained from the model matrix.nl.dfis a vector giving the approximate degrees of freedom for each column ofsmooth. For smoothing splines specified bys(x), the approximatedfwill be the trace of the implicit smoother matrix minus 2.nl.chisqis a vector containing a type of score test for the removal of each of the columns ofsmooth.varis a matrix likesmooth, containing the approximate pointwise variances for the columns ofsmooth.- smooth.frame
This is essentially a subset of the model frame corresponding to the smooth terms, and has the ingredients needed for making predictions from a
Gamobject- residuals
the residuals from the final weighted additive fit; also known as residuals, these are typically not interpretable without rescaling by the weights.
- deviance
up to a constant, minus twice the maximized log-likelihood. Similar to the residual sum of squares. Where sensible, the constant is chosen so that a saturated model has deviance zero.
- null.deviance
The deviance for the null model, comparable with
deviance. The null model will include the offset, and an intercept if there is one in the model- iter
the number of local scoring iterations used to compute the estimates.
- bf.iter
a vector of length
itergiving number of backfitting iterations used at each inner loop.- family
a three-element character vector giving the name of the family, the link, and the variance function; mainly for printing purposes.
- weights
the working weights, that is the weights in the final iteration of the local scoring fit.
- prior.weights
the case weights initially supplied.
- df.residual
the residual degrees of freedom.
- df.null
the residual degrees of freedom for the null model.
The object will also have the components of a lm object:
coefficients, residuals, fitted.values, call,
terms, and some others involving the numerical fit. See
lm.object.
Details
The gam model is fit using the local scoring algorithm, which iteratively
fits weighted additive models by backfitting. The backfitting algorithm is a
Gauss-Seidel method for fitting additive models, by iteratively smoothing
partial residuals. The algorithm separates the parametric from the
nonparametric part of the fit, and fits the parametric part using weighted
linear least squares within the backfitting algorithm. This version of
gam remains faithful to the philosophy of GAM models as outlined in
the references below.
An object gam.slist (currently set to c("lo","s","random"))
lists the smoothers supported by gam. Corresponding to each of these
is a smoothing function gam.lo, gam.s etc that take particular
arguments and produce particular output, custom built to serve as building
blocks in the backfitting algorithm. This allows users to add their own
smoothing methods. See the documentation for these methods for further
information. In addition, the object gam.wlist (currently set to
c("s","lo")) lists the smoothers for which efficient backfitters are
provided. These are invoked if all the smoothing methods are of one kind
(either all "lo" or all "s").
References
Hastie, T. J. (1991) Generalized additive models. Chapter 7 of Statistical Models in S eds J. M. Chambers and T. J. Hastie, Wadsworth & Brooks/Cole.
Hastie, T. and Tibshirani, R. (1990) Generalized Additive Models. London: Chapman and Hall.
Venables, W. N. and Ripley, B. D. (2002) Modern Applied Statistics with S. New York: Springer.
Author
Written by Trevor Hastie, following closely the design in the
"Generalized Additive Models" chapter (Hastie, 1992) in Chambers and Hastie
(1992), and the philosophy in Hastie and Tibshirani (1991). This version of
gam is adapted from the S version to match the glm and
lm functions in R.
Note that this version of gam is different from the function with the
same name in the R library mgcv, which uses only smoothing splines
with a focus on automatic smoothing parameter selection via GCV. To avoid
issues with S3 method handling when both packages are loaded, the object
class in package "gam" is now "Gam".
Examples
data(kyphosis)
gam(Kyphosis ~ s(Age,4) + Number, family = binomial, data=kyphosis,
trace=TRUE)
#> GAM s.wam loop 1: deviance = 66.42095
#> GAM s.wam loop 2: deviance = 63.77252
#> GAM s.wam loop 3: deviance = 63.25199
#> GAM s.wam loop 4: deviance = 63.13399
#> GAM s.wam loop 5: deviance = 63.11016
#> GAM s.wam loop 6: deviance = 63.10748
#> GAM s.wam loop 7: deviance = 63.10727
#> GAM s.wam loop 8: deviance = 63.10725
#> GAM s.wam loop 9: deviance = 63.10725
#> Call:
#> gam(formula = Kyphosis ~ s(Age, 4) + Number, family = binomial,
#> data = kyphosis, trace = TRUE)
#>
#> Degrees of Freedom: 80 total; 75.00002 Residual
#> Residual Deviance: 63.10725
data(airquality)
gam(Ozone^(1/3) ~ lo(Solar.R) + lo(Wind, Temp), data=airquality, na=na.gam.replace)
#> Warning: 37 observations omitted due to missing values in the response
#> Call:
#> gam(formula = Ozone^(1/3) ~ lo(Solar.R) + lo(Wind, Temp), data = airquality,
#> na.action = na.gam.replace)
#>
#> Degrees of Freedom: 115 total; 102.6679 Residual
#> Residual Deviance: 20.05229
gam(Kyphosis ~ poly(Age,2) + s(Start), data=kyphosis, family=binomial, subset=Number>2)
#> Call:
#> gam(formula = Kyphosis ~ poly(Age, 2) + s(Start), family = binomial,
#> data = kyphosis, subset = Number > 2)
#>
#> Degrees of Freedom: 68 total; 61.99976 Residual
#> Residual Deviance: 44.5635
data(gam.data)
Gam.object <- gam(y ~ s(x,6) + z,data=gam.data)
summary(Gam.object)
#>
#> Call: gam(formula = y ~ s(x, 6) + z, data = gam.data)
#> Deviance Residuals:
#> Min 1Q Median 3Q Max
#> -0.59177 -0.18050 0.01291 0.23941 0.51412
#>
#> (Dispersion Parameter for gaussian family taken to be 0.0823)
#>
#> Null Deviance: 57.7496 on 99 degrees of freedom
#> Residual Deviance: 7.5745 on 92.0003 degrees of freedom
#> AIC: 43.7495
#>
#> Number of Local Scoring Iterations: NA
#>
#> Anova for Parametric Effects
#> Df Sum Sq Mean Sq F value Pr(>F)
#> s(x, 6) 1 38.210 38.210 464.0987 <2e-16 ***
#> z 1 0.084 0.084 1.0247 0.3141
#> Residuals 92 7.575 0.082
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
#>
#> Anova for Nonparametric Effects
#> Npar Df Npar F Pr(F)
#> (Intercept)
#> s(x, 6) 5 28.697 < 2.2e-16 ***
#> z
#> ---
#> Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
plot(Gam.object,se=TRUE)
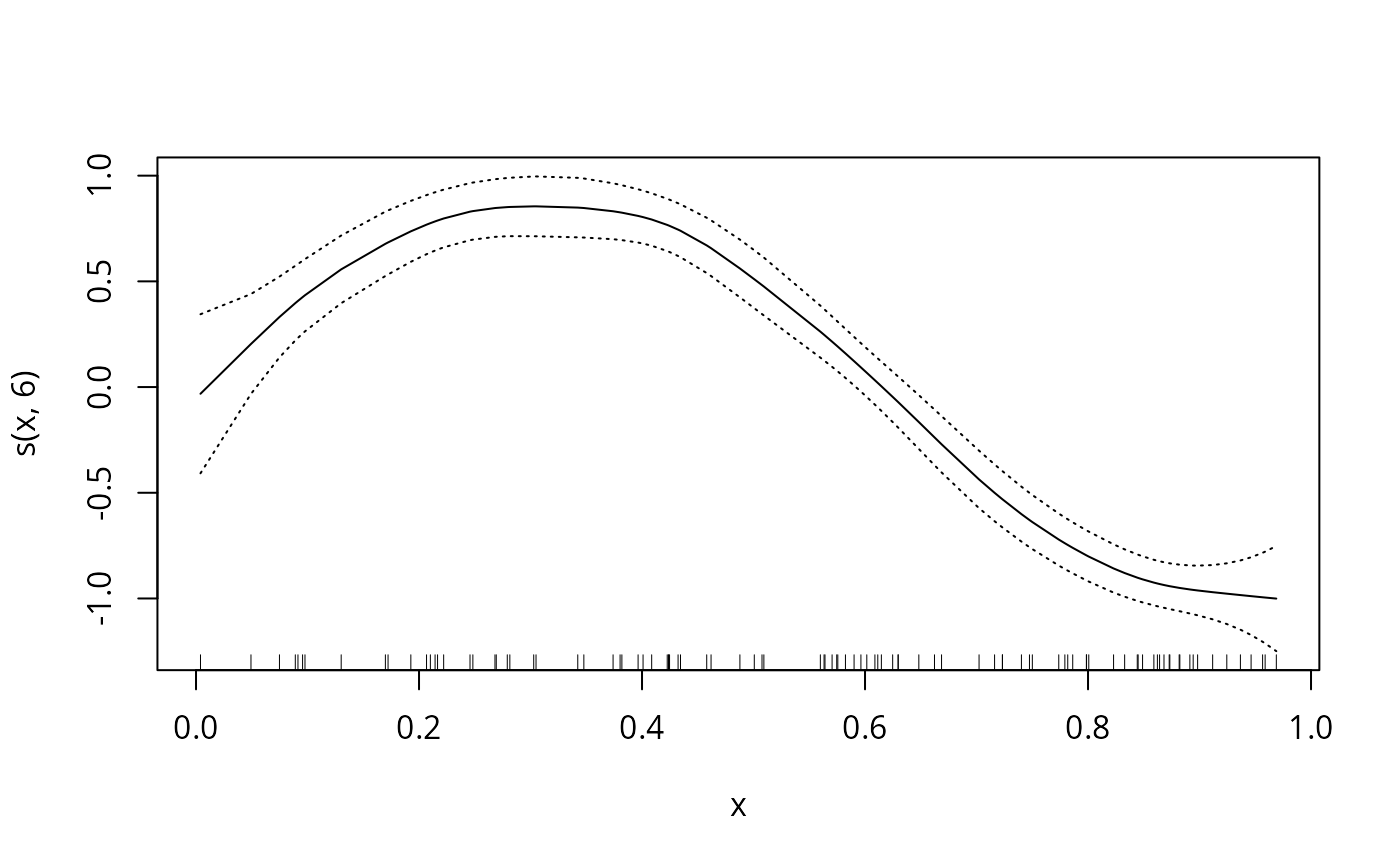
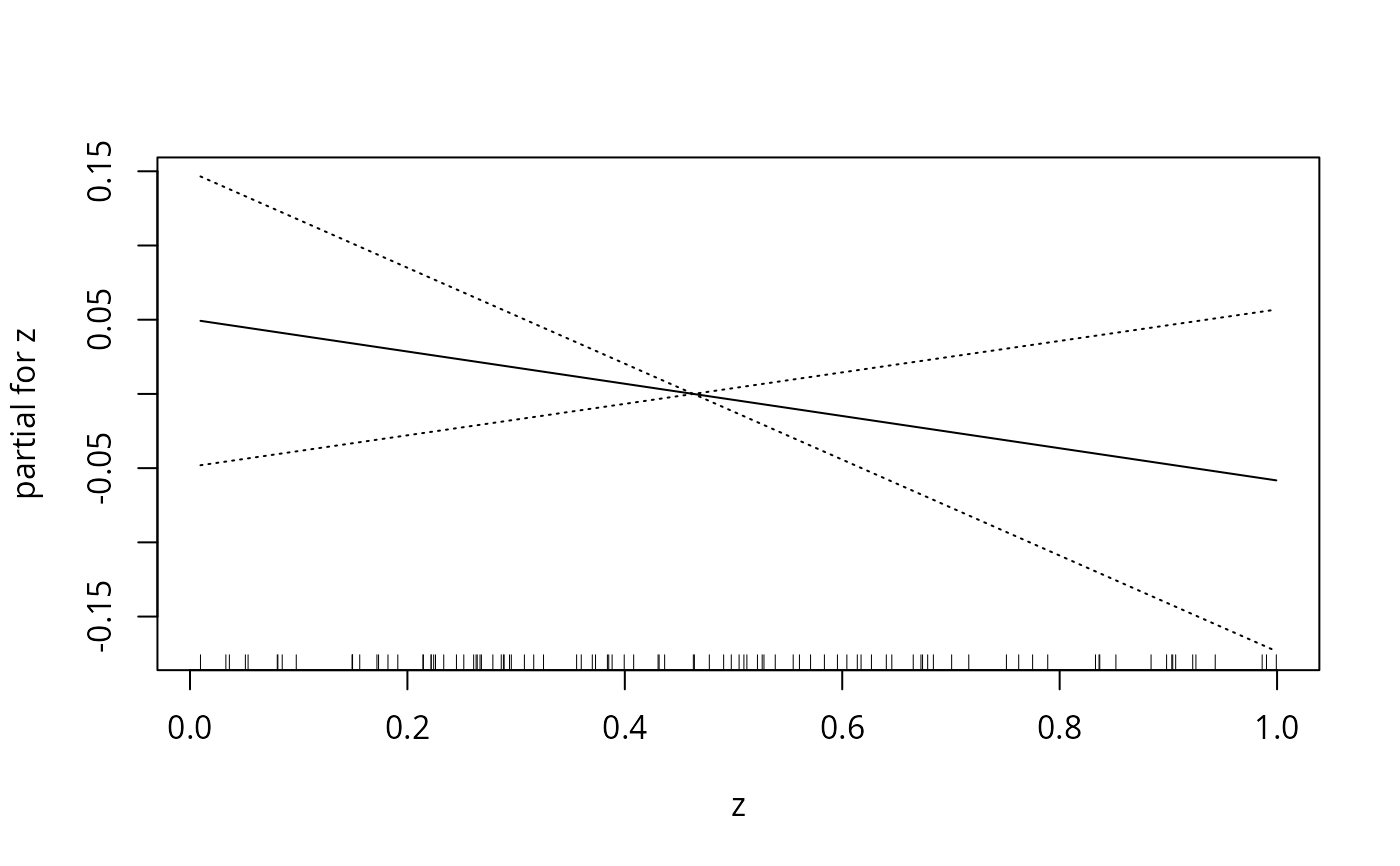 data(gam.newdata)
predict(Gam.object,type="terms",newdata=gam.newdata)
#> s(x, 6) z
#> 1 0.44379143 0.039428080
#> 2 0.75360796 0.028567775
#> 3 0.85468111 0.017707471
#> 4 0.80512984 0.006847166
#> 5 0.51279094 -0.004013139
#> 6 0.07523574 -0.014873443
#> 7 -0.42622549 -0.025733748
#> 8 -0.80005872 -0.036594052
#> 9 -0.96321574 -0.047454357
#> 10 -1.01594919 -0.058314661
#> attr(,"constant")
#> [1] 0.634896
data(gam.newdata)
predict(Gam.object,type="terms",newdata=gam.newdata)
#> s(x, 6) z
#> 1 0.44379143 0.039428080
#> 2 0.75360796 0.028567775
#> 3 0.85468111 0.017707471
#> 4 0.80512984 0.006847166
#> 5 0.51279094 -0.004013139
#> 6 0.07523574 -0.014873443
#> 7 -0.42622549 -0.025733748
#> 8 -0.80005872 -0.036594052
#> 9 -0.96321574 -0.047454357
#> 10 -1.01594919 -0.058314661
#> attr(,"constant")
#> [1] 0.634896