Distribution of One and Two Sample Permutation Tests
dperm.RdDensity, distribution function and quantile function for the distribution of one and two sample permutation tests using the Shift-Algorithm by Streitberg & R\"ohmel.
Usage
dperm(x, scores, m, paired=NULL, tol = 0.01, fact=NULL, density=FALSE,
simulate=FALSE, B=10000)
pperm(q, scores, m, paired=NULL, tol = 0.01, fact=NULL,
alternative=c("less", "greater", "two.sided"), pprob=FALSE,
simulate=FALSE, B=10000)
qperm(p, scores, m, paired=NULL, tol = 0.01, fact=NULL,
simulate=FALSE, B=10000)
rperm(n, scores, m)Arguments
- x, q
vector of quantiles.
- p
vector of probabilities.
- scores
arbitrary scores of the observations of the
x(firstmelements) andysample.- m
sample size of the
xsample. Ifm = length(x)scores of paired observations are assumed.- paired
logical. Indicates if paired observations are used. Needed to discriminate between a paired problem and the distribution of the total sum of the scores (which has mass 1 at the point
sum(scores)).
.
- tol
real. Real valued scores are mapped into integers by rounding after multiplication with an appropriate factor. Make sure that the absolute difference between the each possible test statistic for the original scores and the rounded scores is less than
tol. This might not be possible due to memory/time limitations, a warning is given in this case.- fact
real. If
factis given, real valued scores are mapped into integers usingfactas factor.tolis ignored in this case.- n
number of random observations to generate.
- alternative
character indicating whether the probability \(P(T \le q)\) (
less), \(P(T \ge q)\) (greater) or a two-sided p-value (two.sided) should be computed inpperm.- pprob
logical. Indicates if the probability \(P(T = q)\) should be computed additionally.
- density
logical. When
xis a scalar anddensityisTRUE,dpermreturns the density for all possible statistics less or equalxas a data frame.- simulate
logical. Use conditional Monte-Carlo to compute the distribution.
- B
number of Monte-Carlo replications to be used.
Details
The exact distribution of the sum of the first m scores is
evaluated using the Shift-Algorithm by Streitberg & R\"ohmel under the
hypothesis of exchangeability (or, equivalent, the hypothesis that all
permutations of the scores are equally likely). The algorithm is able
to deal with tied scores, so the conditional distribution can be
evaluated.
The algorithm is defined for positive integer valued scores only.
There are two ways dealing with real valued scores.
First, one can try to find integer valued scores that lead to statistics
which differ not more than tol
from the statistics computed for the original scores. This can be done as
follows.
Without loss of generality let \(a_i > 0\) denote real valued scores in
reverse ordering and
\(f\) a positive factor (this is the fact argument).
Let \(R_i = f \cdot a_i - round(f \cdot a_i)\). Then
$$ \sum_{i=1}^m f \cdot a_i = \sum_{i=1}^m round(f \cdot a_i) - R_i. $$
Clearly, the maximum difference between \(1/f \sum_{i=1}^m f \cdot a_i\) and \(1/f \sum_{i=1}^n round(f \cdot a_i)\) is given by \(|\sum_{i=1}^m R_i|\). Therefore one searches for \(f\) with
$$ |\sum_{i=1}^m R_i| \le \sum_{i=1}^m |R_i| \le tol.$$
If \(f\) induces more that 100.000 columns in the Shift-Algorithm by Streitberg & R\"ohmel, \(f\) is restricted to the largest integer that does not.
The second idea is to map the scores into integers by taking the
integer part of \(a_i N / \max(a_i)\) (Hothorn & Lausen, 2002).
This induces additional ties, but the shape of the
scores is very similar. That means we do not try to approximate something
but use a different test (with integer valued scores), serving for the
same purpose
(due to a similar shape of the scores). However, this has to be done prior
to calling pperm (see the examples).
Exact two-sided p-values are computed as suggested in the StatXact-5
manual, page 225, equation (9.31) and equation (8.18), p. 179 (paired case).
In detail: For the paired case the two-sided p-value is just twice the
one-sided one. For the independent sample case the two sided p-value is
defined as
$$p_2 = P( |T - E(T)| \ge | q - E(T) |)$$
where \(q\) is the quantile passed to pperm.
Value
dperm gives the density, pperm gives the distribution
function and qperm gives the quantile function. If pprob is
true, pperm returns a list with elements
- PVALUE
the probability specified by
alternative.- PPROB
the probability \(P(T = q)\).
rperm is a wrapper to sample.
References
Bernd Streitberg & Joachim R\"ohmel (1986), Exact distributions for permutations and rank tests: An introduction to some recently published algorithms. Statistical Software Newsletter 12(1), 10–17.
Bernd Streitberg & Joachim R\"ohmel (1987), Exakte Verteilungen f\"ur Rang- und Randomisierungstests im allgemeinen $c$-Stichprobenfall. EDV in Medizin und Biologie 18(1), 12–19.
Torsten Hothorn (2001), On exact rank tests in R. R News 1(1), 11–12.
Cyrus R. Mehta & Nitin R. Patel (2001), StatXact-5 for Windows. Manual, Cytel Software Cooperation, Cambridge, USA
Torsten Hothorn & Berthold Lausen (2003), On the exact distribution of maximally selected rank statistics. Computational Statistics & Data Analysis, 43(2), 121-137.
Examples
# exact one-sided p-value of the Wilcoxon test for a tied sample
x <- c(0.5, 0.5, 0.6, 0.6, 0.7, 0.8, 0.9)
y <- c(0.5, 1.0, 1.2, 1.2, 1.4, 1.5, 1.9, 2.0)
r <- cscores(c(x,y), type="Wilcoxon")
pperm(sum(r[seq(along=x)]), r, 7)
#> [1] 0.004351204
# Compare the exact algorithm as implemented in ctest and the
# Shift-Algorithm by Streitberg & Roehmel for untied samples
# Wilcoxon:
n <- 10
x <- rnorm(n, 2)
y <- rnorm(n, 3)
r <- cscores(c(x,y), type="Wilcoxon")
# exact distribution using the Shift-Algorithm
dwexac <- dperm((n*(n+1)/2):(n^2 + n*(n+1)/2), r, n)
sum(dwexac) # should be something near 1 :-)
#> [1] 1
# exact distribution using dwilcox
dw <- dwilcox(0:(n^2), n, n)
# compare the two distributions:
plot(dw, dwexac, main="Wilcoxon", xlab="dwilcox", ylab="dperm")
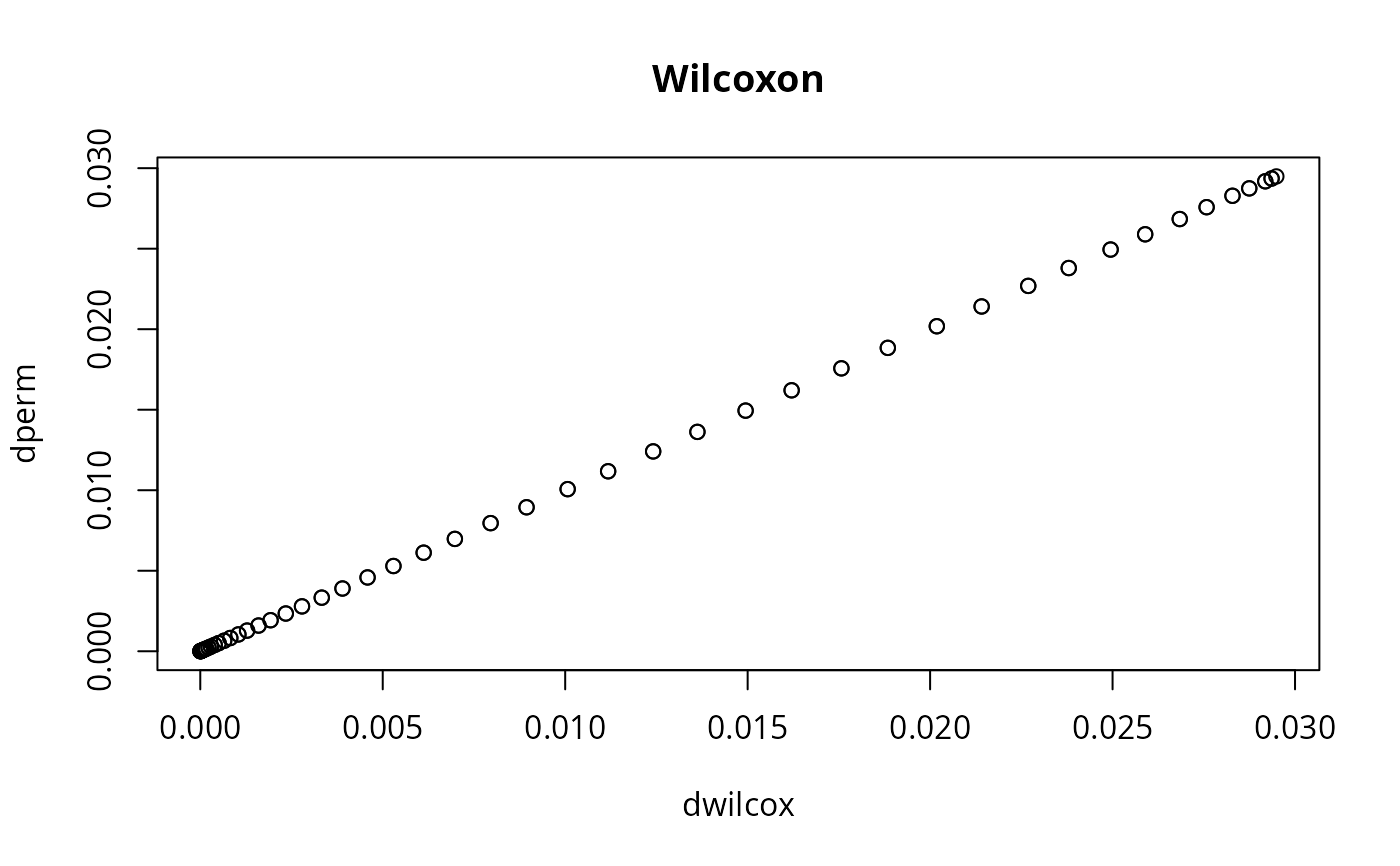 # should give a "perfect" line
# Wilcoxon signed rank test
n <- 10
x <- rnorm(n, 5)
y <- rnorm(n, 5)
r <- cscores(abs(x - y), type="Wilcoxon")
pperm(sum(r[x - y > 0]), r, length(r))
#> [1] 0.7216797
wilcox.test(x,y, paired=TRUE, alternative="less")
#>
#> Wilcoxon signed rank exact test
#>
#> data: x and y
#> V = 33, p-value = 0.7217
#> alternative hypothesis: true location shift is less than 0
#>
psignrank(sum(r[x - y > 0]), length(r))
#> [1] 0.7216797
# Ansari-Bradley
n <- 10
x <- rnorm(n, 2, 1)
y <- rnorm(n, 2, 2)
# exact distribution using the Shift-Algorithm
sc <- cscores(c(x,y), type="Ansari")
dabexac <- dperm(0:(n*(2*n+1)/2), sc, n)
sum(dabexac)
#> [1] 1
# real scores are allowed (but only result in an approximation)
# e.g. v.d. Waerden test
n <- 10
x <- rnorm(n)
y <- rnorm(n)
scores <- cscores(c(x,y), type="NormalQuantile")
X <- sum(scores[seq(along=x)]) # <- v.d. Waerden normal quantile statistic
# critical value, two-sided test
abs(qperm(0.025, scores, length(x)))
#> Warning: cannot hold tol, tolerance: 0.010292
#> [1] 3.871345
# p-values
p1 <- pperm(X, scores, length(x), alternative="two.sided")
#> Warning: cannot hold tol, tolerance: 0.010292
# generate integer valued scores with the same shape as normal quantile
# scores, this no longer v.d.Waerden, but something very similar
scores <- cscores(c(x,y), type="NormalQuantile", int=TRUE)
X <- sum(scores[seq(along=x)])
p2 <- pperm(X, scores, length(x), alternative="two.sided")
# compare p1 and p2
p1 - p2
#> [1] -0.02262443
# should give a "perfect" line
# Wilcoxon signed rank test
n <- 10
x <- rnorm(n, 5)
y <- rnorm(n, 5)
r <- cscores(abs(x - y), type="Wilcoxon")
pperm(sum(r[x - y > 0]), r, length(r))
#> [1] 0.7216797
wilcox.test(x,y, paired=TRUE, alternative="less")
#>
#> Wilcoxon signed rank exact test
#>
#> data: x and y
#> V = 33, p-value = 0.7217
#> alternative hypothesis: true location shift is less than 0
#>
psignrank(sum(r[x - y > 0]), length(r))
#> [1] 0.7216797
# Ansari-Bradley
n <- 10
x <- rnorm(n, 2, 1)
y <- rnorm(n, 2, 2)
# exact distribution using the Shift-Algorithm
sc <- cscores(c(x,y), type="Ansari")
dabexac <- dperm(0:(n*(2*n+1)/2), sc, n)
sum(dabexac)
#> [1] 1
# real scores are allowed (but only result in an approximation)
# e.g. v.d. Waerden test
n <- 10
x <- rnorm(n)
y <- rnorm(n)
scores <- cscores(c(x,y), type="NormalQuantile")
X <- sum(scores[seq(along=x)]) # <- v.d. Waerden normal quantile statistic
# critical value, two-sided test
abs(qperm(0.025, scores, length(x)))
#> Warning: cannot hold tol, tolerance: 0.010292
#> [1] 3.871345
# p-values
p1 <- pperm(X, scores, length(x), alternative="two.sided")
#> Warning: cannot hold tol, tolerance: 0.010292
# generate integer valued scores with the same shape as normal quantile
# scores, this no longer v.d.Waerden, but something very similar
scores <- cscores(c(x,y), type="NormalQuantile", int=TRUE)
X <- sum(scores[seq(along=x)])
p2 <- pperm(X, scores, length(x), alternative="two.sided")
# compare p1 and p2
p1 - p2
#> [1] -0.02262443