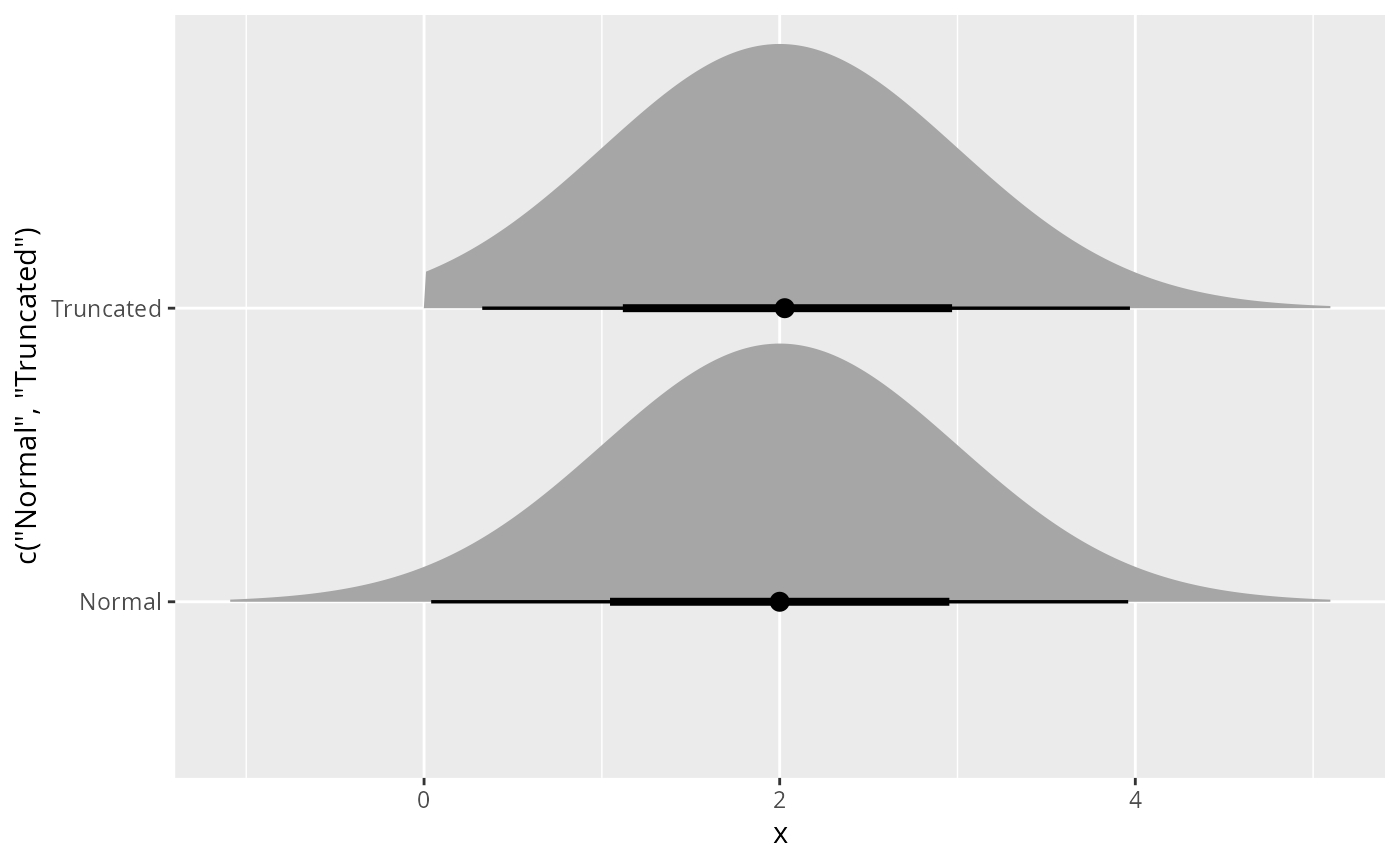
Note that the samples are generated using inverse transform sampling, and the means and variances are estimated from samples.
Details
We recommend reading this documentation on pkgdown which renders math nicely. https://pkg.mitchelloharawild.com/distributional/reference/dist_truncated.html
In the following, let \(X\) be a truncated random variable with
underlying distribution \(Y\), truncation bounds lower = \(a\) and
upper = \(b\), where \(F_Y(x)\) is the c.d.f. of \(Y\) and
\(f_Y(x)\) is the p.d.f. of \(Y\).
Support: \([a, b]\)
Mean: For the general case, the mean is approximated numerically. For a truncated Normal distribution with underlying mean \(\mu\) and standard deviation \(\sigma\), the mean is:
$$ E(X) = \mu + \frac{\phi(\alpha) - \phi(\beta)}{\Phi(\beta) - \Phi(\alpha)} \sigma $$
where \(\alpha = (a - \mu)/\sigma\), \(\beta = (b - \mu)/\sigma\), \(\phi\) is the standard Normal p.d.f., and \(\Phi\) is the standard Normal c.d.f.
Variance: Approximated numerically for all distributions.
Probability density function (p.d.f):
$$ f(x) = \begin{cases} \frac{f_Y(x)}{F_Y(b) - F_Y(a)} & \text{if } a \le x \le b \\ 0 & \text{otherwise} \end{cases} $$
Cumulative distribution function (c.d.f):
$$ F(x) = \begin{cases} 0 & \text{if } x < a \\ \frac{F_Y(x) - F_Y(a)}{F_Y(b) - F_Y(a)} & \text{if } a \le x \le b \\ 1 & \text{if } x > b \end{cases} $$
Quantile function:
$$ Q(p) = F_Y^{-1}(F_Y(a) + p(F_Y(b) - F_Y(a))) $$
clamped to the interval \([a, b]\).
Examples
dist <- dist_truncated(dist_normal(2,1), lower = 0)
dist
#> <distribution[1]>
#> [1] N(2, 1)[0,Inf]
mean(dist)
#> [1] 2.055248
variance(dist)
#> [1] 0.8864519
generate(dist, 10)
#> [[1]]
#> [1] 1.7415534 0.7471783 2.6599636 2.4990165 2.9423018 1.9174482 3.7826920
#> [8] 1.4461079 3.6731440 1.4203806
#>
density(dist, 2)
#> [1] 0.4082296
density(dist, 2, log = TRUE)
#> [1] -0.8959256
cdf(dist, 4)
#> [1] 0.9767203
quantile(dist, 0.7)
#> [1] 2.544133
if(requireNamespace("ggdist")) {
library(ggplot2)
ggplot() +
ggdist::stat_dist_halfeye(
aes(y = c("Normal", "Truncated"),
dist = c(dist_normal(2,1), dist_truncated(dist_normal(2,1), lower = 0)))
)
}