Symmetry Tests
SymmetryTests.RdTesting the symmetry of a numeric repeated measurements variable in a complete block design.
# S3 method for class 'formula'
sign_test(formula, data, subset = NULL, ...)
# S3 method for class 'SymmetryProblem'
sign_test(object, ...)
# S3 method for class 'formula'
wilcoxsign_test(formula, data, subset = NULL, ...)
# S3 method for class 'SymmetryProblem'
wilcoxsign_test(object, zero.method = c("Pratt", "Wilcoxon"), ...)
# S3 method for class 'formula'
friedman_test(formula, data, subset = NULL, ...)
# S3 method for class 'SymmetryProblem'
friedman_test(object, ...)
# S3 method for class 'formula'
quade_test(formula, data, subset = NULL, ...)
# S3 method for class 'SymmetryProblem'
quade_test(object, ...)Arguments
- formula
a formula of the form
y ~ x | blockwhereyis a numeric variable,xis a factor with two (sign_testandwilcoxsign_test) or more levels andblockis an optional factor (which is generated automatically if omitted).- data
an optional data frame containing the variables in the model formula.
- subset
an optional vector specifying a subset of observations to be used. Defaults to
NULL.- object
an object inheriting from class
"SymmetryProblem".- zero.method
a character, the method used to handle zeros: either
"Pratt"(default) or"Wilcoxon"; see ‘Details’.- ...
further arguments to be passed to
symmetry_test().
Details
sign_test(), wilcoxsign_test(), friedman_test() and
quade_test() provide the sign test, the Wilcoxon signed-rank test, the
Friedman test, the Page test and the Quade test. A general description of
these methods is given by Hollander and Wolfe (1999).
The null hypothesis of symmetry is tested. The response variable and the
measurement conditions are given by y and x, respectively, and
block is a factor where each level corresponds to exactly one subject
with repeated measurements. For sign_test and wilcoxsign_test,
formulae of the form y ~ x | block and y ~ x are allowed. The
latter form is interpreted as y is the first and x the second
measurement on the same subject.
If x is an ordered factor, the default scores, 1:nlevels(x), can
be altered using the scores argument (see symmetry_test());
this argument can also be used to coerce nominal factors to class
"ordered". In this case, a linear-by-linear association test is
computed and the direction of the alternative hypothesis can be specified
using the alternative argument. For the Friedman test, this extension
was given by Page (1963) and is known as the Page test.
For wilcoxsign_test(), the default method of handling zeros
(zero.method = "Pratt"), due to Pratt (1959), first rank-transforms the
absolute differences (including zeros) and then discards the ranks
corresponding to the zero-differences. The proposal by Wilcoxon (1949, p. 6)
first discards the zero-differences and then rank-transforms the remaining
absolute differences (zero.method = "Wilcoxon").
The conditional null distribution of the test statistic is used to obtain
\(p\)-values and an asymptotic approximation of the exact distribution is
used by default (distribution = "asymptotic"). Alternatively, the
distribution can be approximated via Monte Carlo resampling or computed
exactly for univariate two-sample problems by setting distribution to
"approximate" or "exact", respectively. See
asymptotic(), approximate() and
exact() for details.
Value
An object inheriting from class "IndependenceTest".
Note
Starting with coin version 1.0-16, the zero.method argument
replaced the (now removed) ties.method argument. The current default
is zero.method = "Pratt" whereas earlier versions had
ties.method = "HollanderWolfe", which is equivalent to
zero.method = "Wilcoxon".
References
Hollander, M. and Wolfe, D. A. (1999). Nonparametric Statistical Methods, Second Edition. New York: John Wiley & Sons.
Page, E. B. (1963). Ordered hypotheses for multiple treatments: a significance test for linear ranks. Journal of the American Statistical Association 58(301), 216–230. doi:10.1080/01621459.1963.10500843
Pratt, J. W. (1959). Remarks on zeros and ties in the Wilcoxon signed rank procedures. Journal of the American Statistical Association 54(287), 655–667. doi:10.1080/01621459.1959.10501526
Quade, D. (1979). Using weighted rankings in the analysis of complete blocks with additive block effects. Journal of the American Statistical Association 74(367), 680–683. doi:10.1080/01621459.1979.10481670
Wilcoxon, F. (1949). Some Rapid Approximate Statistical Procedures. New York: American Cyanamid Company.
Examples
## Example data from ?wilcox.test
y1 <- c(1.83, 0.50, 1.62, 2.48, 1.68, 1.88, 1.55, 3.06, 1.30)
y2 <- c(0.878, 0.647, 0.598, 2.05, 1.06, 1.29, 1.06, 3.14, 1.29)
## One-sided exact sign test
(st <- sign_test(y1 ~ y2, distribution = "exact",
alternative = "greater"))
#>
#> Exact Sign Test
#>
#> data: y by x (pos, neg)
#> stratified by block
#> Z = 1.6667, p-value = 0.08984
#> alternative hypothesis: true mu is greater than 0
#>
midpvalue(st) # mid-p-value
#> [1] 0.0546875
## One-sided exact Wilcoxon signed-rank test
(wt <- wilcoxsign_test(y1 ~ y2, distribution = "exact",
alternative = "greater"))
#>
#> Exact Wilcoxon-Pratt Signed-Rank Test
#>
#> data: y by x (pos, neg)
#> stratified by block
#> Z = 2.0732, p-value = 0.01953
#> alternative hypothesis: true mu is greater than 0
#>
statistic(wt, type = "linear")
#>
#> pos 40
midpvalue(wt) # mid-p-value
#> [1] 0.01660156
## Comparison with R's wilcox.test() function
wilcox.test(y1, y2, paired = TRUE, alternative = "greater")
#>
#> Wilcoxon signed rank exact test
#>
#> data: y1 and y2
#> V = 40, p-value = 0.01953
#> alternative hypothesis: true location shift is greater than 0
#>
## Data with explicit group and block information
dta <- data.frame(y = c(y1, y2), x = gl(2, length(y1)),
block = factor(rep(seq_along(y1), 2)))
## For two samples, the sign test is equivalent to the Friedman test...
sign_test(y ~ x | block, data = dta, distribution = "exact")
#>
#> Exact Sign Test
#>
#> data: y by x (pos, neg)
#> stratified by block
#> Z = 1.6667, p-value = 0.1797
#> alternative hypothesis: true mu is not equal to 0
#>
friedman_test(y ~ x | block, data = dta, distribution = "exact")
#>
#> Exact Friedman Test
#>
#> data: y by x (1, 2)
#> stratified by block
#> chi-squared = 2.7778, p-value = 0.1797
#>
## ...and the signed-rank test is equivalent to the Quade test
wilcoxsign_test(y ~ x | block, data = dta, distribution = "exact")
#>
#> Exact Wilcoxon-Pratt Signed-Rank Test
#>
#> data: y by x (pos, neg)
#> stratified by block
#> Z = 2.0732, p-value = 0.03906
#> alternative hypothesis: true mu is not equal to 0
#>
quade_test(y ~ x | block, data = dta, distribution = "exact")
#>
#> Exact Quade Test
#>
#> data: y by x (1, 2)
#> stratified by block
#> chi-squared = 4.2982, p-value = 0.03906
#>
## Comparison of three methods ("round out", "narrow angle", and "wide angle")
## for rounding first base.
## Hollander and Wolfe (1999, p. 274, Tab. 7.1)
rounding <- data.frame(
times = c(5.40, 5.50, 5.55,
5.85, 5.70, 5.75,
5.20, 5.60, 5.50,
5.55, 5.50, 5.40,
5.90, 5.85, 5.70,
5.45, 5.55, 5.60,
5.40, 5.40, 5.35,
5.45, 5.50, 5.35,
5.25, 5.15, 5.00,
5.85, 5.80, 5.70,
5.25, 5.20, 5.10,
5.65, 5.55, 5.45,
5.60, 5.35, 5.45,
5.05, 5.00, 4.95,
5.50, 5.50, 5.40,
5.45, 5.55, 5.50,
5.55, 5.55, 5.35,
5.45, 5.50, 5.55,
5.50, 5.45, 5.25,
5.65, 5.60, 5.40,
5.70, 5.65, 5.55,
6.30, 6.30, 6.25),
methods = factor(rep(1:3, 22),
labels = c("Round Out", "Narrow Angle", "Wide Angle")),
block = gl(22, 3)
)
## Asymptotic Friedman test
friedman_test(times ~ methods | block, data = rounding)
#>
#> Asymptotic Friedman Test
#>
#> data: times by
#> methods (Round Out, Narrow Angle, Wide Angle)
#> stratified by block
#> chi-squared = 11.143, df = 2, p-value = 0.003805
#>
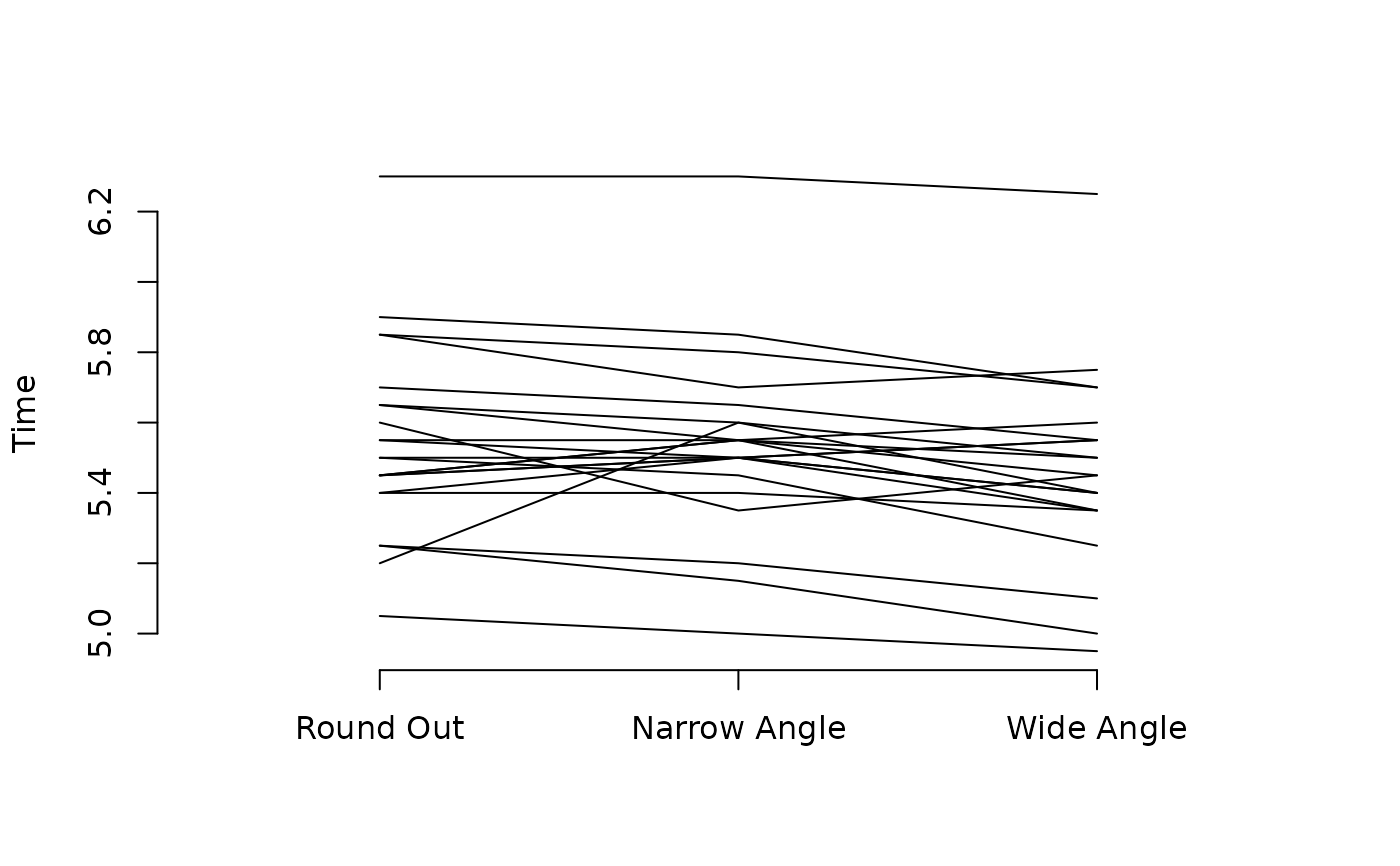
## Parallel coordinates plot
with(rounding, {
matplot(t(matrix(times, ncol = 3, byrow = TRUE)),
type = "l", lty = 1, col = 1, ylab = "Time", xlim = c(0.5, 3.5),
axes = FALSE)
axis(1, at = 1:3, labels = levels(methods))
axis(2)
})
 ## Where do the differences come from?
## Wilcoxon-Nemenyi-McDonald-Thompson test (Hollander and Wolfe, 1999, p. 295)
## Note: all pairwise comparisons
(st <- symmetry_test(times ~ methods | block, data = rounding,
ytrafo = function(data)
trafo(data, numeric_trafo = rank_trafo,
block = rounding$block),
xtrafo = mcp_trafo(methods = "Tukey")))
#>
#> Asymptotic General Symmetry Test
#>
#> data: times by
#> methods (Round Out, Narrow Angle, Wide Angle)
#> stratified by block
#> maxT = 3.2404, p-value = 0.00346
#> alternative hypothesis: two.sided
#>
## Simultaneous test of all pairwise comparisons
## Wide Angle vs. Round Out differ (Hollander and Wolfe, 1999, p. 296)
pvalue(st, method = "single-step") # subset pivotality is violated
#> Warning: p-values may be incorrect due to violation
#> of the subset pivotality condition
#>
#> Narrow Angle - Round Out 0.623910827
#> Wide Angle - Round Out 0.003516106
#> Wide Angle - Narrow Angle 0.053953055
## Strength Index of Cotton
## Hollander and Wolfe (1999, p. 286, Tab. 7.5)
cotton <- data.frame(
strength = c(7.46, 7.17, 7.76, 8.14, 7.63,
7.68, 7.57, 7.73, 8.15, 8.00,
7.21, 7.80, 7.74, 7.87, 7.93),
potash = ordered(rep(c(144, 108, 72, 54, 36), 3),
levels = c(144, 108, 72, 54, 36)),
block = gl(3, 5)
)
## One-sided asymptotic Page test
friedman_test(strength ~ potash | block, data = cotton, alternative = "greater")
#>
#> Asymptotic Page Test
#>
#> data: strength by
#> potash (144 < 108 < 72 < 54 < 36)
#> stratified by block
#> Z = 2.6558, p-value = 0.003956
#> alternative hypothesis: greater
#>
## One-sided approximative (Monte Carlo) Page test
friedman_test(strength ~ potash | block, data = cotton, alternative = "greater",
distribution = approximate(nresample = 10000))
#>
#> Approximative Page Test
#>
#> data: strength by
#> potash (144 < 108 < 72 < 54 < 36)
#> stratified by block
#> Z = 2.6558, p-value = 0.0023
#> alternative hypothesis: greater
#>
## Data from Quade (1979, p. 683)
dta <- data.frame(
y = c(52, 45, 38,
63, 79, 50,
45, 57, 39,
53, 51, 43,
47, 50, 56,
62, 72, 49,
49, 52, 40),
x = factor(rep(LETTERS[1:3], 7)),
b = factor(rep(1:7, each = 3))
)
## Approximative (Monte Carlo) Friedman test
## Quade (1979, p. 683)
friedman_test(y ~ x | b, data = dta,
distribution = approximate(nresample = 10000)) # chi^2 = 6.000
#>
#> Approximative Friedman Test
#>
#> data: y by x (A, B, C)
#> stratified by b
#> chi-squared = 6, p-value = 0.048
#>
## Approximative (Monte Carlo) Quade test
## Quade (1979, p. 683)
(qt <- quade_test(y ~ x | b, data = dta,
distribution = approximate(nresample = 10000))) # W = 8.157
#>
#> Approximative Quade Test
#>
#> data: y by x (A, B, C)
#> stratified by b
#> chi-squared = 8.1571, p-value = 0.0046
#>
## Comparison with R's quade.test() function
quade.test(y ~ x | b, data = dta)
#>
#> Quade test
#>
#> data: y and x and b
#> Quade F = 8.3765, num df = 2, denom df = 12, p-value = 0.005284
#>
## quade.test() uses an F-statistic
b <- nlevels(qt@statistic@block)
A <- sum(qt@statistic@ytrans^2)
B <- sum(statistic(qt, type = "linear")^2) / b
(b - 1) * B / (A - B) # F = 8.3765
#> [1] 8.376528
## Where do the differences come from?
## Wilcoxon-Nemenyi-McDonald-Thompson test (Hollander and Wolfe, 1999, p. 295)
## Note: all pairwise comparisons
(st <- symmetry_test(times ~ methods | block, data = rounding,
ytrafo = function(data)
trafo(data, numeric_trafo = rank_trafo,
block = rounding$block),
xtrafo = mcp_trafo(methods = "Tukey")))
#>
#> Asymptotic General Symmetry Test
#>
#> data: times by
#> methods (Round Out, Narrow Angle, Wide Angle)
#> stratified by block
#> maxT = 3.2404, p-value = 0.00346
#> alternative hypothesis: two.sided
#>
## Simultaneous test of all pairwise comparisons
## Wide Angle vs. Round Out differ (Hollander and Wolfe, 1999, p. 296)
pvalue(st, method = "single-step") # subset pivotality is violated
#> Warning: p-values may be incorrect due to violation
#> of the subset pivotality condition
#>
#> Narrow Angle - Round Out 0.623910827
#> Wide Angle - Round Out 0.003516106
#> Wide Angle - Narrow Angle 0.053953055
## Strength Index of Cotton
## Hollander and Wolfe (1999, p. 286, Tab. 7.5)
cotton <- data.frame(
strength = c(7.46, 7.17, 7.76, 8.14, 7.63,
7.68, 7.57, 7.73, 8.15, 8.00,
7.21, 7.80, 7.74, 7.87, 7.93),
potash = ordered(rep(c(144, 108, 72, 54, 36), 3),
levels = c(144, 108, 72, 54, 36)),
block = gl(3, 5)
)
## One-sided asymptotic Page test
friedman_test(strength ~ potash | block, data = cotton, alternative = "greater")
#>
#> Asymptotic Page Test
#>
#> data: strength by
#> potash (144 < 108 < 72 < 54 < 36)
#> stratified by block
#> Z = 2.6558, p-value = 0.003956
#> alternative hypothesis: greater
#>
## One-sided approximative (Monte Carlo) Page test
friedman_test(strength ~ potash | block, data = cotton, alternative = "greater",
distribution = approximate(nresample = 10000))
#>
#> Approximative Page Test
#>
#> data: strength by
#> potash (144 < 108 < 72 < 54 < 36)
#> stratified by block
#> Z = 2.6558, p-value = 0.0023
#> alternative hypothesis: greater
#>
## Data from Quade (1979, p. 683)
dta <- data.frame(
y = c(52, 45, 38,
63, 79, 50,
45, 57, 39,
53, 51, 43,
47, 50, 56,
62, 72, 49,
49, 52, 40),
x = factor(rep(LETTERS[1:3], 7)),
b = factor(rep(1:7, each = 3))
)
## Approximative (Monte Carlo) Friedman test
## Quade (1979, p. 683)
friedman_test(y ~ x | b, data = dta,
distribution = approximate(nresample = 10000)) # chi^2 = 6.000
#>
#> Approximative Friedman Test
#>
#> data: y by x (A, B, C)
#> stratified by b
#> chi-squared = 6, p-value = 0.048
#>
## Approximative (Monte Carlo) Quade test
## Quade (1979, p. 683)
(qt <- quade_test(y ~ x | b, data = dta,
distribution = approximate(nresample = 10000))) # W = 8.157
#>
#> Approximative Quade Test
#>
#> data: y by x (A, B, C)
#> stratified by b
#> chi-squared = 8.1571, p-value = 0.0046
#>
## Comparison with R's quade.test() function
quade.test(y ~ x | b, data = dta)
#>
#> Quade test
#>
#> data: y and x and b
#> Quade F = 8.3765, num df = 2, denom df = 12, p-value = 0.005284
#>
## quade.test() uses an F-statistic
b <- nlevels(qt@statistic@block)
A <- sum(qt@statistic@ytrans^2)
B <- sum(statistic(qt, type = "linear")^2) / b
(b - 1) * B / (A - B) # F = 8.3765
#> [1] 8.376528