Agglomerative Nesting (AGNES) Object
agnes.object.RdThe objects of class "agnes"
represent an agglomerative hierarchical clustering of a dataset.
GENERATION
This class of objects is returned from agnes.
METHODS
The "agnes" class has methods for the following generic functions:
print, summary, plot, and
as.dendrogram.
In addition, cutree(x, *) can be used to “cut”
the dendrogram in order to produce cluster assignments.
INHERITANCE
The class "agnes" inherits from "twins".
Therefore, the generic functions pltree and
as.hclust are available for agnes objects.
After applying as.hclust(), all its methods are
available, of course.
Value
A legitimate agnes object is a list with the following components:
- order
a vector giving a permutation of the original observations to allow for plotting, in the sense that the branches of a clustering tree will not cross.
- order.lab
a vector similar to
order, but containing observation labels instead of observation numbers. This component is only available if the original observations were labelled.- height
a vector with the distances between merging clusters at the successive stages.
- ac
the agglomerative coefficient, measuring the clustering structure of the dataset.
For each observation i, denote by m(i) its dissimilarity to the first cluster it is merged with, divided by the dissimilarity of the merger in the final step of the algorithm. The
acis the average of all 1 - m(i). It can also be seen as the average width (or the percentage filled) of the banner plot. Becauseacgrows with the number of observations, this measure should not be used to compare datasets of very different sizes.- merge
an (n-1) by 2 matrix, where n is the number of observations. Row i of
mergedescribes the merging of clusters at step i of the clustering. If a number j in the row is negative, then the single observation |j| is merged at this stage. If j is positive, then the merger is with the cluster formed at stage j of the algorithm.- diss
an object of class
"dissimilarity"(seedissimilarity.object), representing the total dissimilarity matrix of the dataset.- data
a matrix containing the original or standardized measurements, depending on the
standoption of the functionagnes. If a dissimilarity matrix was given as input structure, then this component is not available.
Examples
data(agriculture)
ag.ag <- agnes(agriculture)
class(ag.ag)
#> [1] "agnes" "twins"
pltree(ag.ag) # the dendrogram
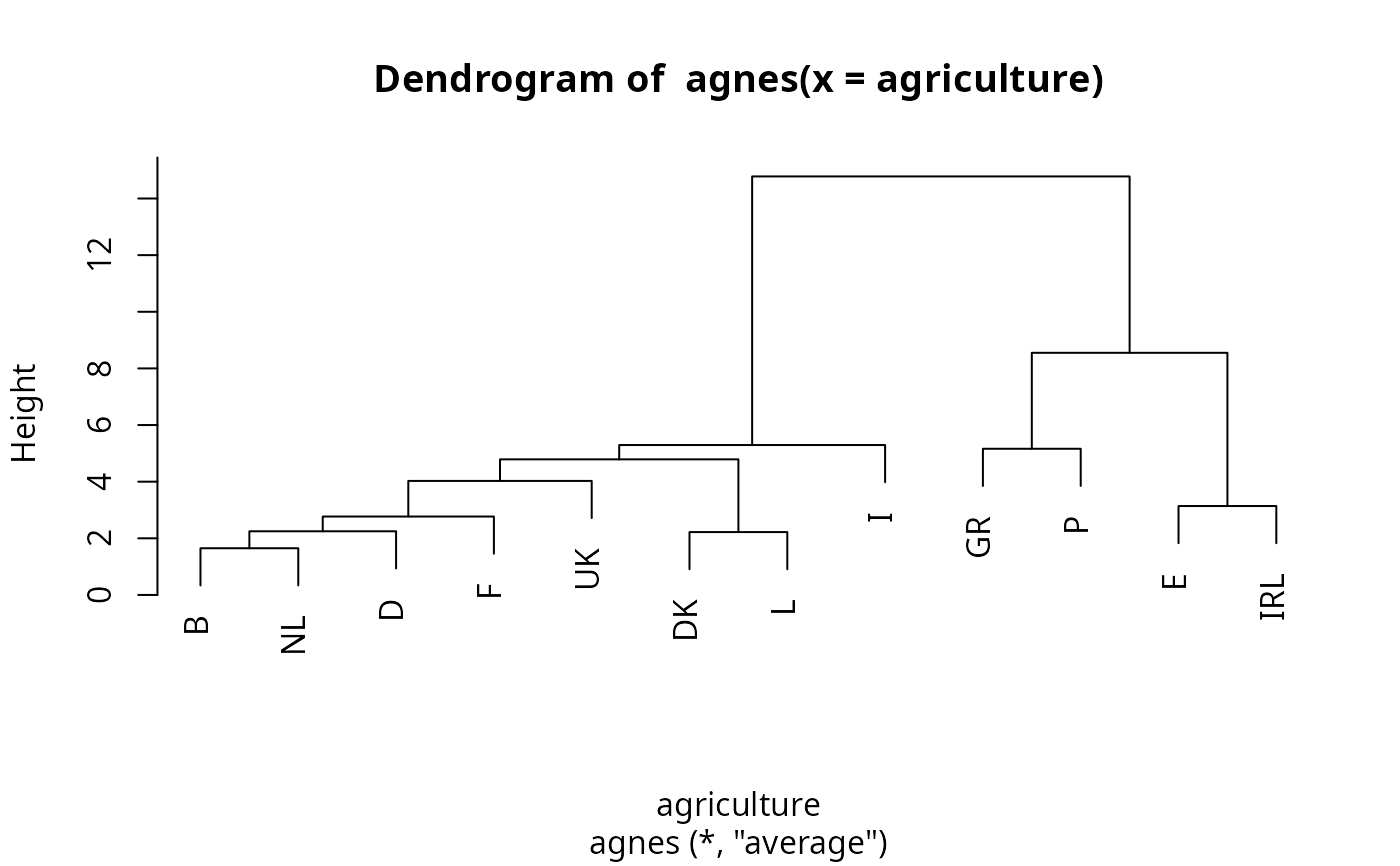 ## cut the dendrogram -> get cluster assignments:
(ck3 <- cutree(ag.ag, k = 3))
#> [1] 1 1 1 2 3 1 3 1 1 1 2 1
(ch6 <- cutree(as.hclust(ag.ag), h = 6))
#> B DK D GR E F IRL I L NL P UK
#> 1 1 1 2 3 1 3 1 1 1 2 1
stopifnot(identical(unname(ch6), ck3))
## cut the dendrogram -> get cluster assignments:
(ck3 <- cutree(ag.ag, k = 3))
#> [1] 1 1 1 2 3 1 3 1 1 1 2 1
(ch6 <- cutree(as.hclust(ag.ag), h = 6))
#> B DK D GR E F IRL I L NL P UK
#> 1 1 1 2 3 1 3 1 1 1 2 1
stopifnot(identical(unname(ch6), ck3))