Marginal Predictions with marginaleffects::avg_predictions()
Source: R/marginal_tidiers.R
tidy_marginal_predictions.RdUse marginaleffects::avg_predictions() to estimate marginal predictions for
each variable of a model and return a tibble tidied in a way that it could
be used by broom.helpers functions.
See marginaleffects::avg_predictions() for a list of supported models.
tidy_marginal_predictions(
x,
variables_list = "auto",
conf.int = TRUE,
conf.level = 0.95,
...
)
variables_to_predict(
model,
interactions = TRUE,
categorical = unique,
continuous = stats::fivenum
)
plot_marginal_predictions(x, variables_list = "auto", conf.level = 0.95, ...)Arguments
- x
(a model object, e.g.
glm)
A model to be tidied.- variables_list
(
listorstring)
A list whose elements will be sequentially passed tovariablesinmarginaleffects::avg_predictions()(see details below); alternatively, it could also be the string"auto"(default) or"no_interaction".- conf.int
(
logical)
Whether or not to include a confidence interval in the tidied output.- conf.level
(
numeric)
The confidence level to use for the confidence interval (between0ans1).- ...
Additional parameters passed to
marginaleffects::avg_predictions().- model
(a model object, e.g.
glm)
A model.- interactions
(
logical)
Should combinations of variables corresponding to interactions be returned?- categorical
(
predictor values)
Default values for categorical variables.- continuous
(
predictor values)
Default values for continuous variables.
Details
Marginal predictions are obtained by calling, for each variable,
marginaleffects::avg_predictions() with the same variable being used for
the variables and the by argument.
Considering a categorical variable named cat, tidy_marginal_predictions()
will call avg_predictions(model, variables = list(cat = unique), by = "cat")
to obtain average marginal predictions for this variable.
Considering a continuous variable named cont, tidy_marginal_predictions()
will call avg_predictions(model, variables = list(cont = "fivenum"), by = "cont")
to obtain average marginal predictions for this variable at the minimum, the
first quartile, the median, the third quartile and the maximum of the observed
values of cont.
By default, average marginal predictions are computed: predictions are made
using a counterfactual grid for each value of the variable of interest,
before averaging the results. Marginal predictions at the mean could be
obtained by indicating newdata = "mean". Other assumptions are possible,
see the help file of marginaleffects::avg_predictions().
tidy_marginal_predictions() will compute marginal predictions for each
variable or combination of variables, before stacking the results in a unique
tibble. This is why tidy_marginal_predictions() has a variables_list
argument consisting of a list of specifications that will be passed
sequentially to the variables argument of marginaleffects::avg_predictions().
The helper function variables_to_predict() could be used to automatically
generate a suitable list to be used with variables_list. By default, all
unique values are retained for categorical variables and fivenum (i.e.
Tukey's five numbers, minimum, quartiles and maximum) for continuous variables.
When interactions = FALSE, variables_to_predict() will return a list of
all individual variables used in the model. If interactions = FALSE, it
will search for higher order combinations of variables (see
model_list_higher_order_variables()).
variables_list's default value, "auto", calls
variables_to_predict(interactions = TRUE) while "no_interaction" is a
shortcut for variables_to_predict(interactions = FALSE).
You can also provide custom specifications (see examples).
plot_marginal_predictions() works in a similar way and returns a list of
plots that could be combined with patchwork::wrap_plots() (see examples).
For more information, see vignette("marginal_tidiers", "broom.helpers").
See also
marginaleffects::avg_predictions()
Other marginal_tieders:
tidy_all_effects(),
tidy_avg_comparisons(),
tidy_avg_slopes(),
tidy_ggpredict(),
tidy_marginal_contrasts(),
tidy_margins()
Examples
# example code
# \donttest{
# Average Marginal Predictions
df <- Titanic |>
dplyr::as_tibble() |>
tidyr::uncount(n) |>
dplyr::mutate(Survived = factor(Survived, c("No", "Yes")))
mod <- glm(
Survived ~ Class + Age + Sex,
data = df, family = binomial
)
tidy_marginal_predictions(mod)
#> variable term estimate std.error statistic p.value s.value
#> 1 Class 1st 0.5181374 0.027913630 18.56217 6.502960e-77 253.08737
#> 2 Class 2nd 0.3229724 0.024270457 13.30722 2.101515e-40 131.80569
#> 3 Class 3rd 0.2115172 0.013107421 16.13721 1.397105e-58 192.18939
#> 4 Class Crew 0.3504908 0.015030218 23.31908 2.839372e-120 397.12580
#> 5 Age Adult 0.3140462 0.008827755 35.57486 3.429343e-277 918.39615
#> 6 Age Child 0.5112612 0.047737203 10.70991 9.144970e-27 86.49908
#> 7 Sex Female 0.7255784 0.021732729 33.38644 2.157383e-244 809.44117
#> 8 Sex Male 0.2182251 0.009932243 21.97138 5.410102e-107 353.01065
#> conf.low conf.high
#> 1 0.4634277 0.5728471
#> 2 0.2754031 0.3705416
#> 3 0.1858272 0.2372073
#> 4 0.3210322 0.3799495
#> 5 0.2967441 0.3313483
#> 6 0.4176980 0.6048244
#> 7 0.6829830 0.7681737
#> 8 0.1987582 0.2376919
tidy_plus_plus(mod, tidy_fun = tidy_marginal_predictions)
#> # A tibble: 8 × 20
#> term variable var_label var_class var_type var_nlevels contrasts
#> <chr> <chr> <chr> <chr> <chr> <int> <chr>
#> 1 1st Class Class character categorical 4 contr.treatment
#> 2 2nd Class Class character categorical 4 contr.treatment
#> 3 3rd Class Class character categorical 4 contr.treatment
#> 4 Crew Class Class character categorical 4 contr.treatment
#> 5 Adult Age Age character dichotomous 2 contr.treatment
#> 6 Child Age Age character dichotomous 2 contr.treatment
#> 7 Female Sex Sex character dichotomous 2 contr.treatment
#> 8 Male Sex Sex character dichotomous 2 contr.treatment
#> # ℹ 13 more variables: contrasts_type <chr>, reference_row <lgl>, label <chr>,
#> # n_obs <dbl>, n_event <dbl>, estimate <dbl>, std.error <dbl>,
#> # statistic <dbl>, p.value <dbl>, s.value <dbl>, conf.low <dbl>,
#> # conf.high <dbl>, label_attr <chr>
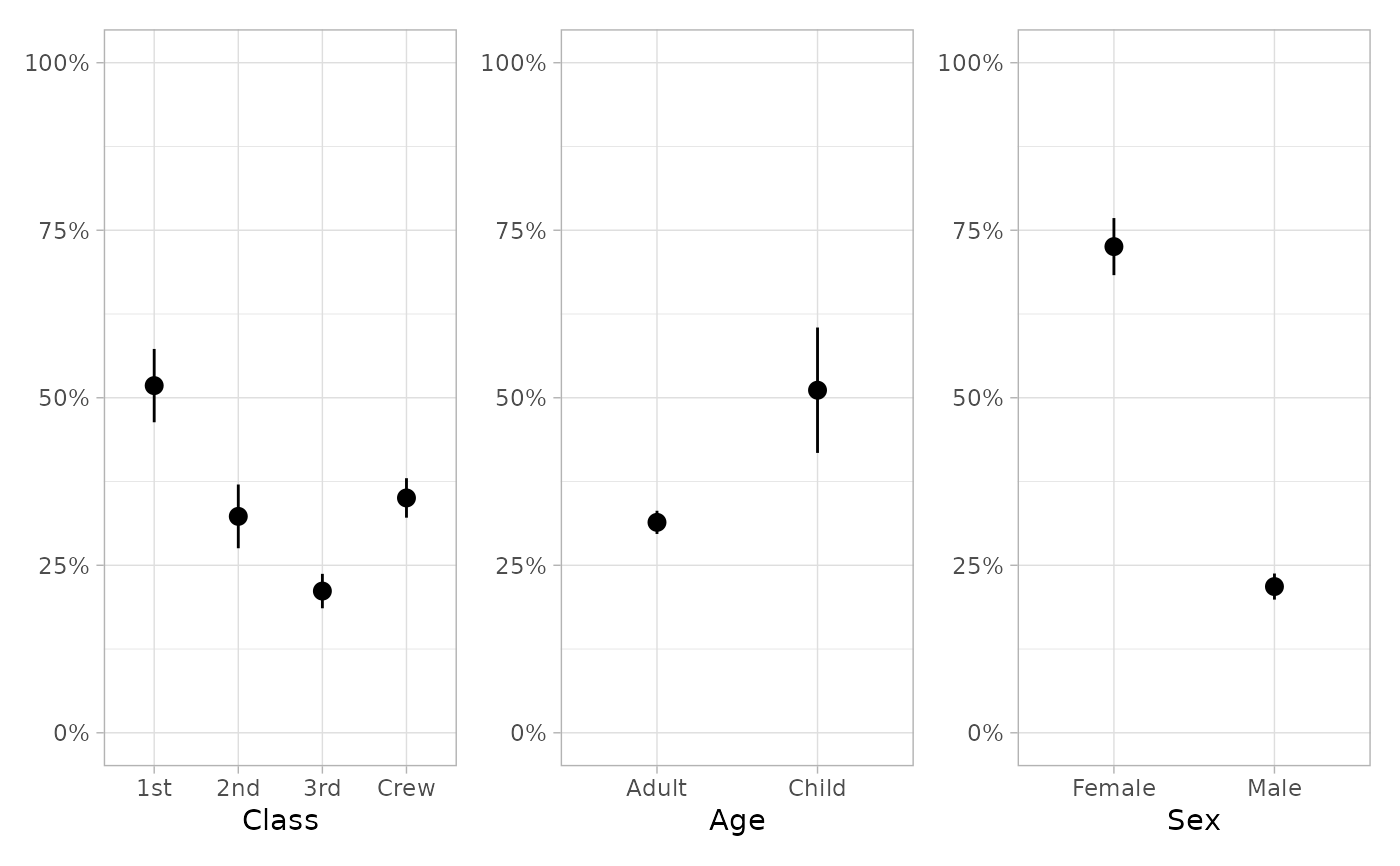
if (require("patchwork")) {
plot_marginal_predictions(mod) |> patchwork::wrap_plots()
plot_marginal_predictions(mod) |>
patchwork::wrap_plots() &
ggplot2::scale_y_continuous(limits = c(0, 1), label = scales::percent)
}
#> Loading required package: patchwork
 mod2 <- lm(Petal.Length ~ poly(Petal.Width, 2) + Species, data = iris)
tidy_marginal_predictions(mod2)
#> variable term estimate std.error statistic p.value s.value
#> 1 Petal.Width 0.1 2.302462 0.28351129 8.121235 4.614622e-16 50.94464
#> 2 Petal.Width 0.3 2.626585 0.20194705 13.006308 1.126554e-38 126.06135
#> 3 Petal.Width 1.3 3.997316 0.09875592 40.476722 0.000000e+00 Inf
#> 4 Petal.Width 1.8 4.526502 0.14311589 31.628228 1.511205e-219 726.90655
#> 5 Petal.Width 2.5 5.092442 0.19970017 25.500436 1.949411e-143 474.07268
#> 6 Species setosa 2.681553 0.22806207 11.757995 6.424184e-32 103.61819
#> 7 Species versicolor 3.998581 0.10599504 37.724221 0.000000e+00 Inf
#> 8 Species virginica 4.593867 0.15720110 29.222868 9.935198e-188 621.20993
#> conf.low conf.high
#> 1 1.746790 2.858134
#> 2 2.230776 3.022394
#> 3 3.803758 4.190874
#> 4 4.246000 4.807004
#> 5 4.701036 5.483847
#> 6 2.234559 3.128546
#> 7 3.790834 4.206327
#> 8 4.285758 4.901975
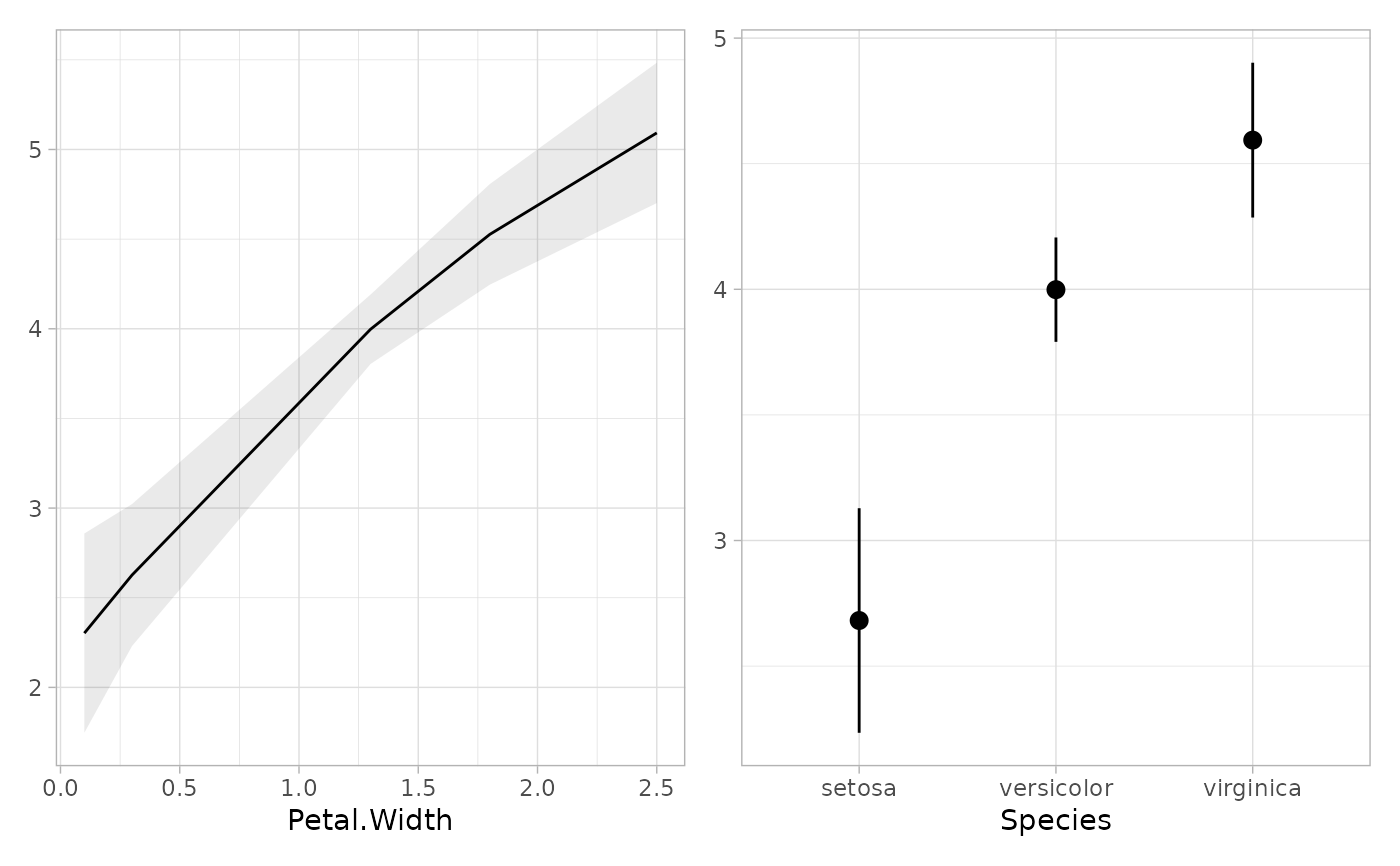
if (require("patchwork")) {
plot_marginal_predictions(mod2) |> patchwork::wrap_plots()
}
mod2 <- lm(Petal.Length ~ poly(Petal.Width, 2) + Species, data = iris)
tidy_marginal_predictions(mod2)
#> variable term estimate std.error statistic p.value s.value
#> 1 Petal.Width 0.1 2.302462 0.28351129 8.121235 4.614622e-16 50.94464
#> 2 Petal.Width 0.3 2.626585 0.20194705 13.006308 1.126554e-38 126.06135
#> 3 Petal.Width 1.3 3.997316 0.09875592 40.476722 0.000000e+00 Inf
#> 4 Petal.Width 1.8 4.526502 0.14311589 31.628228 1.511205e-219 726.90655
#> 5 Petal.Width 2.5 5.092442 0.19970017 25.500436 1.949411e-143 474.07268
#> 6 Species setosa 2.681553 0.22806207 11.757995 6.424184e-32 103.61819
#> 7 Species versicolor 3.998581 0.10599504 37.724221 0.000000e+00 Inf
#> 8 Species virginica 4.593867 0.15720110 29.222868 9.935198e-188 621.20993
#> conf.low conf.high
#> 1 1.746790 2.858134
#> 2 2.230776 3.022394
#> 3 3.803758 4.190874
#> 4 4.246000 4.807004
#> 5 4.701036 5.483847
#> 6 2.234559 3.128546
#> 7 3.790834 4.206327
#> 8 4.285758 4.901975
if (require("patchwork")) {
plot_marginal_predictions(mod2) |> patchwork::wrap_plots()
}
 tidy_marginal_predictions(
mod2,
variables_list = variables_to_predict(mod2, continuous = "threenum")
)
#> variable term estimate std.error statistic p.value
#> 1 Petal.Width 0.437095664372987 2.839141 0.15360160 18.48380 2.788266e-76
#> 2 Petal.Width 1.19933333333333 3.878182 0.08698259 44.58572 0.000000e+00
#> 3 Petal.Width 1.96157100229368 4.675245 0.15291913 30.57332 2.770493e-205
#> 4 Species setosa 2.681553 0.22806207 11.75799 6.424184e-32
#> 5 Species versicolor 3.998581 0.10599504 37.72422 0.000000e+00
#> 6 Species virginica 4.593867 0.15720110 29.22287 9.935198e-188
#> s.value conf.low conf.high
#> 1 250.9872 2.538088 3.140195
#> 2 Inf 3.707699 4.048664
#> 3 679.5251 4.375529 4.974961
#> 4 103.6182 2.234559 3.128546
#> 5 Inf 3.790834 4.206327
#> 6 621.2099 4.285758 4.901975
tidy_marginal_predictions(
mod2,
variables_list = list(
list(Petal.Width = c(0, 1, 2, 3)),
list(Species = unique)
)
)
#> variable term estimate std.error statistic p.value s.value
#> 1 Petal.Width 0 2.134153 0.32883856 6.489972 8.585236e-11 33.43935
#> 2 Petal.Width 1 3.629827 0.06654813 54.544386 0.000000e+00 Inf
#> 3 Petal.Width 2 4.709023 0.15519349 30.342916 3.115795e-202 669.38988
#> 4 Petal.Width 3 5.371741 0.31274123 17.176311 3.995257e-66 217.24897
#> 5 Species setosa 2.681553 0.22806207 11.757995 6.424184e-32 103.61819
#> 6 Species versicolor 3.998581 0.10599504 37.724221 0.000000e+00 Inf
#> 7 Species virginica 4.593867 0.15720110 29.222868 9.935198e-188 621.20993
#> conf.low conf.high
#> 1 1.489641 2.778665
#> 2 3.499395 3.760259
#> 3 4.404849 5.013197
#> 4 4.758779 5.984702
#> 5 2.234559 3.128546
#> 6 3.790834 4.206327
#> 7 4.285758 4.901975
tidy_marginal_predictions(
mod2,
variables_list = list(list(Species = unique, Petal.Width = 1:3))
)
#> variable term estimate std.error statistic
#> 1 Species:Petal.Width setosa * 1 2.553380 0.25261804 10.107670
#> 2 Species:Petal.Width setosa * 2 3.632576 0.37561389 9.671036
#> 3 Species:Petal.Width setosa * 3 4.295293 0.42138341 10.193314
#> 4 Species:Petal.Width versicolor * 1 3.870408 0.08240086 46.970478
#> 5 Species:Petal.Width versicolor * 2 4.949603 0.11523718 42.951446
#> 6 Species:Petal.Width versicolor * 3 5.612321 0.35269177 15.912822
#> 7 Species:Petal.Width virginica * 1 4.465694 0.16675860 26.779393
#> 8 Species:Petal.Width virginica * 2 5.544890 0.05494211 100.922404
#> 9 Species:Petal.Width virginica * 3 6.207607 0.27675645 22.429857
#> p.value s.value conf.low conf.high
#> 1 5.108488e-24 77.37338 2.058257 3.048502
#> 2 4.003040e-22 71.08132 2.896386 4.368765
#> 3 2.123987e-24 78.63950 3.469397 5.121190
#> 4 0.000000e+00 Inf 3.708905 4.031910
#> 5 0.000000e+00 Inf 4.723743 5.175464
#> 6 5.163313e-57 186.98160 4.921058 6.303584
#> 7 5.616443e-158 522.37498 4.138853 4.792535
#> 8 0.000000e+00 Inf 5.437205 5.652574
#> 9 2.012845e-111 367.72478 5.665175 6.750040
# Model with interactions
mod3 <- glm(
Survived ~ Sex * Age + Class,
data = df, family = binomial
)
tidy_marginal_predictions(mod3)
#> variable term estimate std.error statistic p.value
#> 1 Class 1st 0.5122895 0.027808505 18.422043 8.743958e-76
#> 2 Class 2nd 0.3202040 0.023739078 13.488478 1.828371e-41
#> 3 Class 3rd 0.2095767 0.012826120 16.339834 5.139221e-60
#> 4 Class Crew 0.3587744 0.014931459 24.028089 1.414862e-127
#> 5 Sex:Age Female * Adult 0.7419417 0.021989818 33.740241 1.486478e-249
#> 6 Sex:Age Female * Child 0.7217706 0.059574135 12.115503 8.742642e-34
#> 7 Sex:Age Male * Adult 0.2032339 0.009817932 20.700274 3.444078e-95
#> 8 Sex:Age Male * Child 0.5723894 0.060149442 9.516122 1.797624e-21
#> s.value conf.low conf.high
#> 1 249.3382 0.4577858 0.5667931
#> 2 135.3285 0.2736763 0.3667318
#> 3 196.9541 0.1844379 0.2347154
#> 4 421.3842 0.3295093 0.3880395
#> 5 826.5882 0.6988425 0.7850410
#> 6 109.8175 0.6050074 0.8385338
#> 7 313.7991 0.1839911 0.2224767
#> 8 68.9144 0.4544987 0.6902802
tidy_marginal_predictions(mod3, "no_interaction")
#> variable term estimate std.error statistic p.value s.value
#> 1 Sex Female 0.7407251 0.021107078 35.09368 8.414977e-270 893.8476
#> 2 Sex Male 0.2191225 0.009889623 22.15681 8.967724e-109 358.9254
#> 3 Age Adult 0.3142777 0.008706739 36.09591 2.629051e-285 945.3550
#> 4 Age Child 0.6033792 0.050063958 12.05217 1.889174e-33 108.7059
#> 5 Class 1st 0.5122895 0.027808505 18.42204 8.743958e-76 249.3382
#> 6 Class 2nd 0.3202040 0.023739078 13.48848 1.828371e-41 135.3285
#> 7 Class 3rd 0.2095767 0.012826120 16.33983 5.139221e-60 196.9541
#> 8 Class Crew 0.3587744 0.014931459 24.02809 1.414862e-127 421.3842
#> conf.low conf.high
#> 1 0.6993560 0.7820942
#> 2 0.1997392 0.2385058
#> 3 0.2972128 0.3313426
#> 4 0.5052556 0.7015027
#> 5 0.4577858 0.5667931
#> 6 0.2736763 0.3667318
#> 7 0.1844379 0.2347154
#> 8 0.3295093 0.3880395
if (require("patchwork")) {
plot_marginal_predictions(mod3) |>
patchwork::wrap_plots()
plot_marginal_predictions(mod3, "no_interaction") |>
patchwork::wrap_plots()
}
tidy_marginal_predictions(
mod2,
variables_list = variables_to_predict(mod2, continuous = "threenum")
)
#> variable term estimate std.error statistic p.value
#> 1 Petal.Width 0.437095664372987 2.839141 0.15360160 18.48380 2.788266e-76
#> 2 Petal.Width 1.19933333333333 3.878182 0.08698259 44.58572 0.000000e+00
#> 3 Petal.Width 1.96157100229368 4.675245 0.15291913 30.57332 2.770493e-205
#> 4 Species setosa 2.681553 0.22806207 11.75799 6.424184e-32
#> 5 Species versicolor 3.998581 0.10599504 37.72422 0.000000e+00
#> 6 Species virginica 4.593867 0.15720110 29.22287 9.935198e-188
#> s.value conf.low conf.high
#> 1 250.9872 2.538088 3.140195
#> 2 Inf 3.707699 4.048664
#> 3 679.5251 4.375529 4.974961
#> 4 103.6182 2.234559 3.128546
#> 5 Inf 3.790834 4.206327
#> 6 621.2099 4.285758 4.901975
tidy_marginal_predictions(
mod2,
variables_list = list(
list(Petal.Width = c(0, 1, 2, 3)),
list(Species = unique)
)
)
#> variable term estimate std.error statistic p.value s.value
#> 1 Petal.Width 0 2.134153 0.32883856 6.489972 8.585236e-11 33.43935
#> 2 Petal.Width 1 3.629827 0.06654813 54.544386 0.000000e+00 Inf
#> 3 Petal.Width 2 4.709023 0.15519349 30.342916 3.115795e-202 669.38988
#> 4 Petal.Width 3 5.371741 0.31274123 17.176311 3.995257e-66 217.24897
#> 5 Species setosa 2.681553 0.22806207 11.757995 6.424184e-32 103.61819
#> 6 Species versicolor 3.998581 0.10599504 37.724221 0.000000e+00 Inf
#> 7 Species virginica 4.593867 0.15720110 29.222868 9.935198e-188 621.20993
#> conf.low conf.high
#> 1 1.489641 2.778665
#> 2 3.499395 3.760259
#> 3 4.404849 5.013197
#> 4 4.758779 5.984702
#> 5 2.234559 3.128546
#> 6 3.790834 4.206327
#> 7 4.285758 4.901975
tidy_marginal_predictions(
mod2,
variables_list = list(list(Species = unique, Petal.Width = 1:3))
)
#> variable term estimate std.error statistic
#> 1 Species:Petal.Width setosa * 1 2.553380 0.25261804 10.107670
#> 2 Species:Petal.Width setosa * 2 3.632576 0.37561389 9.671036
#> 3 Species:Petal.Width setosa * 3 4.295293 0.42138341 10.193314
#> 4 Species:Petal.Width versicolor * 1 3.870408 0.08240086 46.970478
#> 5 Species:Petal.Width versicolor * 2 4.949603 0.11523718 42.951446
#> 6 Species:Petal.Width versicolor * 3 5.612321 0.35269177 15.912822
#> 7 Species:Petal.Width virginica * 1 4.465694 0.16675860 26.779393
#> 8 Species:Petal.Width virginica * 2 5.544890 0.05494211 100.922404
#> 9 Species:Petal.Width virginica * 3 6.207607 0.27675645 22.429857
#> p.value s.value conf.low conf.high
#> 1 5.108488e-24 77.37338 2.058257 3.048502
#> 2 4.003040e-22 71.08132 2.896386 4.368765
#> 3 2.123987e-24 78.63950 3.469397 5.121190
#> 4 0.000000e+00 Inf 3.708905 4.031910
#> 5 0.000000e+00 Inf 4.723743 5.175464
#> 6 5.163313e-57 186.98160 4.921058 6.303584
#> 7 5.616443e-158 522.37498 4.138853 4.792535
#> 8 0.000000e+00 Inf 5.437205 5.652574
#> 9 2.012845e-111 367.72478 5.665175 6.750040
# Model with interactions
mod3 <- glm(
Survived ~ Sex * Age + Class,
data = df, family = binomial
)
tidy_marginal_predictions(mod3)
#> variable term estimate std.error statistic p.value
#> 1 Class 1st 0.5122895 0.027808505 18.422043 8.743958e-76
#> 2 Class 2nd 0.3202040 0.023739078 13.488478 1.828371e-41
#> 3 Class 3rd 0.2095767 0.012826120 16.339834 5.139221e-60
#> 4 Class Crew 0.3587744 0.014931459 24.028089 1.414862e-127
#> 5 Sex:Age Female * Adult 0.7419417 0.021989818 33.740241 1.486478e-249
#> 6 Sex:Age Female * Child 0.7217706 0.059574135 12.115503 8.742642e-34
#> 7 Sex:Age Male * Adult 0.2032339 0.009817932 20.700274 3.444078e-95
#> 8 Sex:Age Male * Child 0.5723894 0.060149442 9.516122 1.797624e-21
#> s.value conf.low conf.high
#> 1 249.3382 0.4577858 0.5667931
#> 2 135.3285 0.2736763 0.3667318
#> 3 196.9541 0.1844379 0.2347154
#> 4 421.3842 0.3295093 0.3880395
#> 5 826.5882 0.6988425 0.7850410
#> 6 109.8175 0.6050074 0.8385338
#> 7 313.7991 0.1839911 0.2224767
#> 8 68.9144 0.4544987 0.6902802
tidy_marginal_predictions(mod3, "no_interaction")
#> variable term estimate std.error statistic p.value s.value
#> 1 Sex Female 0.7407251 0.021107078 35.09368 8.414977e-270 893.8476
#> 2 Sex Male 0.2191225 0.009889623 22.15681 8.967724e-109 358.9254
#> 3 Age Adult 0.3142777 0.008706739 36.09591 2.629051e-285 945.3550
#> 4 Age Child 0.6033792 0.050063958 12.05217 1.889174e-33 108.7059
#> 5 Class 1st 0.5122895 0.027808505 18.42204 8.743958e-76 249.3382
#> 6 Class 2nd 0.3202040 0.023739078 13.48848 1.828371e-41 135.3285
#> 7 Class 3rd 0.2095767 0.012826120 16.33983 5.139221e-60 196.9541
#> 8 Class Crew 0.3587744 0.014931459 24.02809 1.414862e-127 421.3842
#> conf.low conf.high
#> 1 0.6993560 0.7820942
#> 2 0.1997392 0.2385058
#> 3 0.2972128 0.3313426
#> 4 0.5052556 0.7015027
#> 5 0.4577858 0.5667931
#> 6 0.2736763 0.3667318
#> 7 0.1844379 0.2347154
#> 8 0.3295093 0.3880395
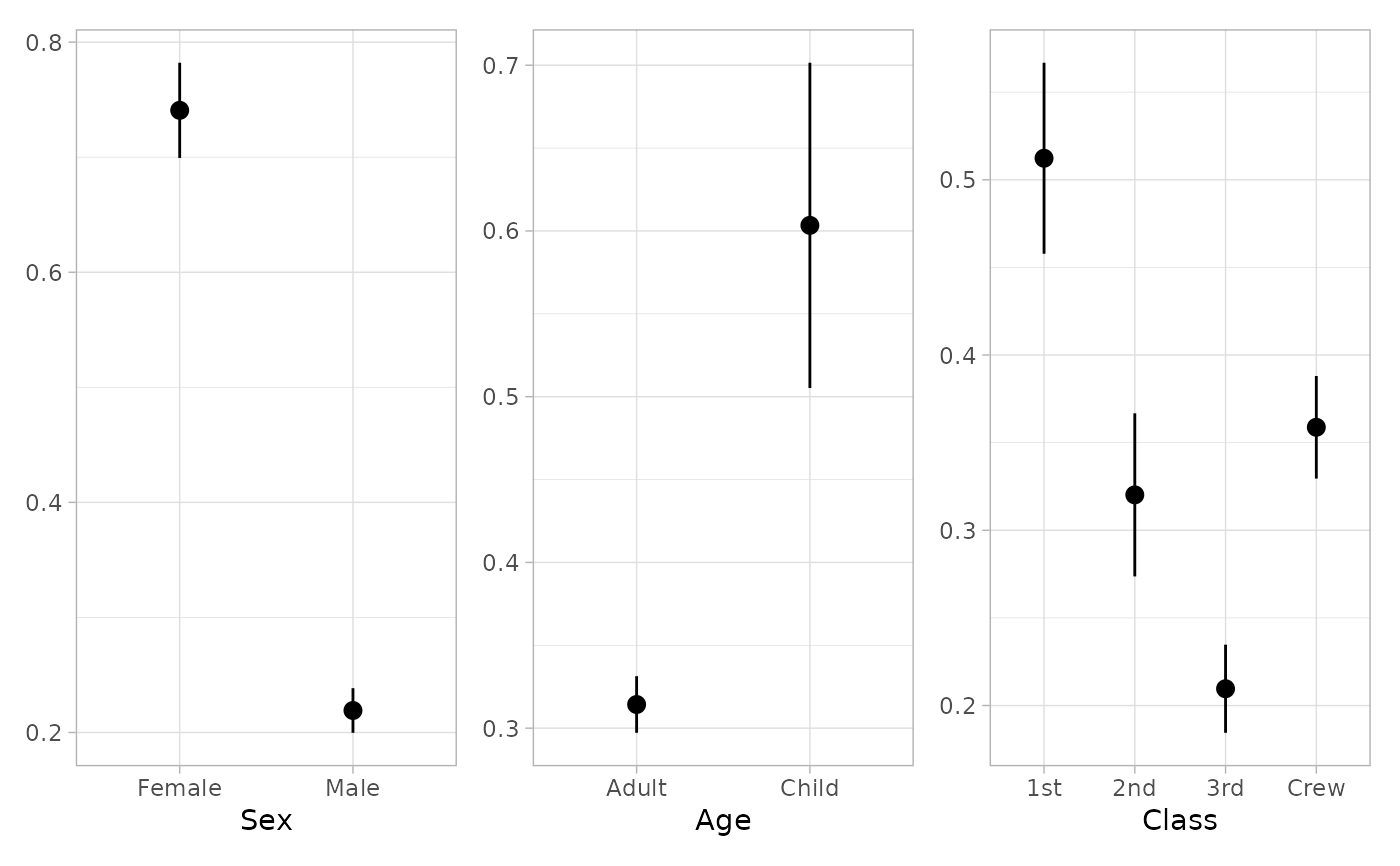
if (require("patchwork")) {
plot_marginal_predictions(mod3) |>
patchwork::wrap_plots()
plot_marginal_predictions(mod3, "no_interaction") |>
patchwork::wrap_plots()
}
 tidy_marginal_predictions(
mod3,
variables_list = list(
list(Class = unique, Sex = "Female"),
list(Age = unique)
)
)
#> variable term estimate std.error statistic p.value
#> 1 Class:Sex 1st * Female 0.8980680 0.015940375 56.33920 0.000000e+00
#> 2 Class:Sex 2nd * Female 0.7580905 0.030941812 24.50052 1.458340e-132
#> 3 Class:Sex 3rd * Female 0.5904013 0.031062754 19.00673 1.500210e-80
#> 4 Class:Sex Crew * Female 0.7978123 0.026646078 29.94108 5.749357e-197
#> 5 Age Adult 0.3142777 0.008706739 36.09591 2.629051e-285
#> 6 Age Child 0.6033792 0.050063958 12.05217 1.889174e-33
#> s.value conf.low conf.high
#> 1 Inf 0.8668255 0.9293106
#> 2 437.9502 0.6974457 0.8187354
#> 3 265.1691 0.5295195 0.6512832
#> 4 651.8964 0.7455869 0.8500376
#> 5 945.3550 0.2972128 0.3313426
#> 6 108.7059 0.5052556 0.7015027
# Marginal Predictions at the Mean
tidy_marginal_predictions(mod, newdata = "mean")
#> variable term estimate std.error statistic p.value s.value
#> 1 Class 1st 0.4070382 0.03286740 12.384253 3.180310e-35 114.59832
#> 2 Class 2nd 0.1987193 0.02474435 8.030897 9.676261e-16 49.87640
#> 3 Class 3rd 0.1039594 0.01182240 8.793426 1.450667e-18 59.25799
#> 4 Class Crew 0.2254997 0.01405834 16.040279 6.685371e-58 189.93082
#> 5 Age Adult 0.2254997 0.01405834 16.040279 6.685371e-58 189.93082
#> 6 Age Child 0.4570172 0.06370387 7.174088 7.279069e-13 40.32131
#> 7 Sex Female 0.7660538 0.02841756 26.957057 4.714993e-160 529.27124
#> 8 Sex Male 0.2254997 0.01405834 16.040279 6.685371e-58 189.93082
#> conf.low conf.high
#> 1 0.34261928 0.4714571
#> 2 0.15022129 0.2472174
#> 3 0.08078793 0.1271309
#> 4 0.19794588 0.2530536
#> 5 0.19794588 0.2530536
#> 6 0.33215989 0.5818745
#> 7 0.71035641 0.8217512
#> 8 0.19794588 0.2530536
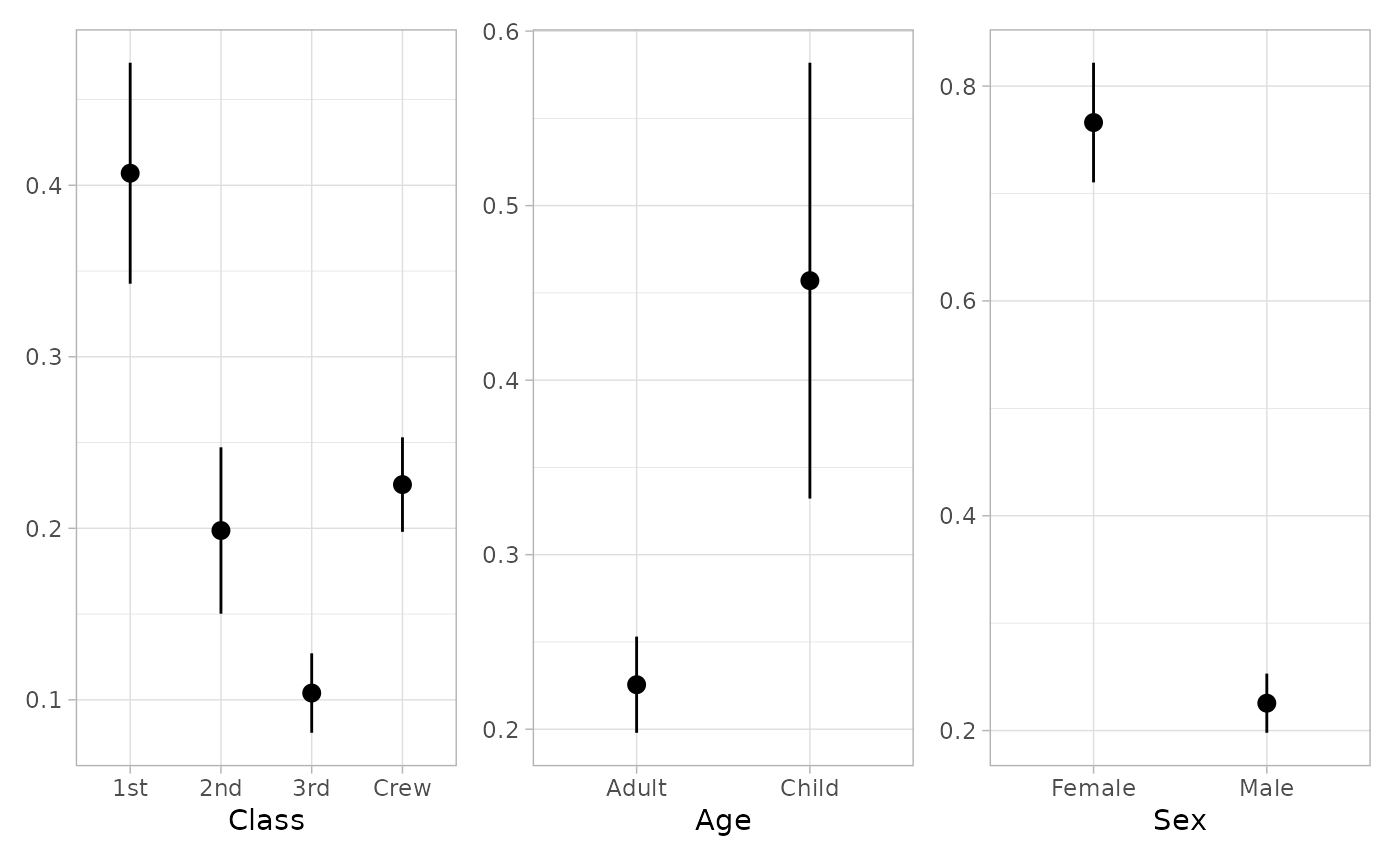
if (require("patchwork")) {
plot_marginal_predictions(mod, newdata = "mean") |>
patchwork::wrap_plots()
}
tidy_marginal_predictions(
mod3,
variables_list = list(
list(Class = unique, Sex = "Female"),
list(Age = unique)
)
)
#> variable term estimate std.error statistic p.value
#> 1 Class:Sex 1st * Female 0.8980680 0.015940375 56.33920 0.000000e+00
#> 2 Class:Sex 2nd * Female 0.7580905 0.030941812 24.50052 1.458340e-132
#> 3 Class:Sex 3rd * Female 0.5904013 0.031062754 19.00673 1.500210e-80
#> 4 Class:Sex Crew * Female 0.7978123 0.026646078 29.94108 5.749357e-197
#> 5 Age Adult 0.3142777 0.008706739 36.09591 2.629051e-285
#> 6 Age Child 0.6033792 0.050063958 12.05217 1.889174e-33
#> s.value conf.low conf.high
#> 1 Inf 0.8668255 0.9293106
#> 2 437.9502 0.6974457 0.8187354
#> 3 265.1691 0.5295195 0.6512832
#> 4 651.8964 0.7455869 0.8500376
#> 5 945.3550 0.2972128 0.3313426
#> 6 108.7059 0.5052556 0.7015027
# Marginal Predictions at the Mean
tidy_marginal_predictions(mod, newdata = "mean")
#> variable term estimate std.error statistic p.value s.value
#> 1 Class 1st 0.4070382 0.03286740 12.384253 3.180310e-35 114.59832
#> 2 Class 2nd 0.1987193 0.02474435 8.030897 9.676261e-16 49.87640
#> 3 Class 3rd 0.1039594 0.01182240 8.793426 1.450667e-18 59.25799
#> 4 Class Crew 0.2254997 0.01405834 16.040279 6.685371e-58 189.93082
#> 5 Age Adult 0.2254997 0.01405834 16.040279 6.685371e-58 189.93082
#> 6 Age Child 0.4570172 0.06370387 7.174088 7.279069e-13 40.32131
#> 7 Sex Female 0.7660538 0.02841756 26.957057 4.714993e-160 529.27124
#> 8 Sex Male 0.2254997 0.01405834 16.040279 6.685371e-58 189.93082
#> conf.low conf.high
#> 1 0.34261928 0.4714571
#> 2 0.15022129 0.2472174
#> 3 0.08078793 0.1271309
#> 4 0.19794588 0.2530536
#> 5 0.19794588 0.2530536
#> 6 0.33215989 0.5818745
#> 7 0.71035641 0.8217512
#> 8 0.19794588 0.2530536
if (require("patchwork")) {
plot_marginal_predictions(mod, newdata = "mean") |>
patchwork::wrap_plots()
}
 # }
# }